For molecular modelers, analyzing atom trajectories along defined pathways can be crucial for revealing complex molecular mechanisms like ligand binding/unbinding or reaction dynamics. SAMSON, the integrative molecular design platform, simplifies this process with the Export Along Paths extension. Whether you’re preparing data for reaction coordinate studies, free energy calculations, or visualizing ligand pathways, this tool allows you to export atomic coordinates efficiently and with precision. Here’s an in-depth look at how to use it.
Why Should You Export Atom Trajectories?
Molecular modeling often requires generating detailed data to understand interactions and dynamics. Exported trajectories are used to:
- Create reaction coordinate files for accurate free energy profiling.
- Visualize ligand pathways to study ligand unbinding or entry dynamics.
- Focus on the dynamics of specific atoms, such as a ligand or binding site residues.
SAMSON’s Export Along Paths extension provides you with a user-friendly way to achieve this.
Getting Started
To begin, ensure the extension is added to your SAMSON workspace:
- Log into your SAMSON Connect account.
- Visit the Export Along Paths Extension page and click Add.
- Restart SAMSON to activate the extension.
Steps to Export Atom Trajectories
1. Load a Sample System
If you’re new to exporting trajectories, SAMSON offers a pre-configured sample system that simplifies the learning process:
- Click Home > Download and paste the sample document link: https://www.samson-connect.net/documents/046f1acd-c799-40f6-8185-cb4847eff795.
- Click Download.
This sample system contains Lactose permease (1PV7), the ligand Thiodigalactosid (TDG), and unbinding paths generated with Ligand Path Finder. This setup is ideal for practice and experimentation.

2. Open the Export Along Paths Tool
Access this tool via the menu: Home > Apps > All > Export Along Paths. Alternatively, use the command shortcut (Shift+E) to locate it through the search interface.
You’ll see an easy-to-navigate interface guiding you through the export process: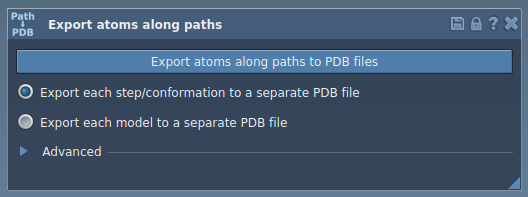
3. Export Atom Trajectories
Option A: Export All Atoms
- Select one or more paths in the Document view.
- Choose the export mode (e.g., all frames in a single PDB file or each frame as a separate PDB file).
- Click Export atoms along paths to PDB files, and specify a destination folder and file prefix.
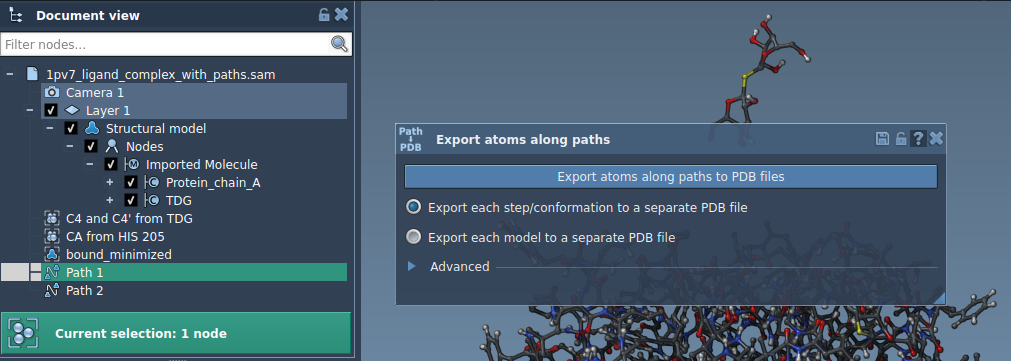
Option B: Export a Subset of Atoms
- Expand the Advanced section in the app interface.
- Select a subset (e.g., ligand
TDG) in the Document view, and add it to the export list. - Optionally, define multiple models (sets of atoms) for export and rename them as needed.
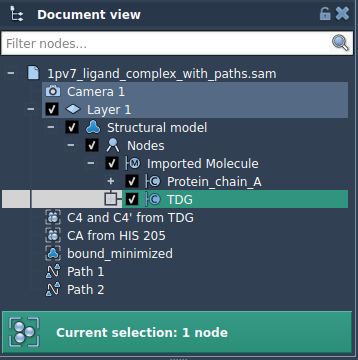
When ready, select the paths to export and click Export atoms along paths to PDB files. Define a destination folder and file prefix to complete the process.
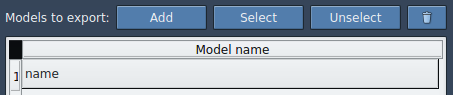
Learn More
Exporting atom trajectories is an essential step for various molecular modeling applications. To delve deeper into SAMSON’s capabilities and advanced export options, visit the full documentation at this link.
SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON at https://www.samson-connect.net.





