Molecular docking can be a critical step in understanding protein-ligand interactions and predicting binding affinities. However, setting up ligands properly—whether single ligands or libraries—can often seem daunting. In this blog post, we’ll take a deep dive into the process of ligand setup using the AutoDock Vina Extended SAMSON Extension, empowering you to streamline your workflow and avoid common pitfalls.
Why Proper Ligand Setup Matters
The accuracy of molecular docking not only depends on the software but also on how well the ligand has been prepared. Issues like incorrect bond rotation settings, missing hydrogens, or non-optimized 2D ligands can significantly impact docking results. Thankfully, the AutoDock Vina Extended SAMSON Extension provides intuitive tools to address these challenges while maintaining flexibility for advanced configurations.
Setting Up a Single Ligand
The process begins in the Set ligand section of the AutoDock Vina Extended app. To set up a single ligand, select the ligand (for example, 2AZ8-IA) within the Document view and click the associated Set button. After this, the system provides visible rotatable bond controllers in the Viewport.
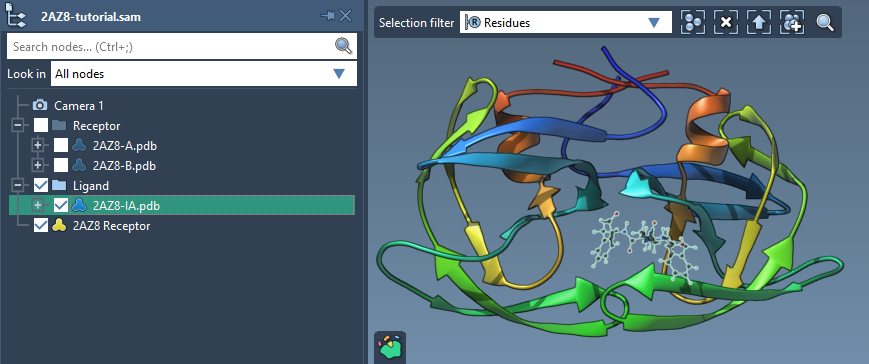
Customizing Rotatable Bonds
Rotatable bonds are represented by green cylinders superimposed on bonds, making it easy to identify and toggle their rotation state. Bonds can be toggled between rotatable (green) and locked (red) simply by clicking on them. This customization lets you precisely control the flexibility of your ligand, catering to docking scenarios where certain bonds must remain fixed for biological reasons.
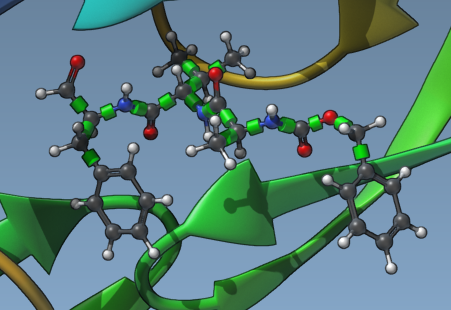
You can also universally lock specific bond types via the Locked bonds settings menu. Once you define bond types to lock, enable the Lock specific ligand bonds option for these to be applied uniformly across your ligand.
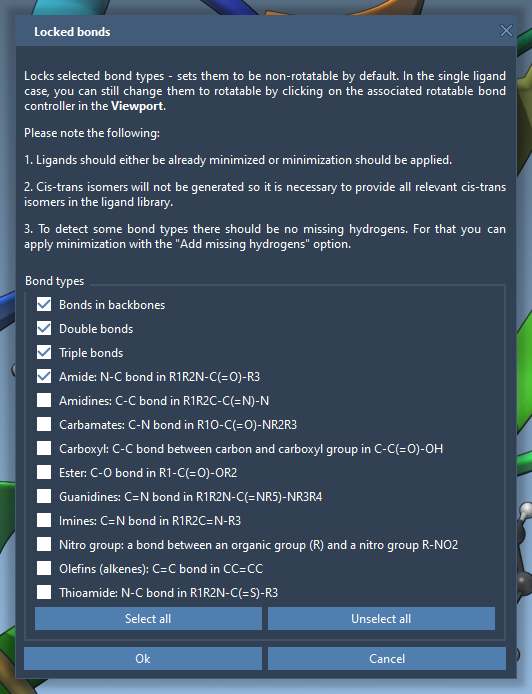
Preparing Ligand Libraries
If you’re working with a collection rather than a single ligand, AutoDock Vina Extended can help you set up a ligand library with ease. This approach supports formats such as MOL2 and SDF, and multiple ligands can be stored in a single file. Simply select Ligand library in the Set ligand section, navigate to your library directory, and AutoDock Vina Extended will import the ligands.
By default, all bonds in the library will be treated as rotatable. You can adjust this setting globally using the same Lock specific ligand bonds options you would for single ligands. If hydrogens are absent, you can instruct the system to add them during setup by enabling Add missing hydrogens in the Minimization menu.
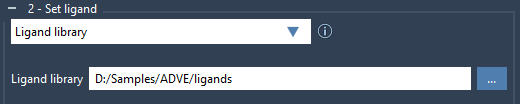
Optimizing Ligands via Minimization
Before docking, ensuring that ligands are in energetically favorable conformations is vital. The Minimize option, available during setup, uses pre-defined presets to optimize ligands while also checking for polar hydrogens—a requirement for accurate docking. The minimization also stops automatically once specific energy thresholds or step limits are met, making it highly efficient for large libraries or complex molecules.

Conclusion
The ligand setup features in AutoDock Vina Extended allow for meticulous preparation without adding complexity. Whether you’re dealing with a single compound or a diverse library, the extension ensures your ligands are ready for high-quality docking simulations.
To explore advanced workflows and features of AutoDock Vina Extended, refer to the official documentation.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON at samson-connect.net.





