For molecular modelers, accurately determining transition paths and saddle points between two conformational states can be quite challenging. These are critical calculations for understanding reaction mechanisms or conformational changes in molecules, yet they often come with computational complexities and setup headaches. Fortunately, the Parallel Nudged Elastic Band (P-NEB) method offers a robust way to streamline these optimizations within the SAMSON platform.
In this blog post, we’ll introduce how the P-NEB method works, explain key features, and show how to use it effectively to minimize energy pathways. By the end of this guide, you’ll be equipped to apply P-NEB to your own molecular systems with confidence.
What is P-NEB?
The P-NEB app is a versatile tool that uses the Nudged Elastic Band (NEB) method, with parallel processing capabilities, to optimize transition paths. This is particularly useful for finding the minimum energy path (MEP) connecting two states of a system, such as ligand unbinding pathways in proteins or conformational transitions in organic molecules.
The app maintains equal spacing of intermediate images through simulated spring forces, ensuring accurate interpolation. It also provides the option to apply the “climbing image” strategy to pinpoint saddle points more accurately, which are crucial for transition state analysis.
Preparation Steps
Before launching into P-NEB, there are some preparatory steps to follow:
- Make sure to install the P-NEB Extension via SAMSON Connect.
- Prepare your molecular system by relaxing the initial and final states using a minimizer such as FIRE.
- Generate a sequence of intermediate conformations between relaxed states using linear interpolation or tools like Ligand Path Finder.
If the intermediate conformations already form a continuous path, the process will be faster and more efficient.
Running the P-NEB App
To start optimization:
- In SAMSON, open the P-NEB app from Home > Apps > All > P-NEB.
- Set the key parameters:
- Spring constant: typically set to 1.00.
- Number of loops: enter the number of iterations, e.g., 100.
- Interaction model: choose a force field such as “Universal Force Field (UFF).”
- Optimizer: select “FIRE” for efficient optimization.
- Enable Parallel execution to speed up the process.
- Optionally, leave the “Climbing image method” unchecked initially for a smoother first pass.
- Select either a path or a set of conformations in the Document view and hit the Run button in the app.
The app then initializes and begins optimization, with progress visualized in SAMSON’s status bar. Once complete, a new optimized path or set of conformations appears in the Document view for further examination.
Benefits and Features
Here’s why P-NEB is valuable:
- Efficiency: Using parallel processing, P-NEB significantly speeds up calculations.
- Flexibility: Apply P-NEB to continuous paths or unconnected conformations.
- Insightful Output: The app not only produces optimized results but also provides a detailed summary of the computation in its interface.
If you aim to improve ligand unbinding pathways or any conformational transition paths, the P-NEB method offers a structured, efficient approach to achieve accurate results.
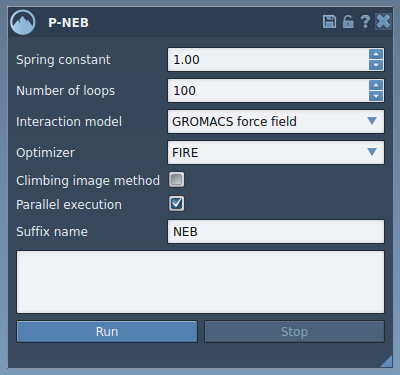
This image shows the main interface of P-NEB, where all the optimization settings can be configured before running computations. As you become familiar with the parameters, you can fine-tune them for your specific use case.
Get Started Today
The P-NEB app simplifies a complicated yet critical task in molecular modeling. From ligand unbindings to dynamic conformational studies, this tool is invaluable for reducing energy pathways while maintaining accuracy.
To dive deeper into the technical details, visit the official documentation page and explore how P-NEB can be applied to your systems!
SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON at SAMSON Connect.





