Preparing data for complex molecular dynamics analyses often involves a significant amount of effort to extract atomic trajectories along defined pathways. For molecular modelers performing reaction coordinate studies or free energy calculations, this process can be tedious. Did you know SAMSON provides a streamlined solution for this via the “Export Along Paths” extension?
The Problem
When studying molecular interactions, such as ligand unbinding or protein conformational changes, researchers often need to analyze atomic paths. Traditional methods to export this data may involve extensive scripting or external utilities, which can become time-intensive and error-prone. The challenge escalates when the scope includes specific atom groups or repeated testing of alternate paths.
The SAMSON Solution
SAMSON’s “Export Along Paths” extension directly addresses this challenge by allowing you to export atomic positions along predefined paths efficiently. Whether you need the coordinates for all atoms or a specific subset, like ligands or binding site atoms, this powerful tool ensures an intuitive and flexible workflow for extracting the data you need.
How It Works
Once you have installed the “Export Along Paths” extension, a simple 3-step workflow lets you export the desired data:
- Step 1: Open the “Export Along Paths” App
Access the app through Home > Apps > All > Export Along Paths or via the shortcut (Shift + E).

- Step 2: Choose Your Export Parameters
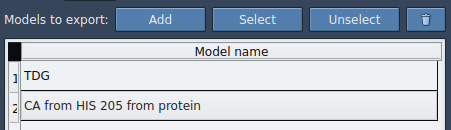
Within the app interface, you can select one or more predefined paths and decide whether to export all atoms or only a subset. For example, a ligand such asTDGcan be chosen in the Document view. You can rename models, add multiple sets of atoms, and customize the frequency of frames to export. This gives unparalleled flexibility.
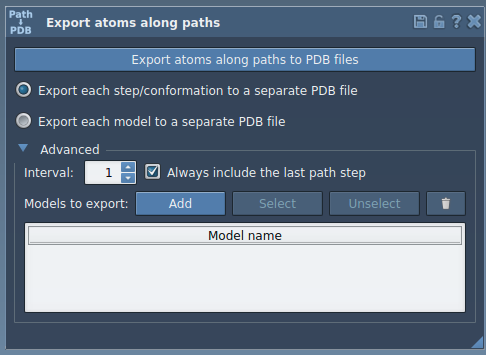
- Step 3: Export and Save
After setting your preferences, you can select the desired output format: either a single PDB file containing all trajectory frames or separate files per frame. Click Export atoms along paths to PDB files. You’ll be prompted to choose a destination folder and file prefix. Voila – you’re done!
Why Use the Export Along Paths Extension?
- Streamlined Reaction Coordinate Files: Generate files for free energy profiling with minimal manual overhead.
- Enhanced Sampling Analysis: Efficiently export ligand entry and exit trajectories for further computational studies.
- Customized Data Selection: Choose specific subsets, like a ligand or protein residues, without complex filtering.
- Flexible Formats: Save trajectories in ways that suit your downstream workflows.
For molecular researchers looking to streamline this critical step in their workflows, the “Export Along Paths” extension is a convenient and reliable solution. Learn more about this functionality and how it fits into your computational modeling by visiting the official documentation page.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Download SAMSON here.





