Molecular modelers often face the challenge of determining realistic transition paths between molecular states, such as ligand binding or unbinding, protein conformational changes, or other dynamic processes. These paths can provide key insights into the mechanisms and energetics of molecular interactions. However, finding and refining meaningful transition paths is not always straightforward.
If you have an initial set of molecular conformations or a rough pathway but need to refine it to generate a more physically meaningful result, SAMSON’s Parallel Nudged Elastic Band (P-NEB) method provides an accessible and systematic approach to achieve this goal. Here’s how it works and how you can apply it to your modeling tasks.
What is P-NEB?
In SAMSON, the P-NEB app is an extension that optimizes transition paths by iteratively refining intermediate molecular conformations. It uses the Nudged Elastic Band (NEB) method, which maintains uniform spacing between neighboring conformations through spring forces. It also minimizes the system’s energy to produce a transition path that adheres to physical principles.
This method is particularly relevant for studying paths between known energy minima, for example, ligand unbinding trajectories or protein conformational transitions.
Why Use P-NEB?
Imagine you have a trajectory of a molecule moving through a complex environment (e.g., a ligand leaving a protein active site). A rough approximation of the path might reveal the general direction of the motion, but such paths often suffer inaccuracies due to uneven distribution of conformations or high-energy barriers. P-NEB resolves these problems by:
- Improving the distribution of molecular states along the path.
- Providing a way to refine trajectories into smoother, physically meaningful transitions.
- Allowing integration of force fields like Universal Force Field (UFF) for accurate energy calculations.
How to Get Started with P-NEB
Here’s a step-by-step walkthrough of how you can use the P-NEB app to refine a transition path in SAMSON:
1. Install the Necessary Extensions
Before starting, ensure that you’ve added the P-NEB extension and the FIRE state updater to SAMSON. If needed, you can install these directly from SAMSON’s Extensions platform.
2. Load Your Input Model
To conduct an optimization, you must have a candidate path or set of molecular conformations prepared. For example, you can load pre-existing paths, such as a sample Zinc ligand unbinding trajectory or a protein-ligand complex path. These can be downloaded from SAMSON Connect as illustrated below:

3. Launch the P-NEB App
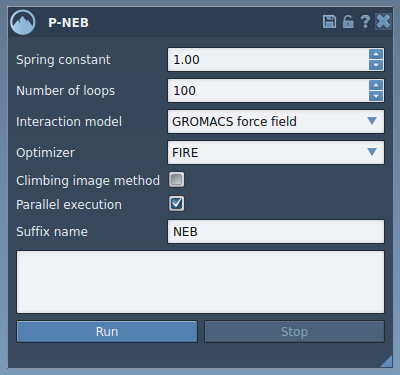
The P-NEB interface is accessible via Home > Apps > All > P-NEB in SAMSON. The interface allows you to fine-tune various optimization parameters, such as:
- Spring Constant: Governs spring forces between conformations, typically set to 1.00.
- Number of Loops: Determines the number of optimization iterations. A good starting value is 100.
- Climbing Image Method: Helps locate saddle points; you can enable this for further refinement.
- Parallel Execution: Check this option to utilize parallel threads, improving computation speed.
4. Optimize Your Path
Once your parameters are configured, click the Run button in the P-NEB app to start optimization. The system will process the data using the Universal Force Field, and computation progress can be tracked via the status bar. Upon completion, the refined transition path will appear in the SAMSON document view:
You can visually inspect the optimized path, animate it to see the refined trajectory, or use it for subsequent free-energy calculations or simulations.
Tips for Effective Use
For optimal performance, consider the following tips:
- If you start with a set of conformations rather than a path, you can first group these conformations into a continuous path using the option Conformation > Create path from conformations in the context menu.
- You can experiment with the “Climbing Image Method” to locate the exact saddle point on the path for advanced applications.
For a detailed guide on P-NEB, visit the official documentation page.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON at SAMSON Connect.





