Molecular modeling often requires exploring how small changes in a molecule’s structure affect its properties, like binding affinity or interactions with target proteins. However, manually generating molecular analogs for such studies can be tedious and time-consuming. Enter positional analogue scanning, a streamlined method available in SAMSON that simplifies this process and empowers molecular modelers to quickly evaluate the impact of structural modifications. This blog post will guide you through the essentials of using positional analogue scanning with the SMILES Manager in SAMSON.
What is Positional Analogue Scanning?
Positional analogue scanning involves systematically modifying parts of a molecule to generate new analogs. This approach is particularly useful for understanding how specific atomic or functional group changes influence a molecule’s properties. With SAMSON’s SMILES Manager extension, you can carry out this process efficiently and directly within the platform. For example, you could evaluate changes in binding affinities or key interactions of your molecular designs without switching between tools.
Step-by-Step Guide to Positional Analogue Scanning in SAMSON
1. Define Your Starting Molecule
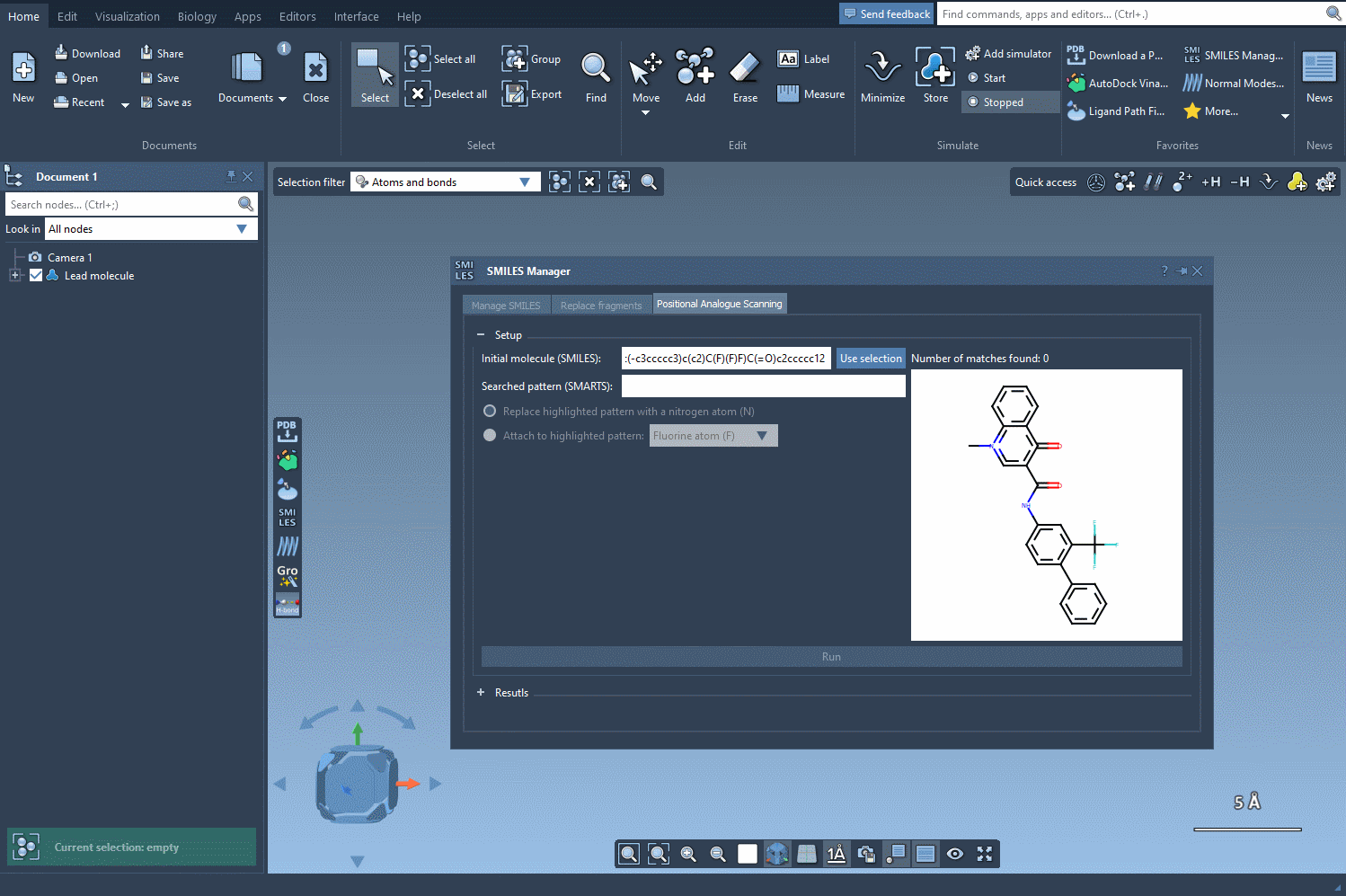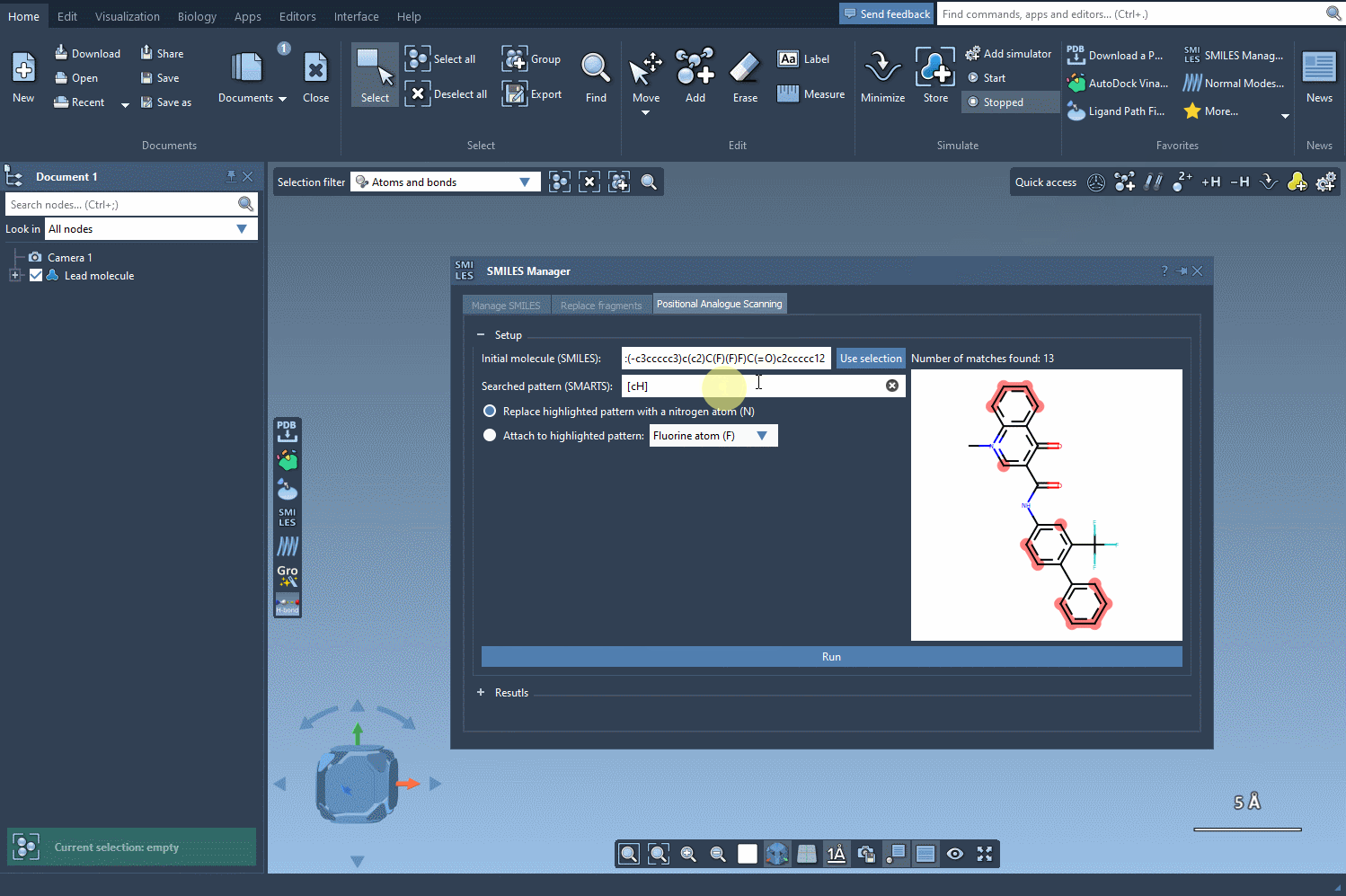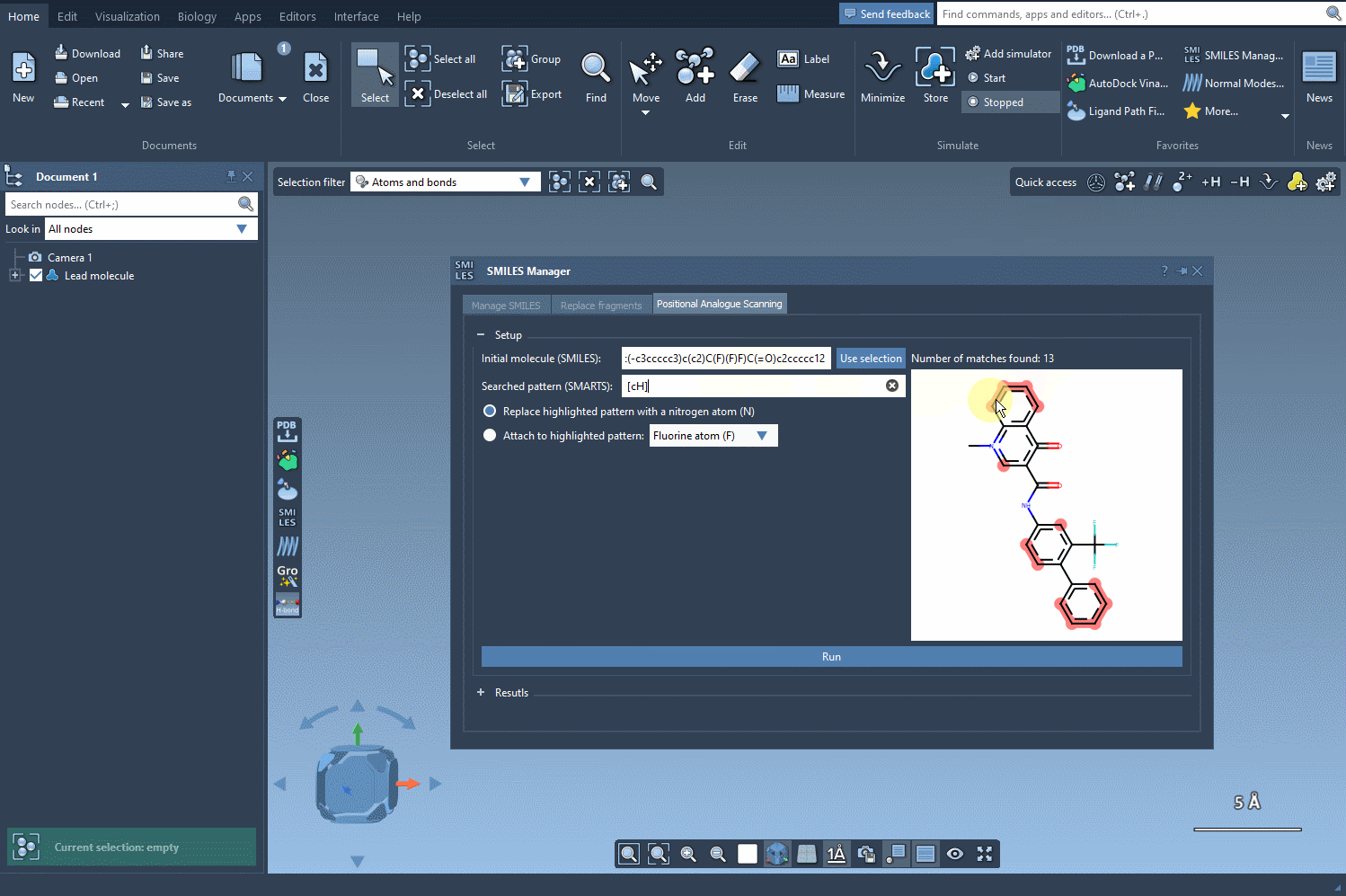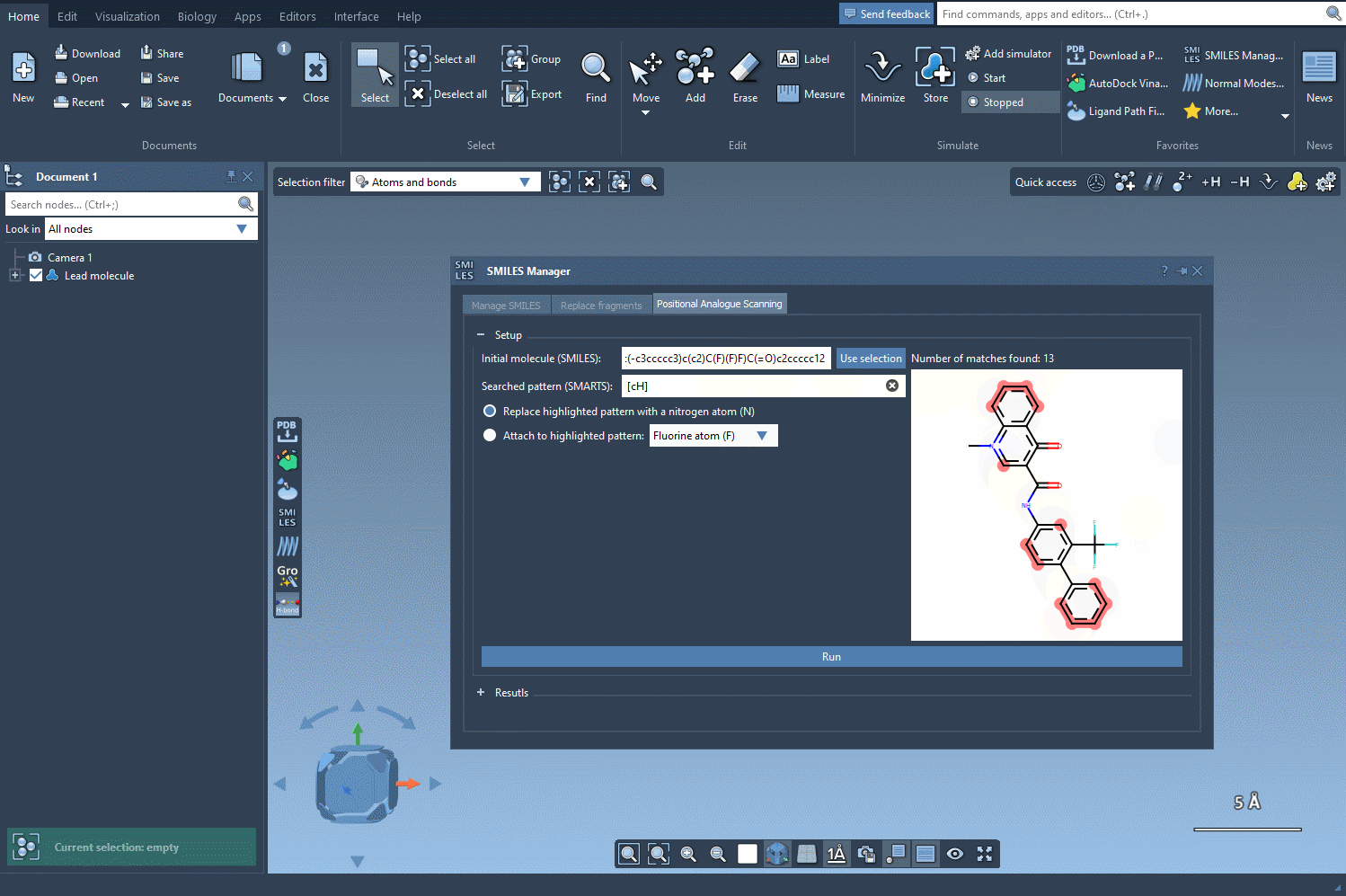
Begin by selecting your molecule of interest. You can either input its SMILES code or select it directly within the SAMSON interface and press the Use selection button. A visual demonstration is available below:
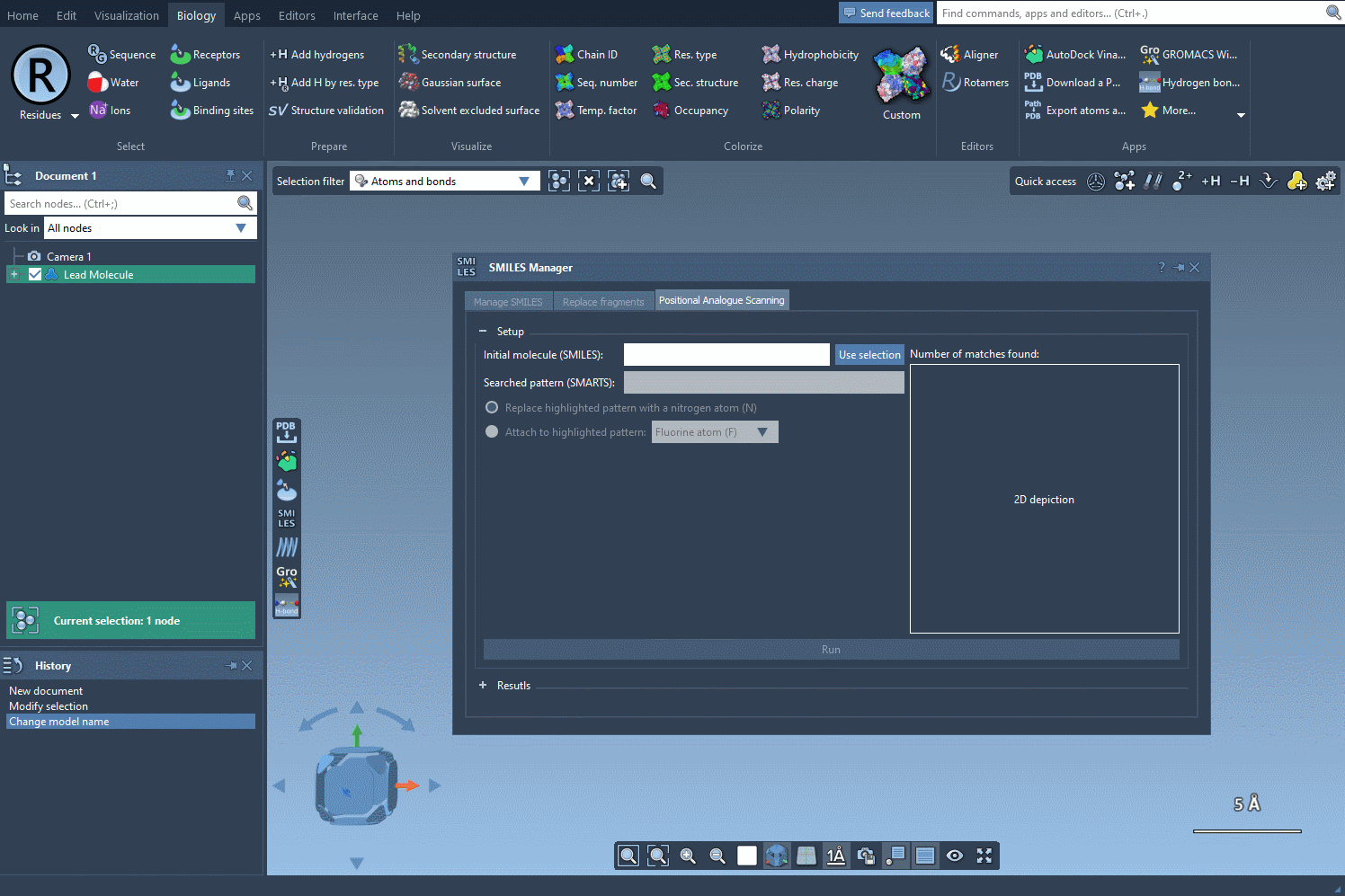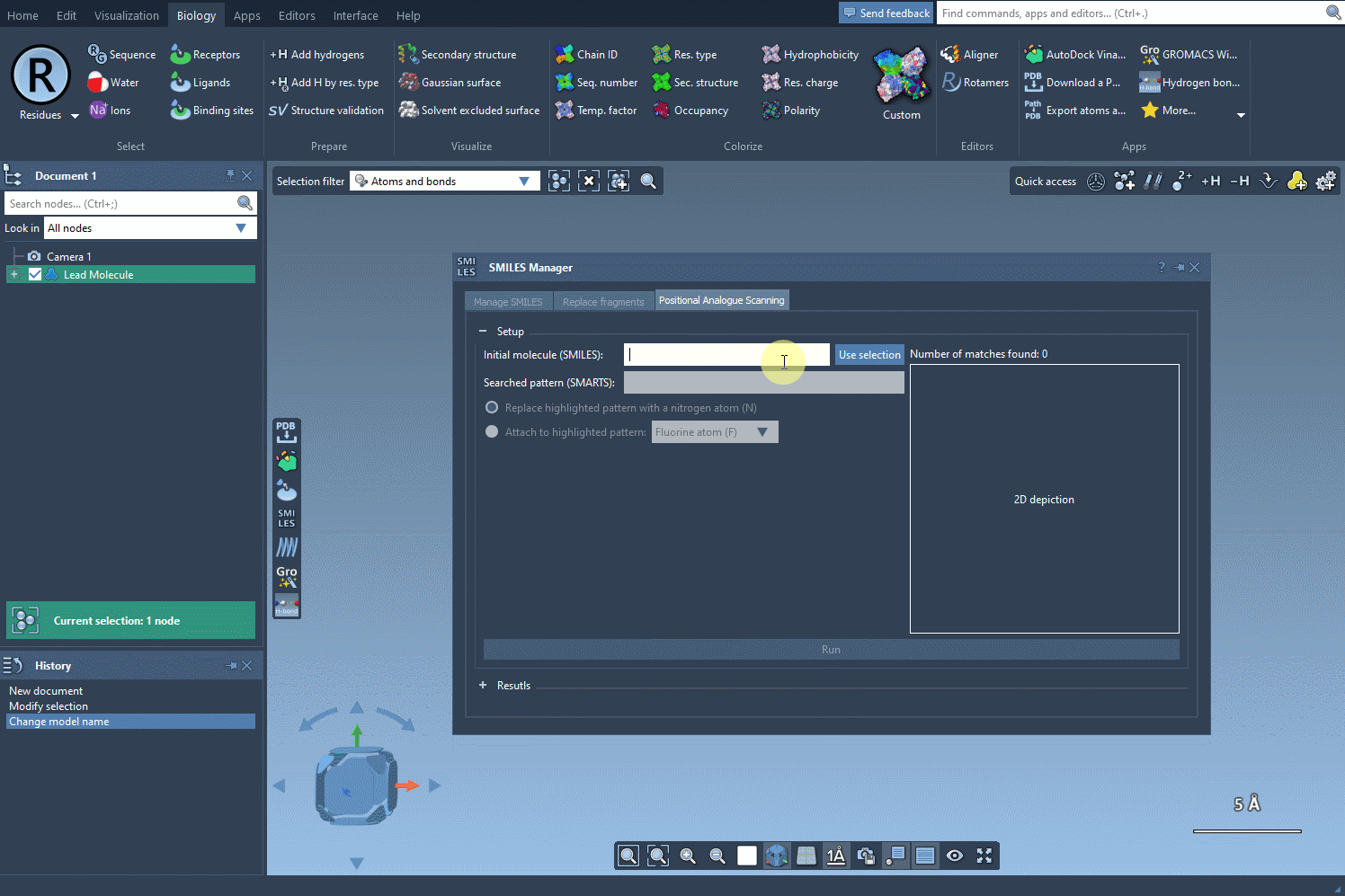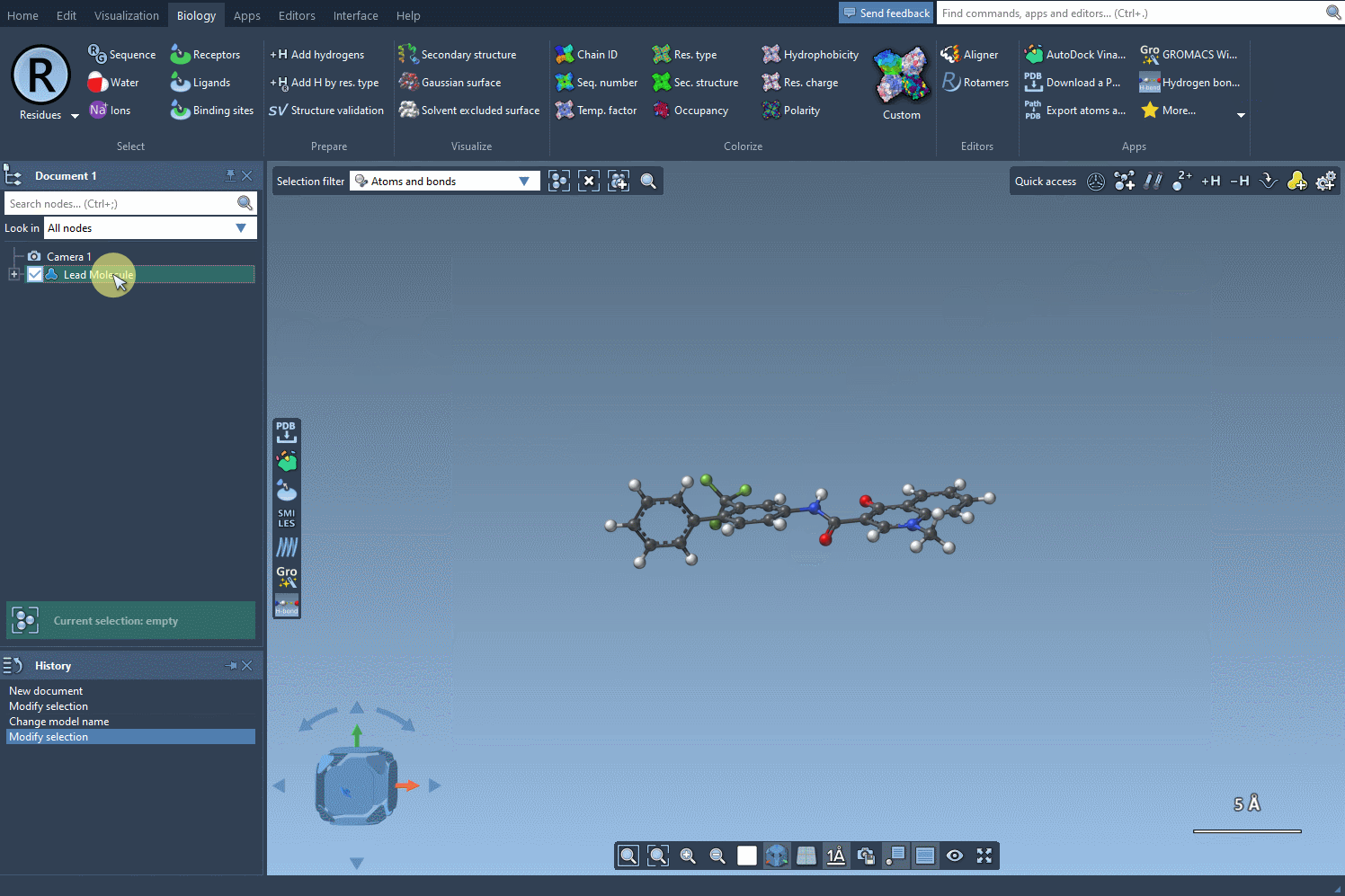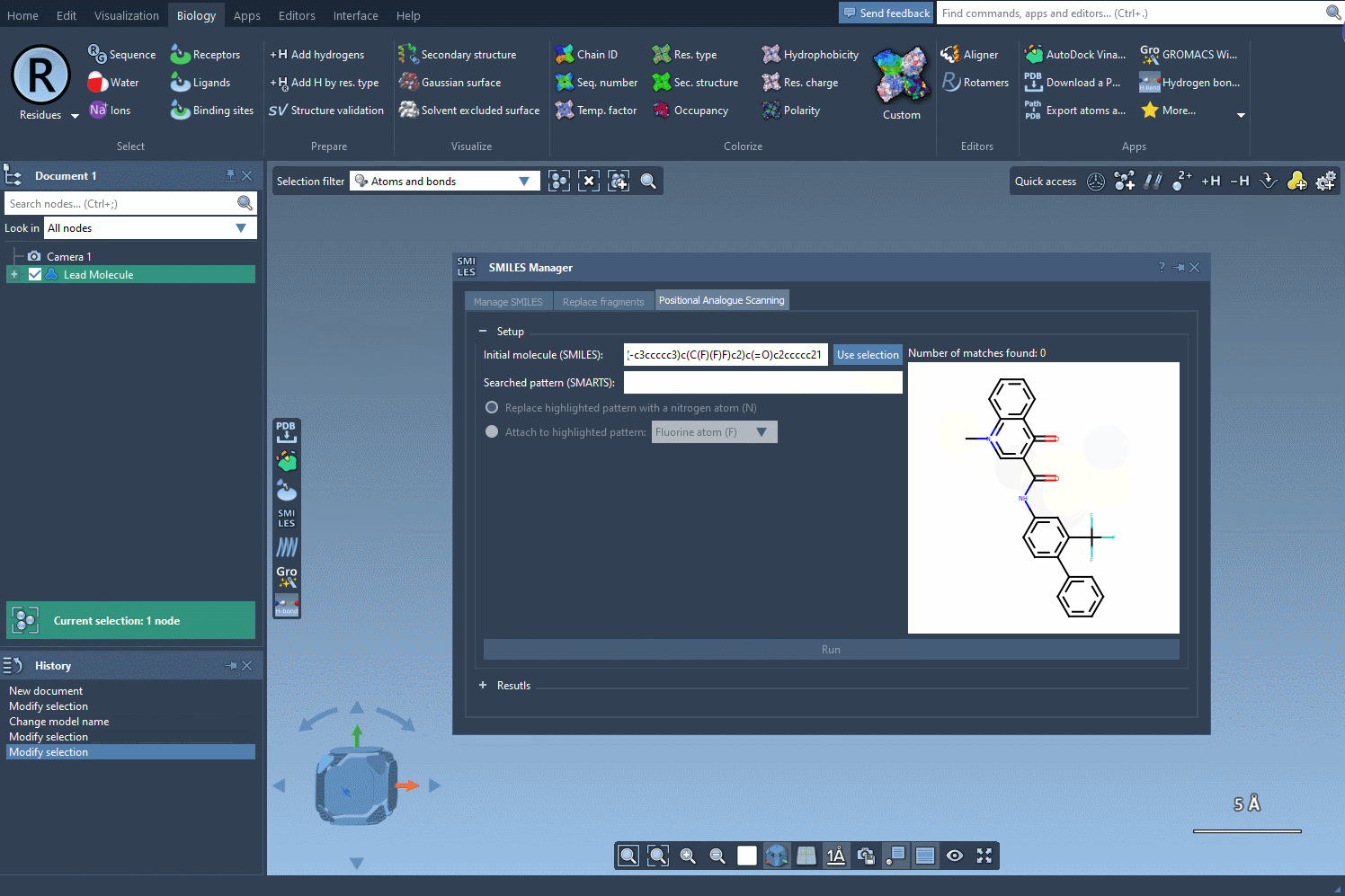
2. Specify the Pattern to Modify
The next step is to define the pattern within your molecule to alter. For instance, you might want to modify aromatic carbon atoms. To do this, enter the SMARTS code for your pattern, such as [cH], in the relevant field. SAMSON will automatically highlight all matching occurrences in your molecule:
3. Modify and Generate Analogs
Once the pattern is identified, choose how to modify it. You can replace the pattern with atoms like nitrogen (N) or attach new groups such as fluorine (F) or methyl (CH3). Click Run to generate a full set of molecular analogs, complete with SMILES codes and 2D depictions. Here’s what this process looks like within SAMSON:
Explore and Refine Your Results
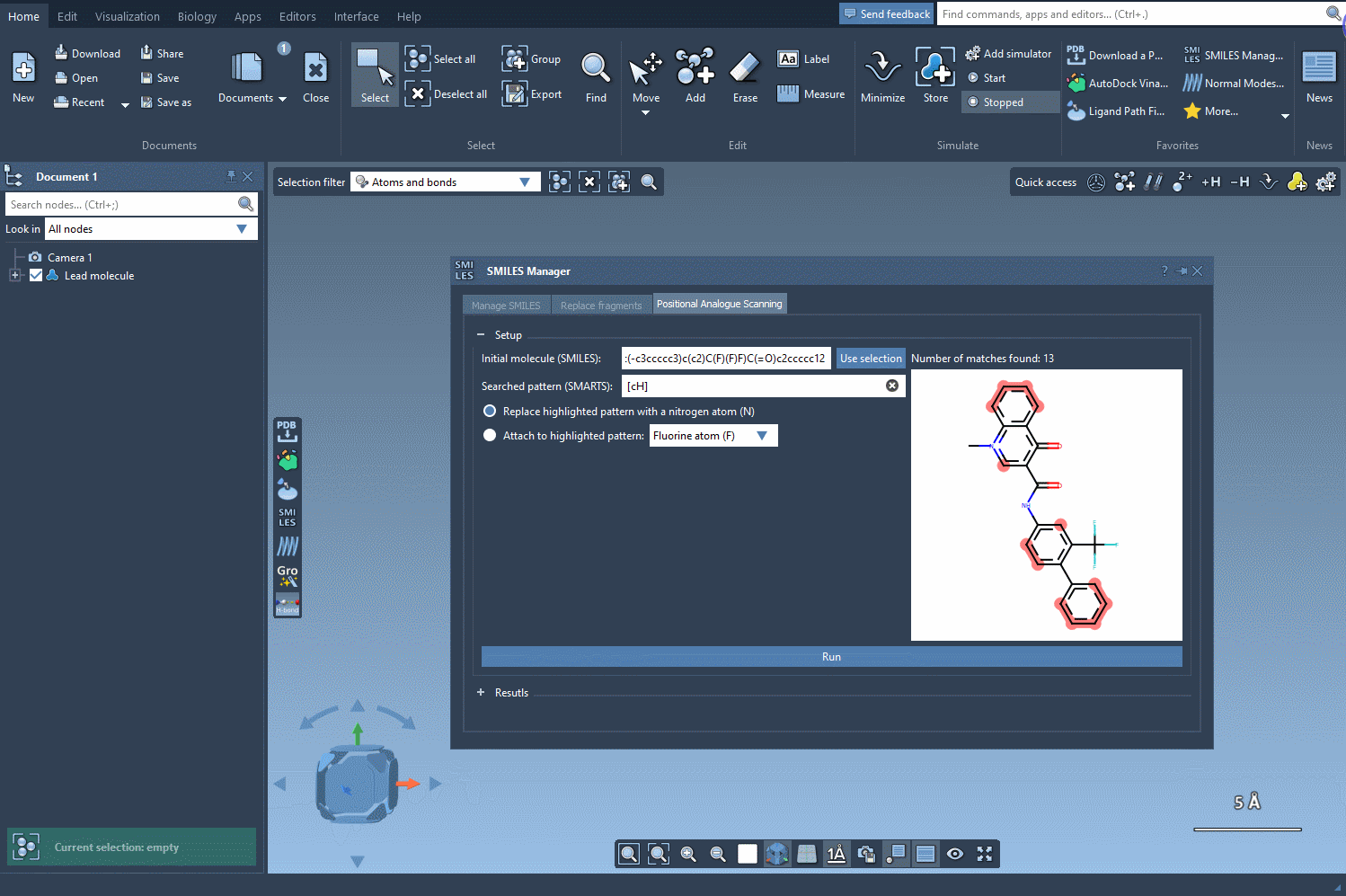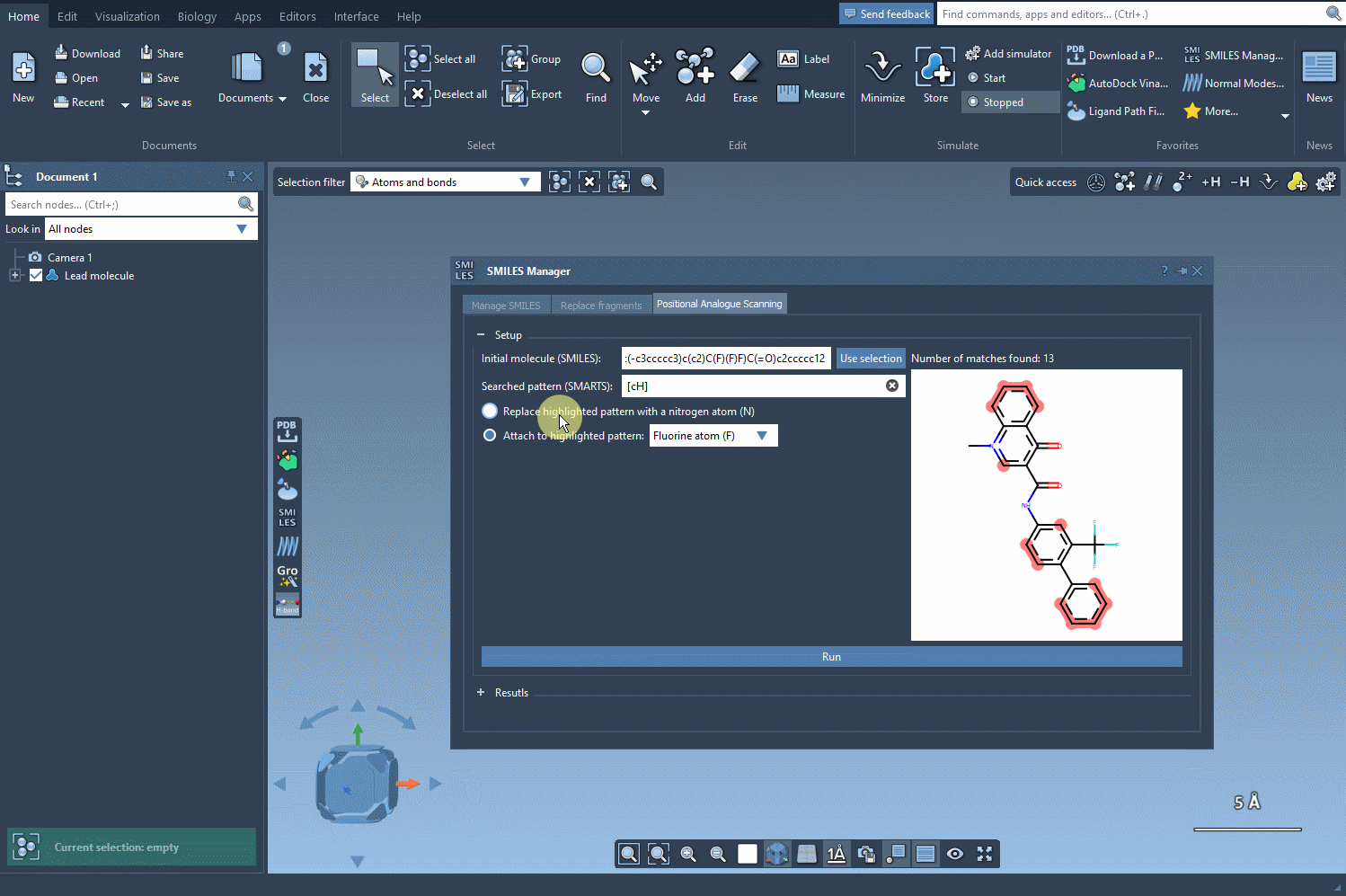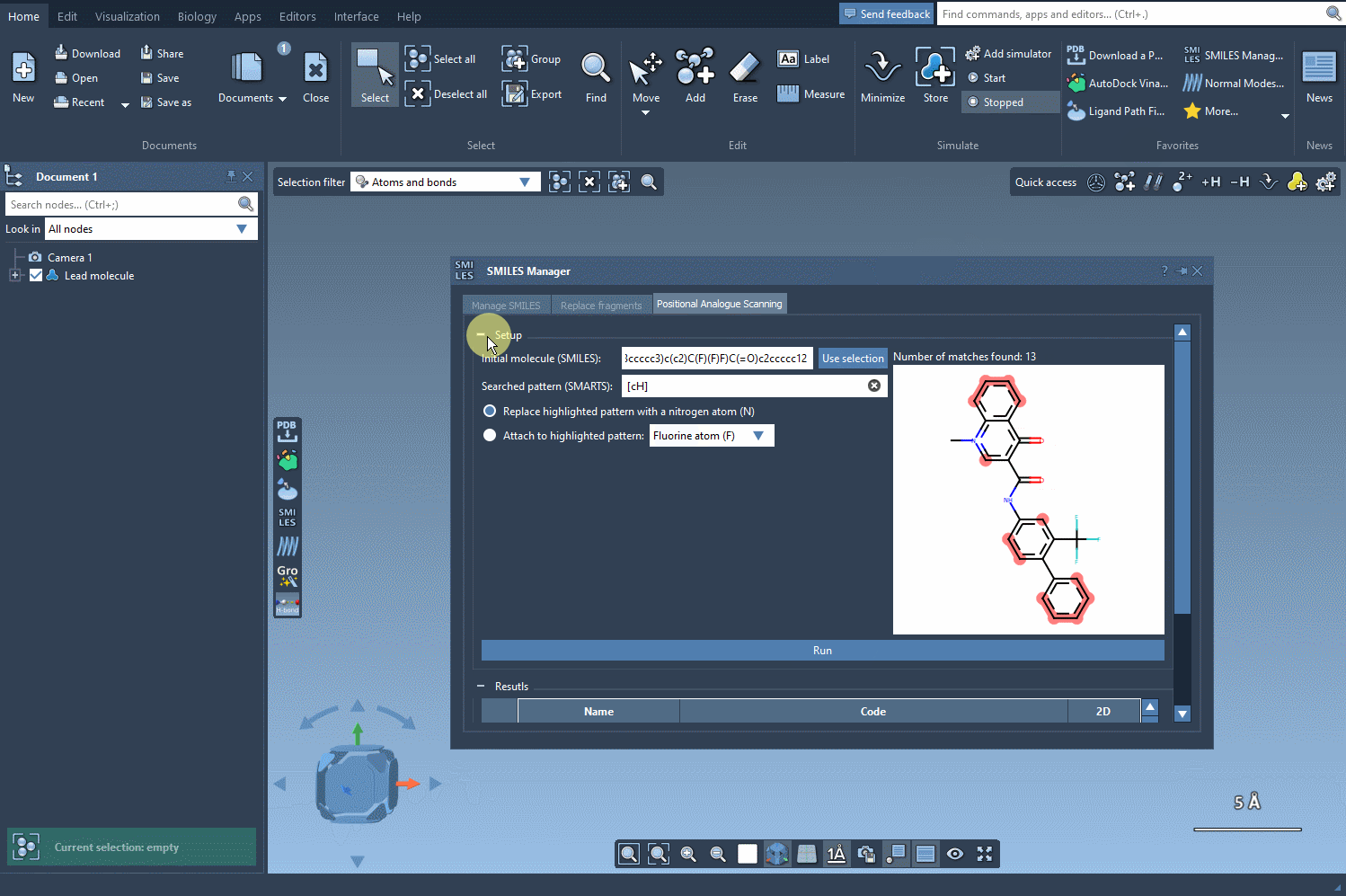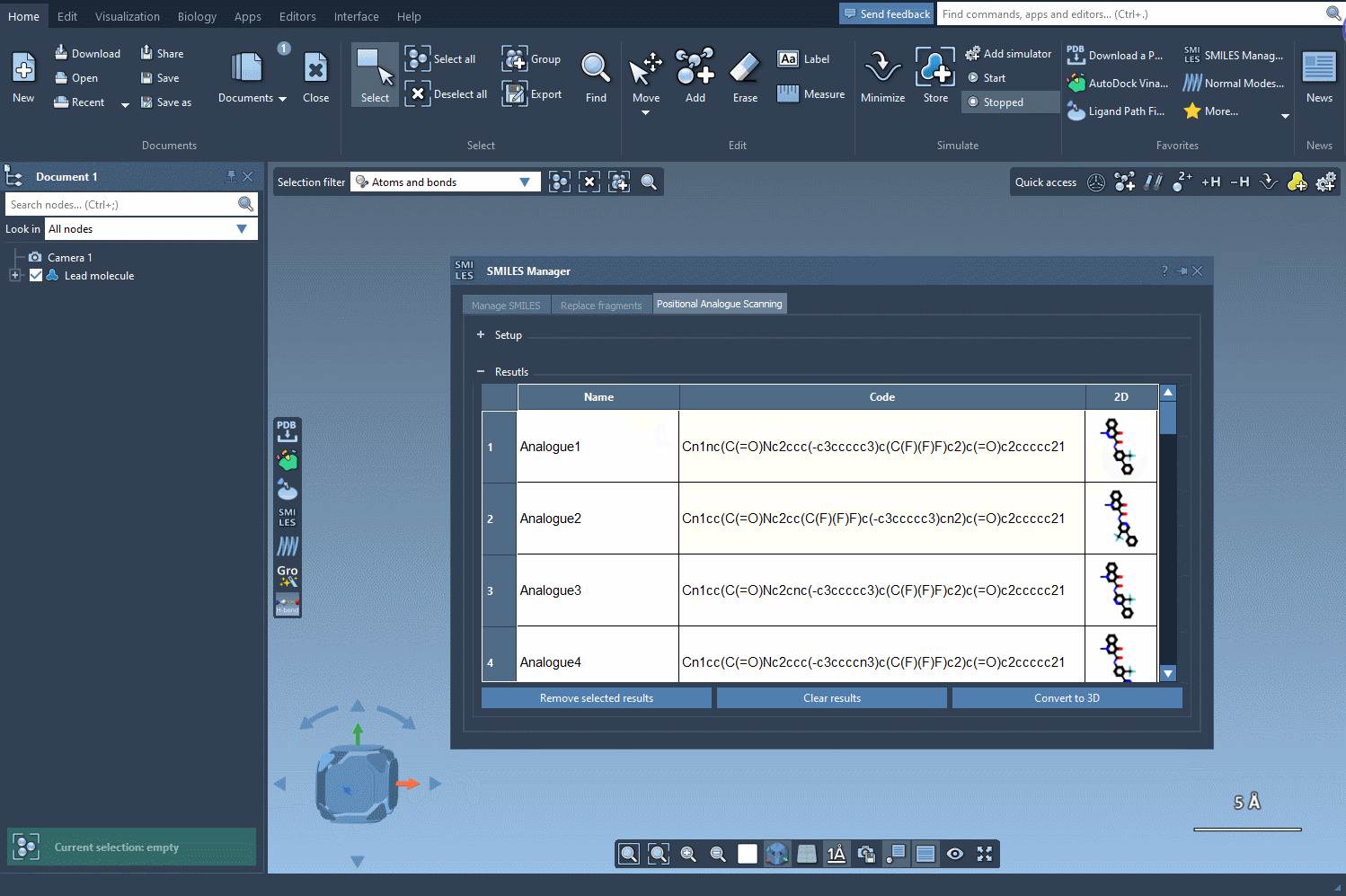
The results table provides several options to analyze and refine your analogs:
- Edit Analog Details: Double-click any cell to rename analogs or modify their SMILES codes.
- View 2D Depictions: Expand 2D images by double-clicking them or accessing the context menu.
- Generate 3D Structures: Transform selected analogs into 3D models for further study.
- Trim the Table: Remove selected analogs or clear the entire results table to start fresh.
Why Use Positional Analogue Scanning?
By leveraging positional analogue scanning, molecular modelers can save time and ensure a systematic exploration of structural modifications. The generated analogs and their properties can be studied directly within SAMSON, paving the way for deeper analyses such as docking experiments using the Autodock Vina Extended extension. This integrated approach eliminates the need for disparate tools and allows for a seamless molecular design experience.
Curious to see what else you can do? Dive deeper into the full documentation at this link.
SAMSON and all SAMSON Extensions are free for non-commercial use. Download SAMSON now at SAMSON Connect.





