In molecular modeling, exploring the effect of modifications to a molecule can be an intensive and time-consuming process. Whether you’re designing molecules for drug discovery or evaluating substituent effects, manually creating analogue series can slow down your workflow. The positional analogue scanning feature in the SMILES Manager of SAMSON provides a highly efficient and intuitive solution to this challenge.
What Is Positional Analogue Scanning?
Positional analogue scanning allows you to generate a series of molecular analogues based on a specific starting molecule and a defined substructure pattern. You start with your molecule of interest, identify key parts (using SMARTS pattern definitions), and systemically replace or modify these parts to create new analogues. This approach is particularly useful for tasks like:
- Exploring potential substituent effects.
- Preparing molecules for docking studies.
- Comparing analogue properties to select candidates for further study.
How It Works in SAMSON
Step 1: Setting the Starting Point
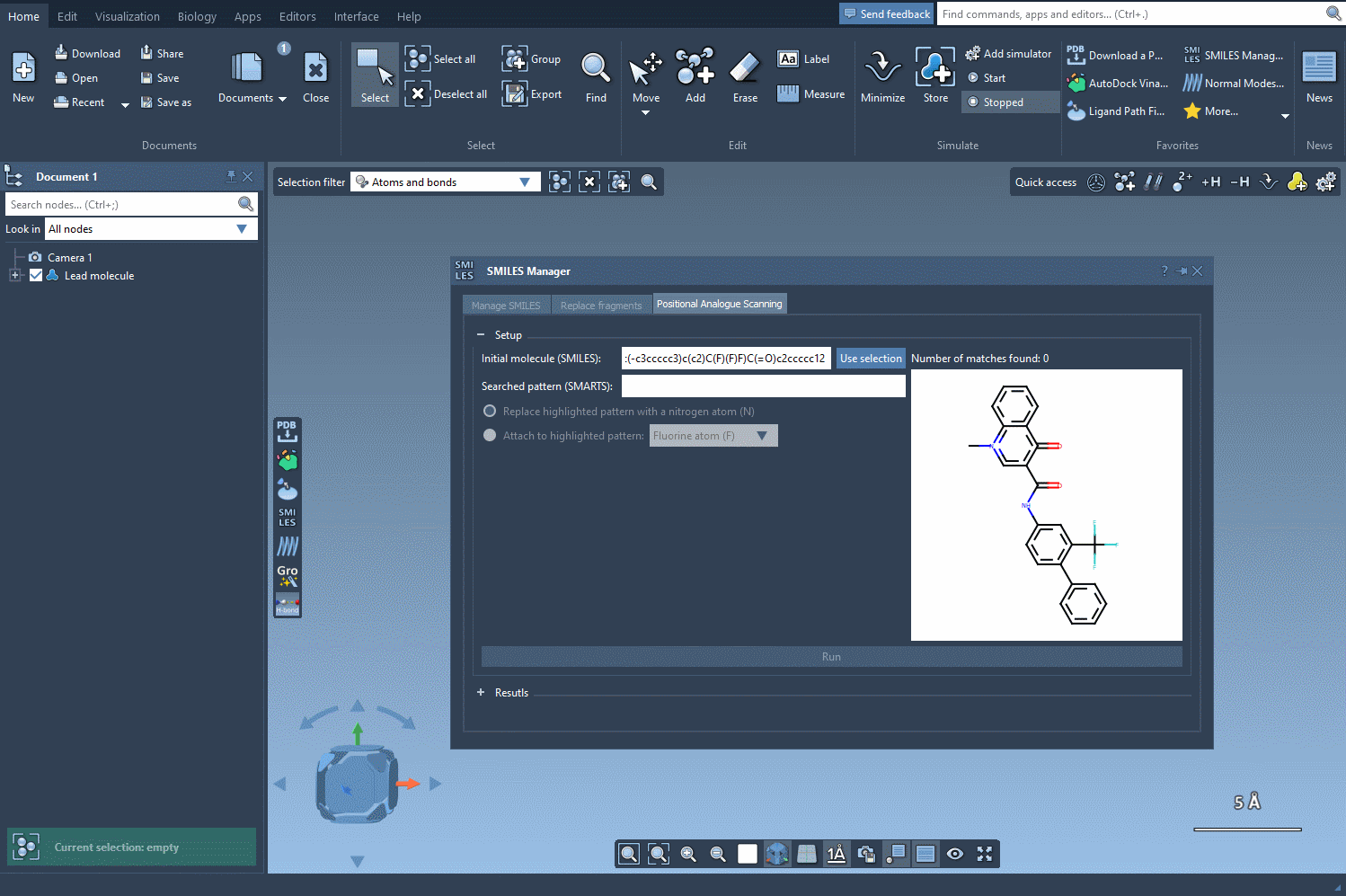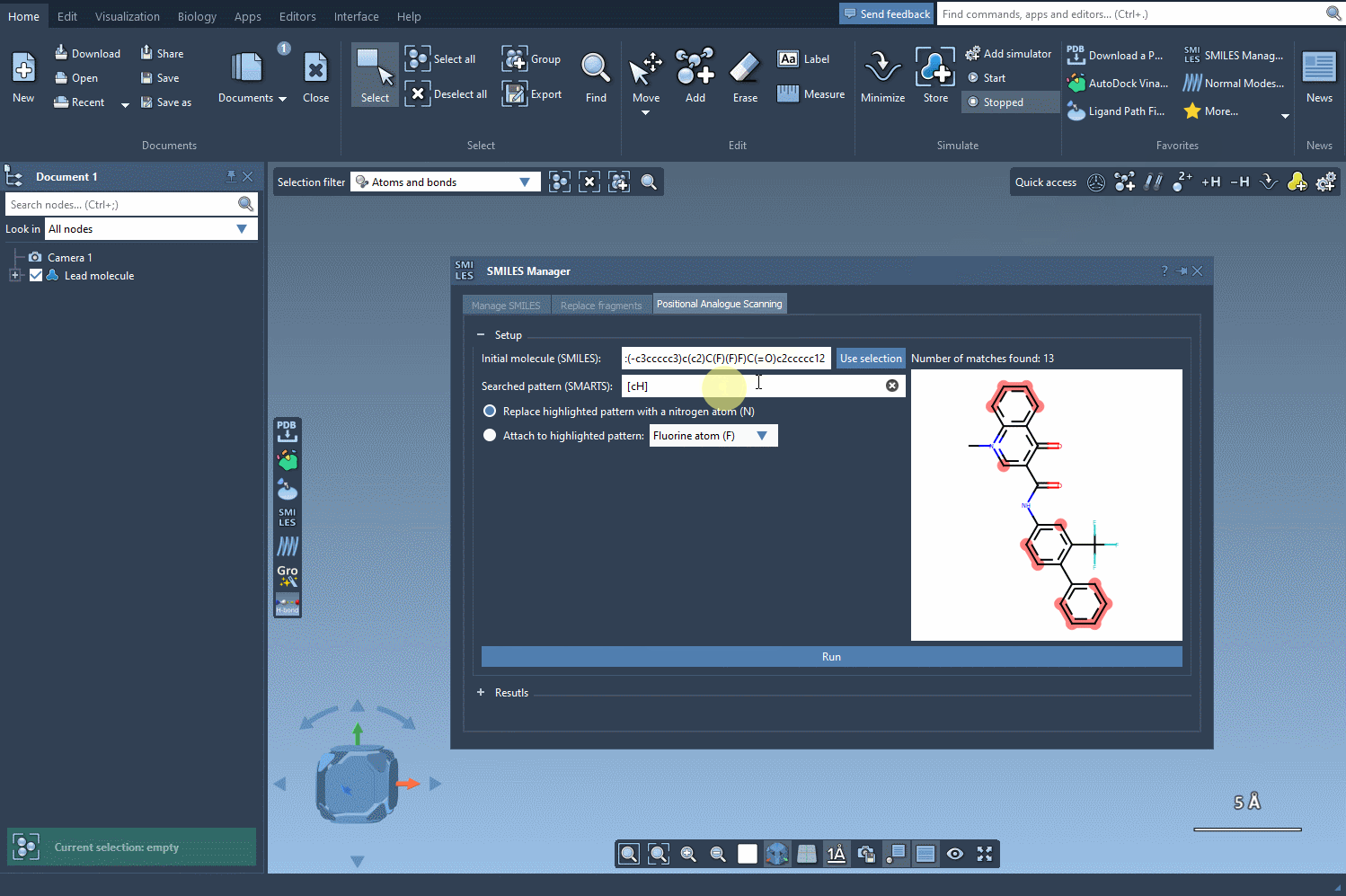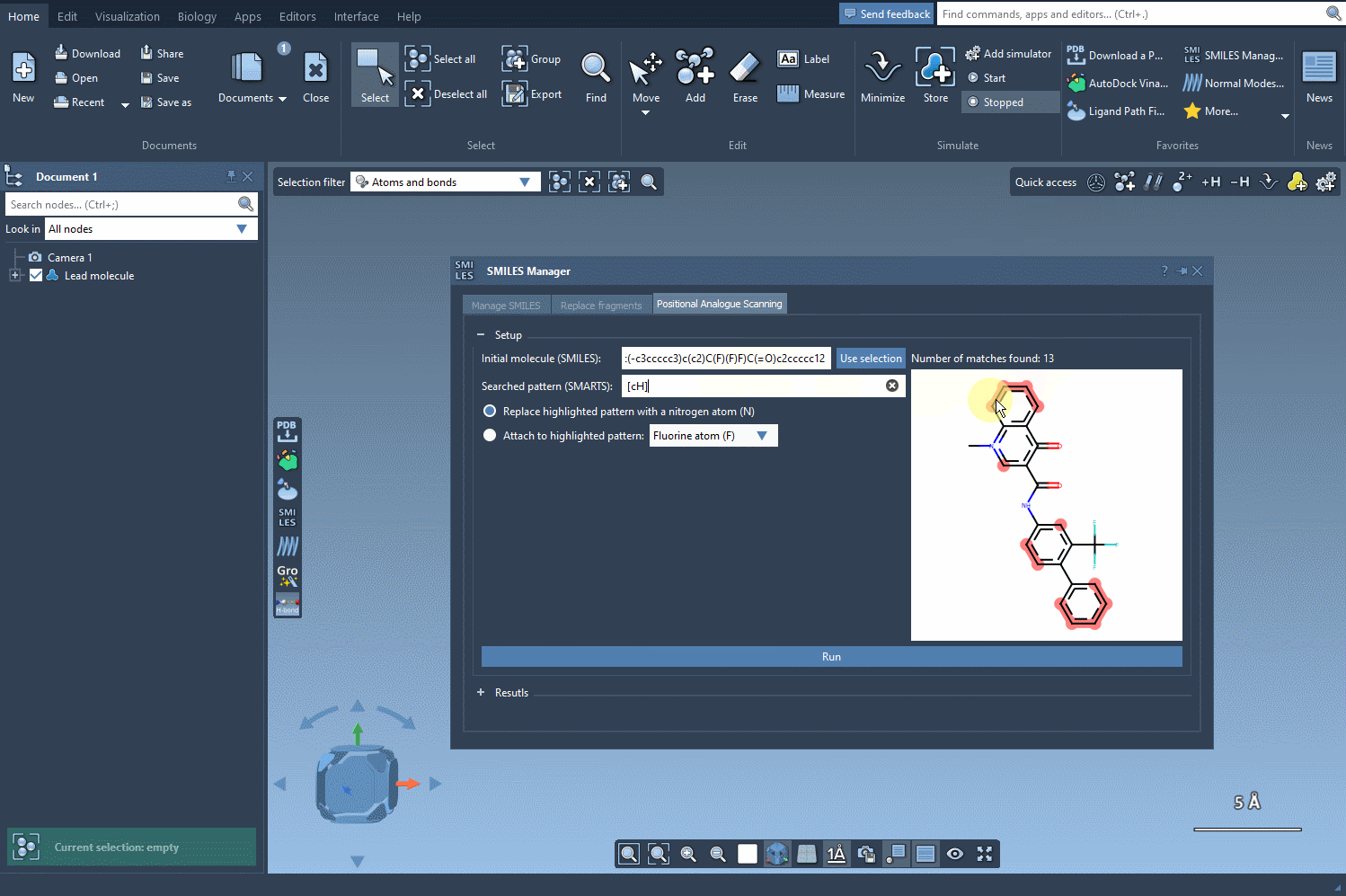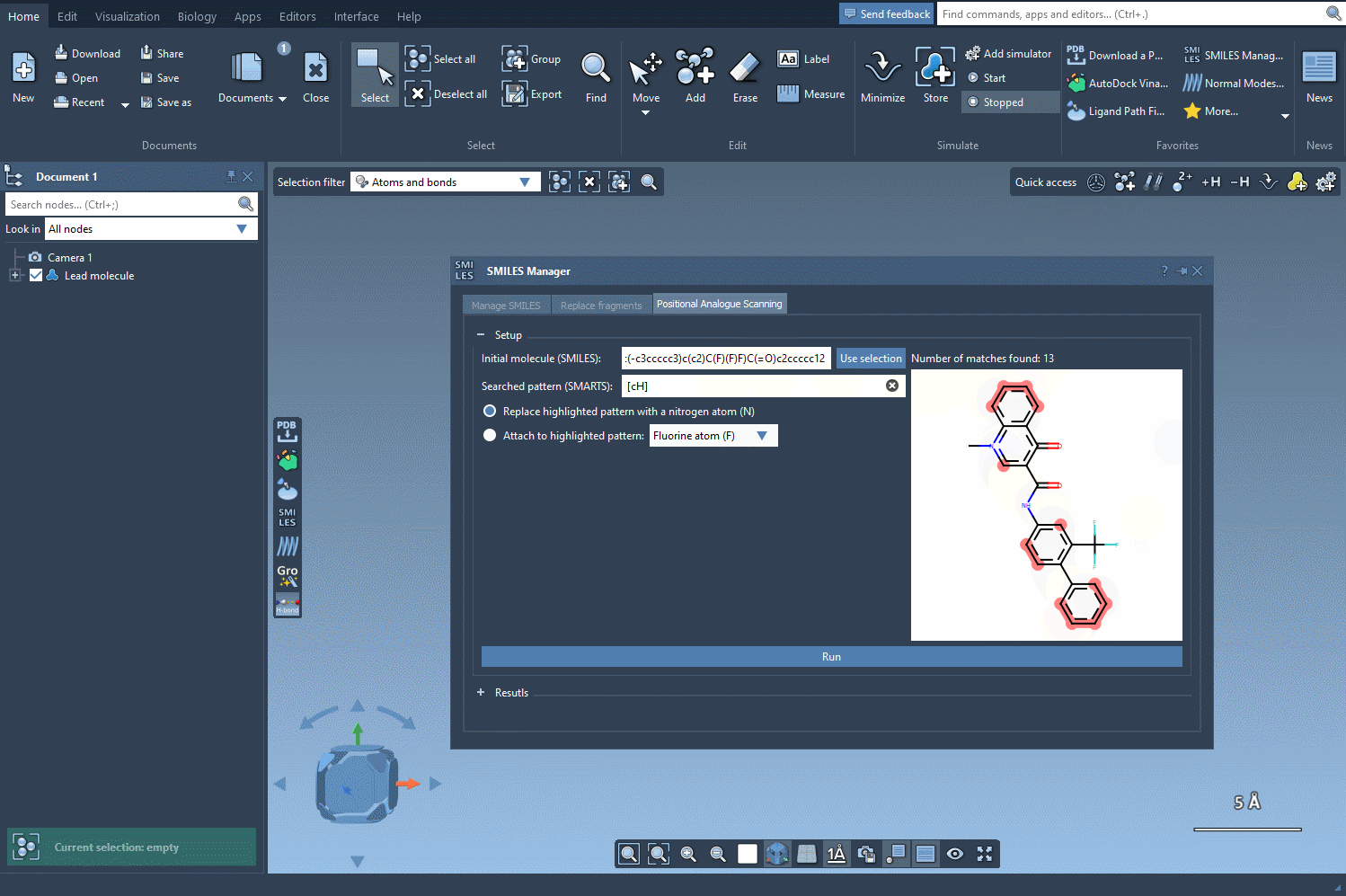
To begin, choose the initial molecule you wish to analyze. You can either input its SMILES code manually, or select it directly in SAMSON and press the Use selection button. This ensures a smooth and flexible start to the process. Check out the image below to see how it’s done:
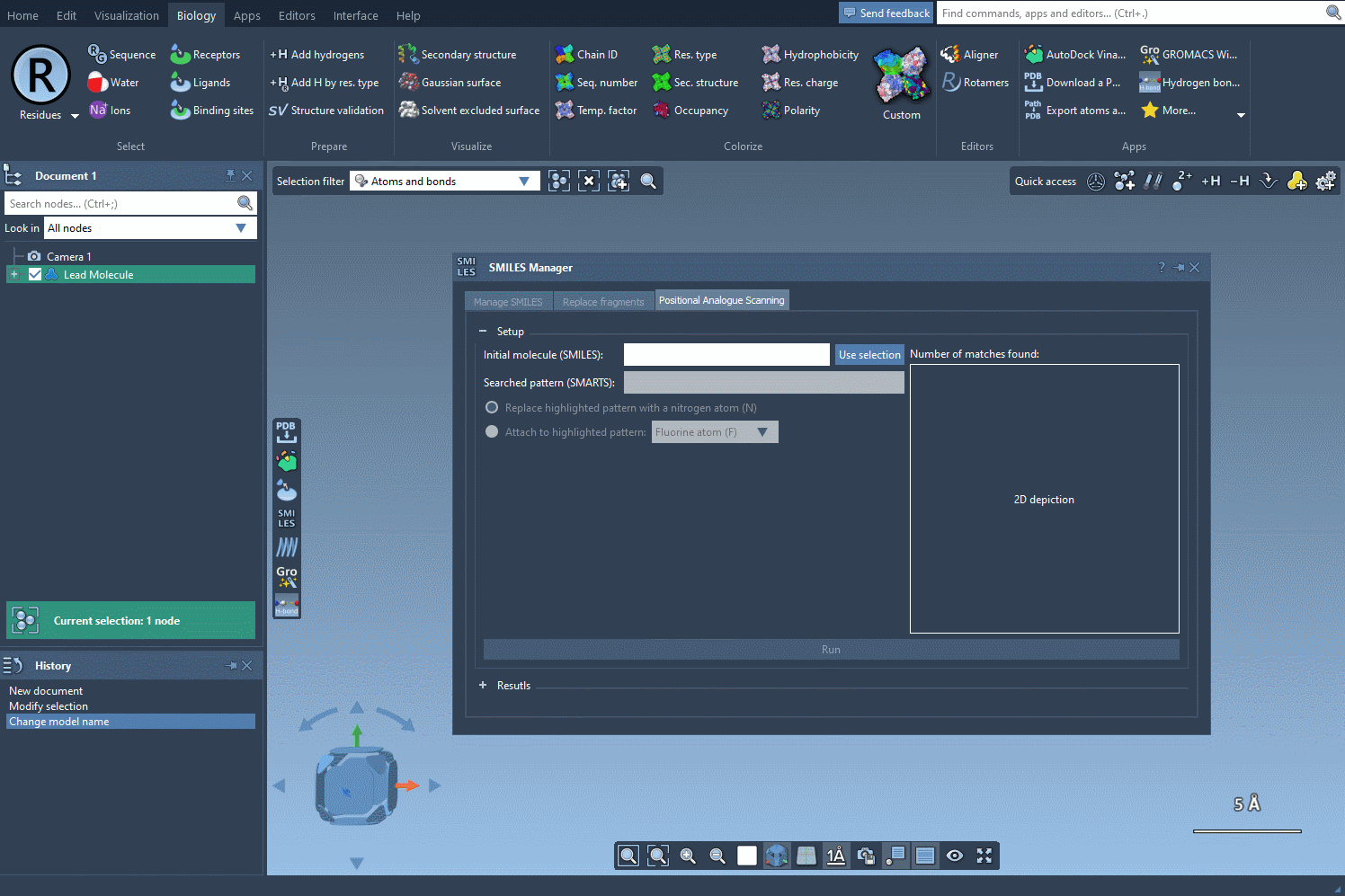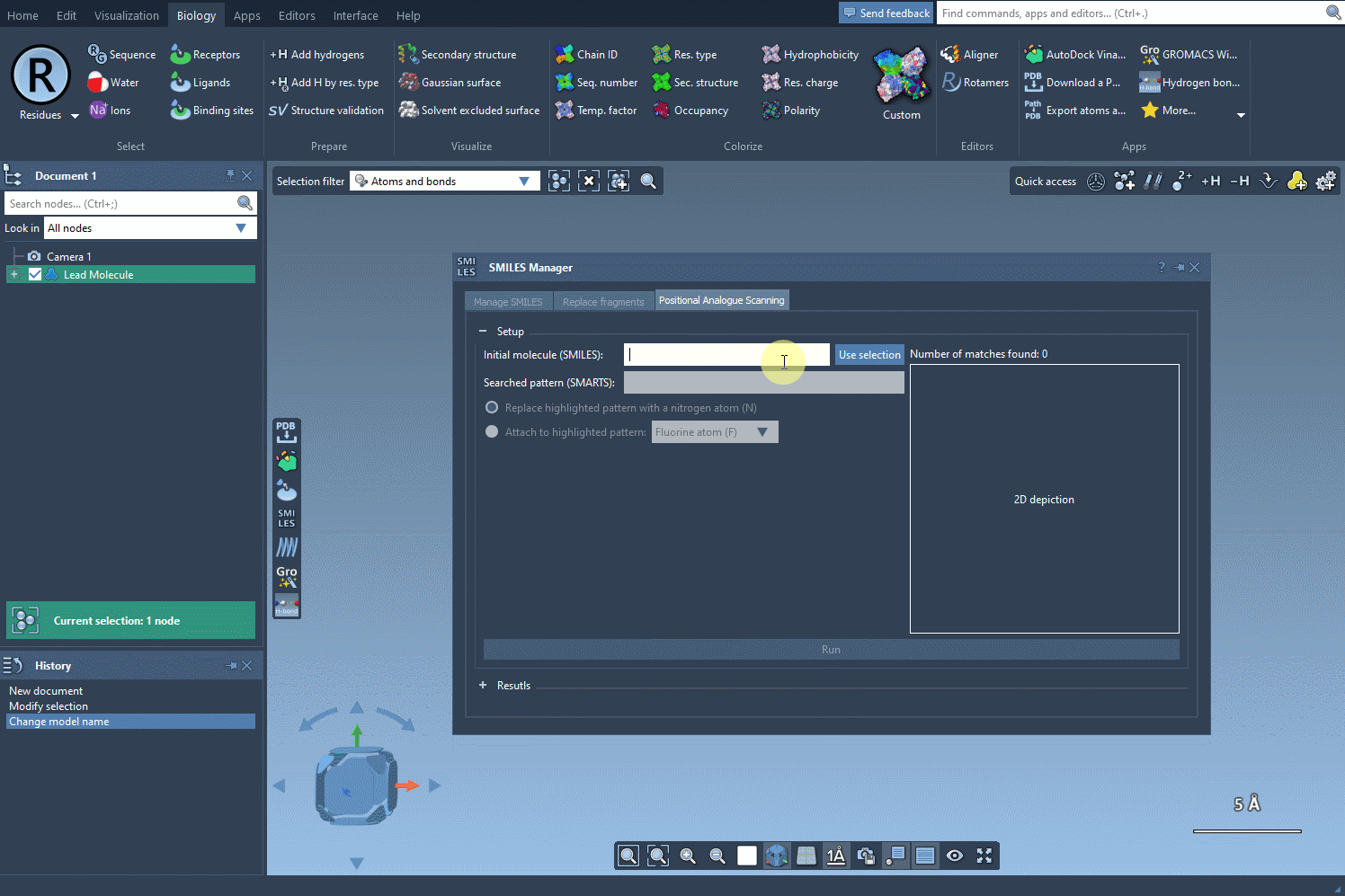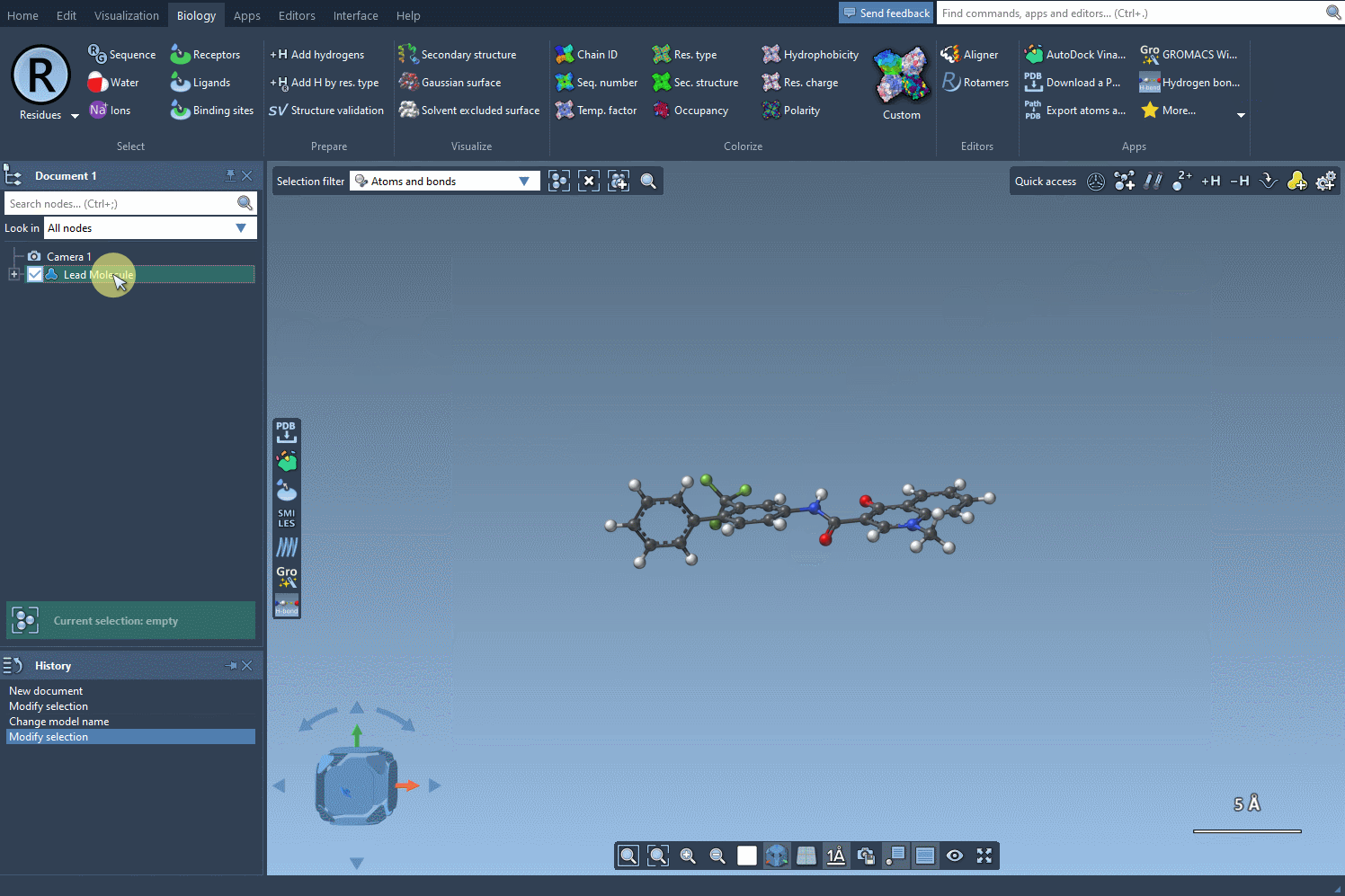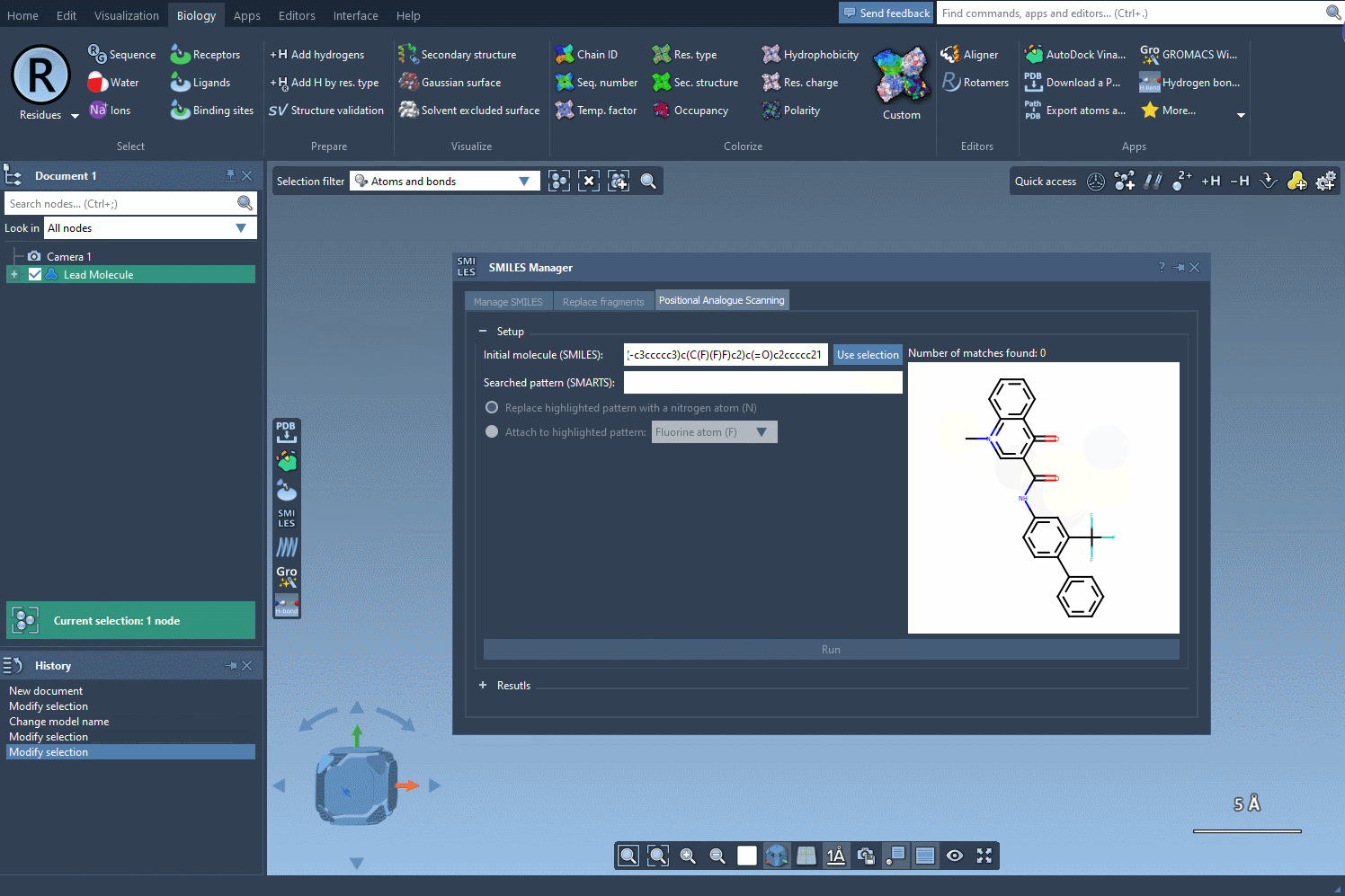
Step 2: Identifying the Target Substructure
Next, define the substructure pattern you want to modify using SMARTS codes. For instance, the aromatic carbon pattern, [cH], can be used to identify substitution points within a molecule. These identified patterns are highlighted directly in the molecule, helping you visually verify your choice. See how this step works:
Step 3: Replacing or Attaching Groups
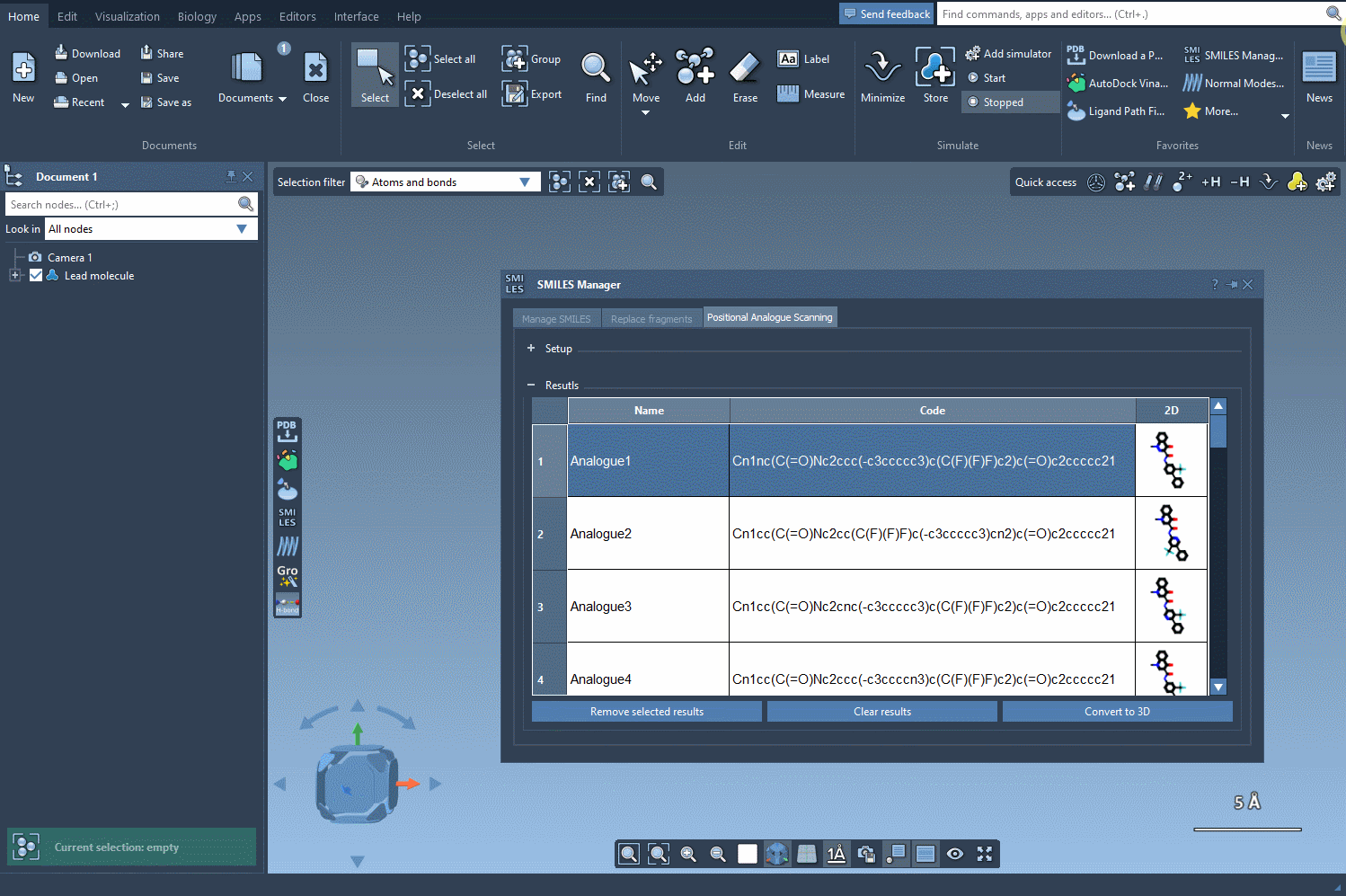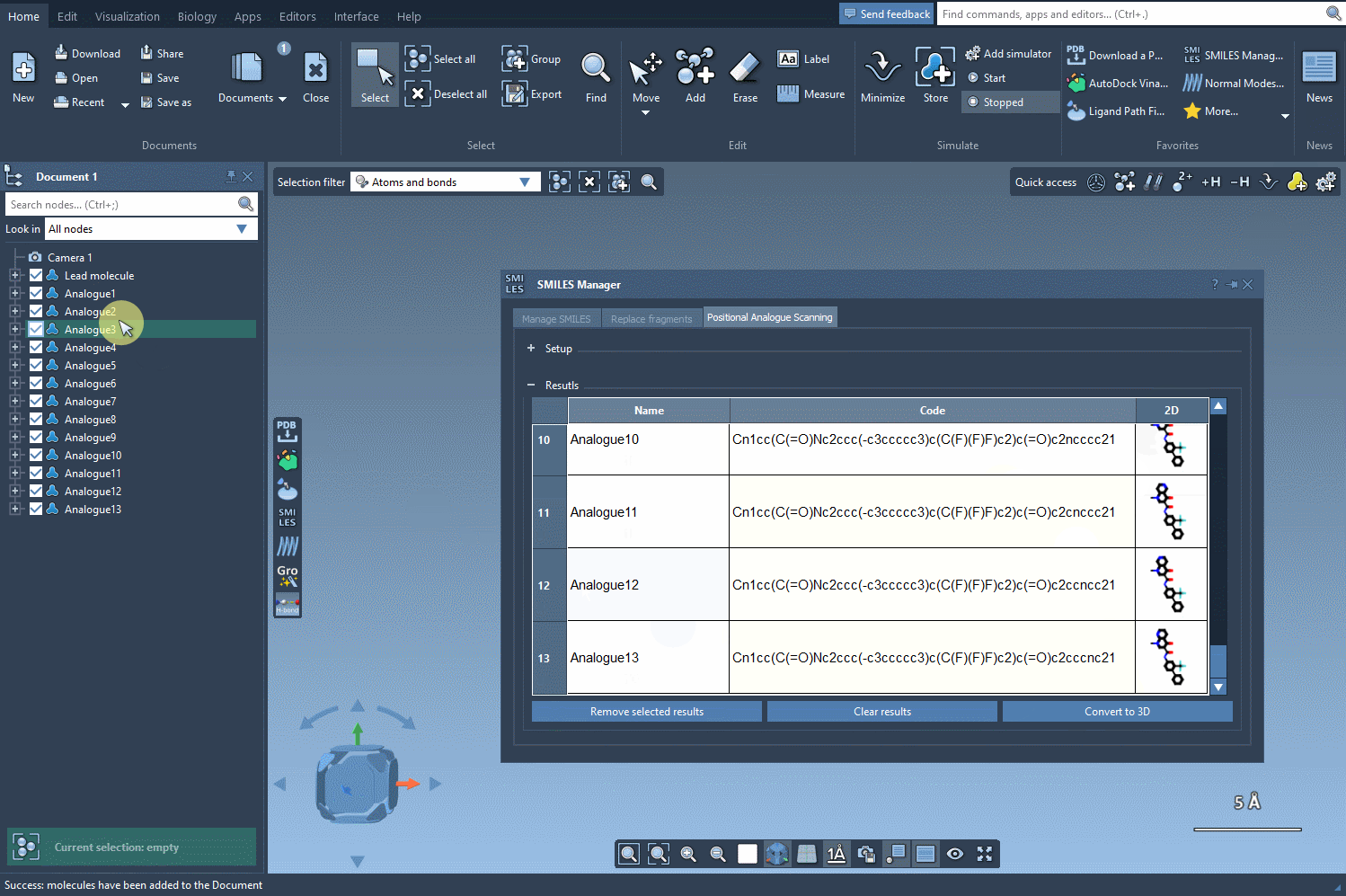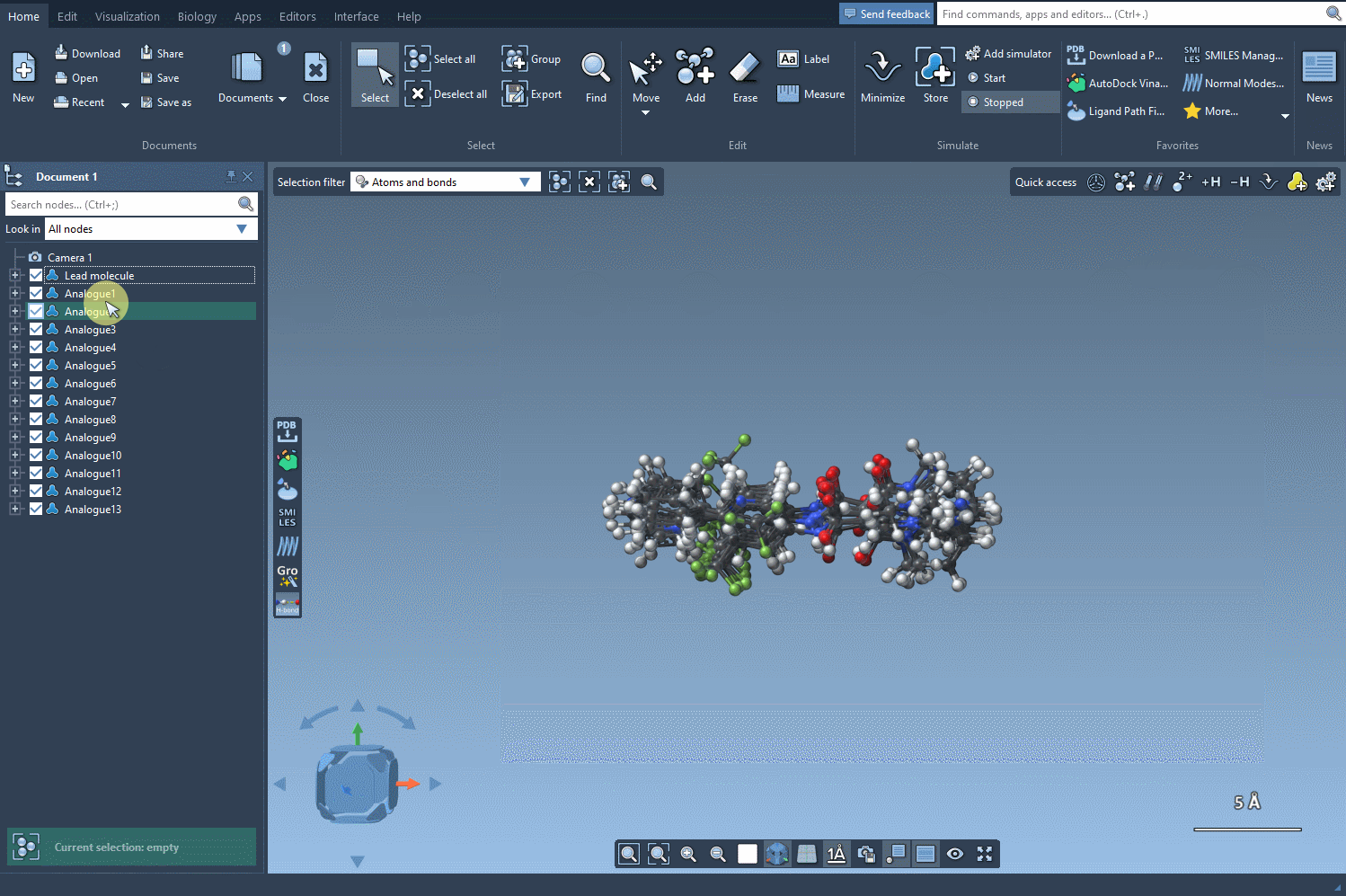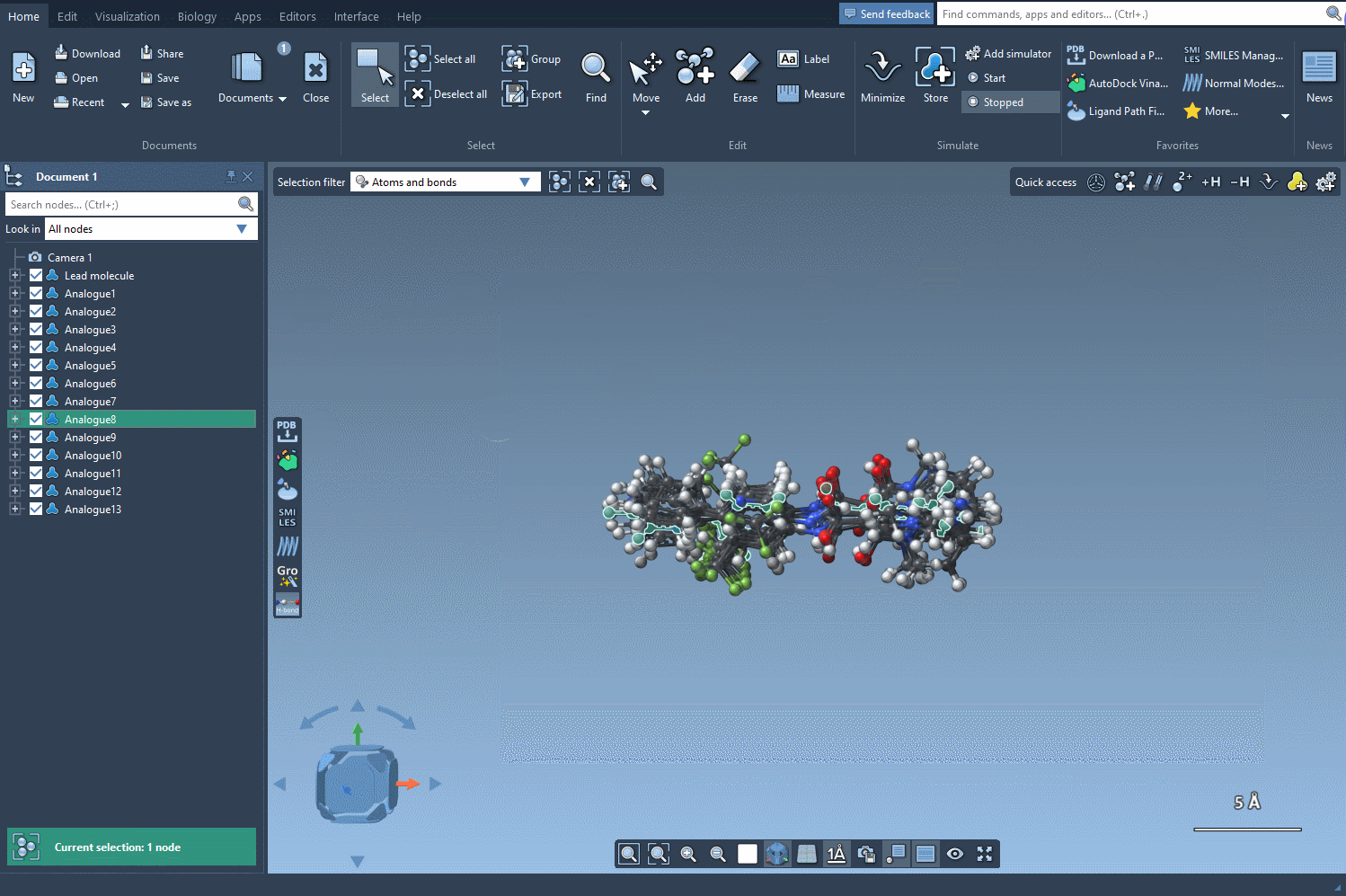
Once the substructure is identified, decide whether you want to replace it or attach a new group. The SMILES Manager allows substitutions such as replacing your substructure with a nitrogen atom (N) or attaching groups like a fluorine atom (F) or a methyl group (CH3). Clicking Run generates systematic analogues almost instantly, providing both their SMILES codes and 2D depictions. The generation process is illustrated below:
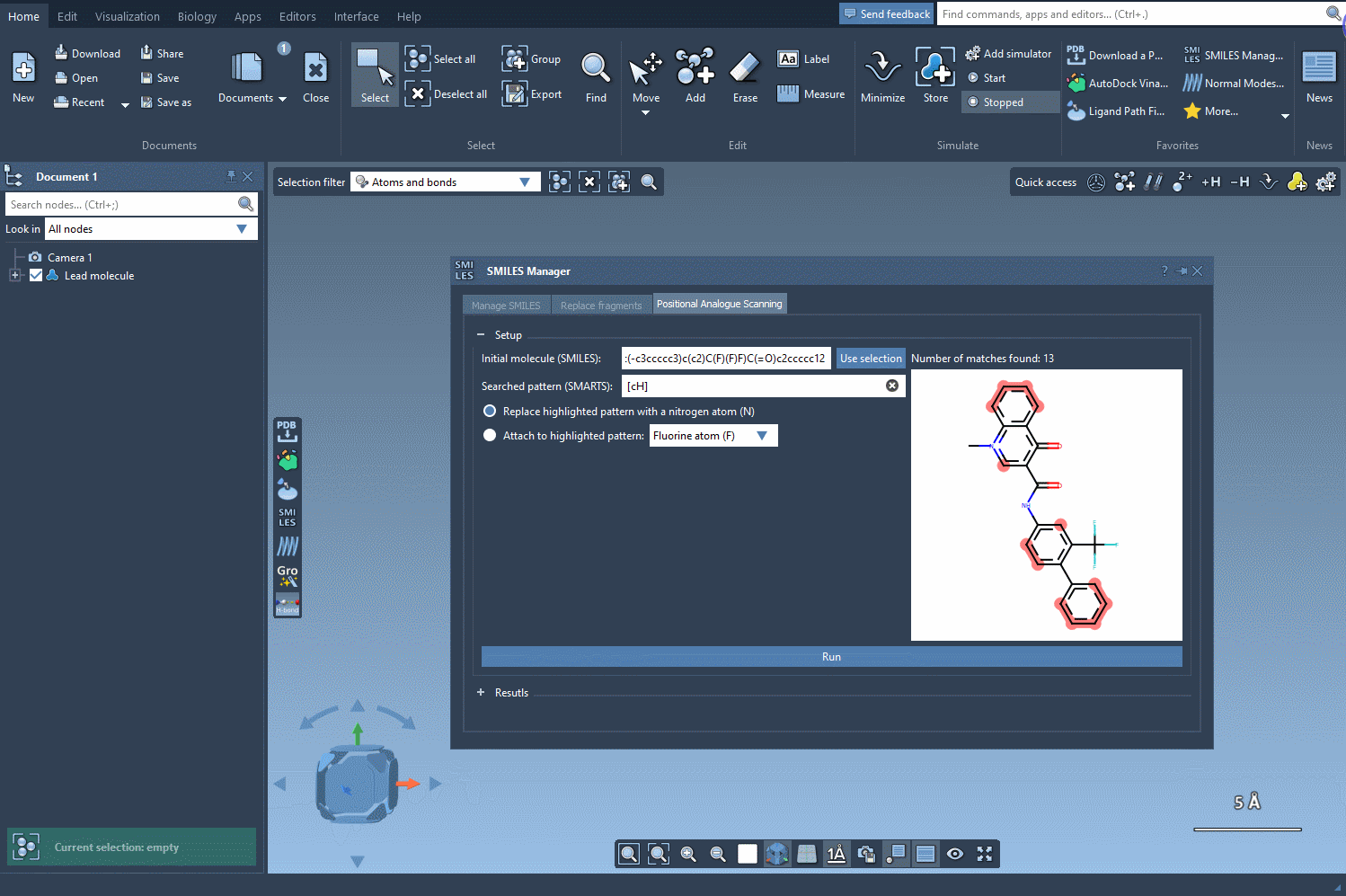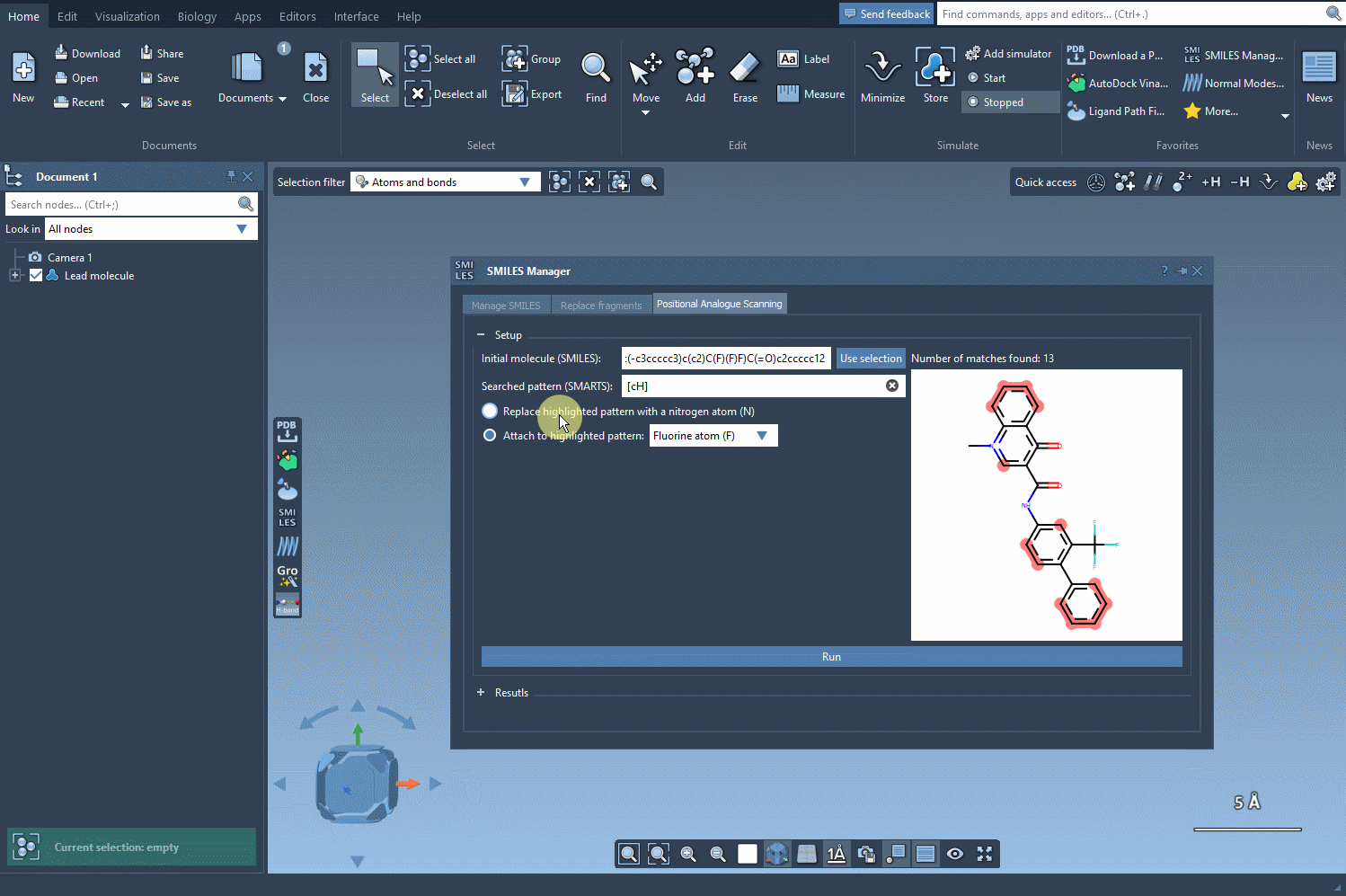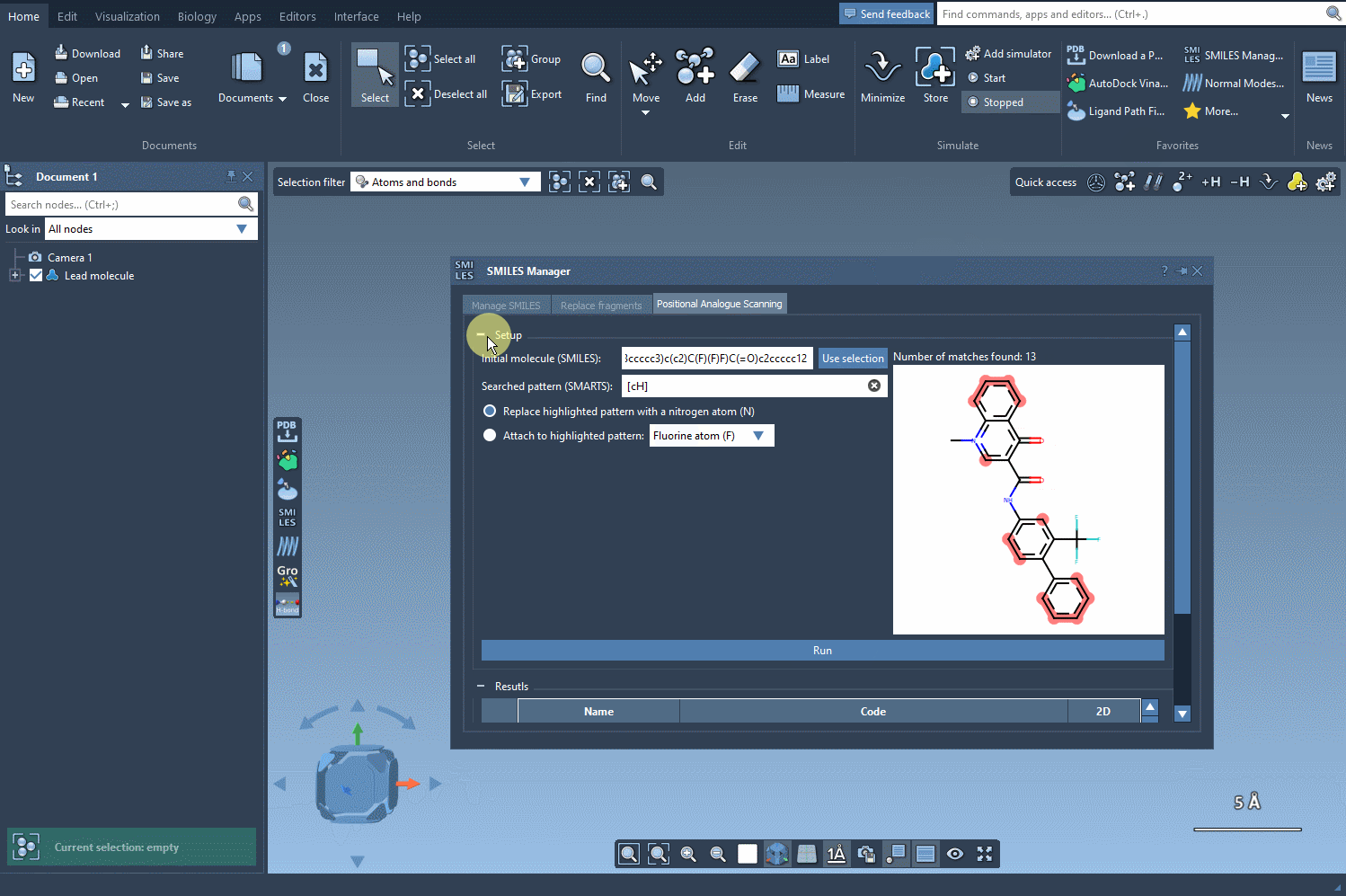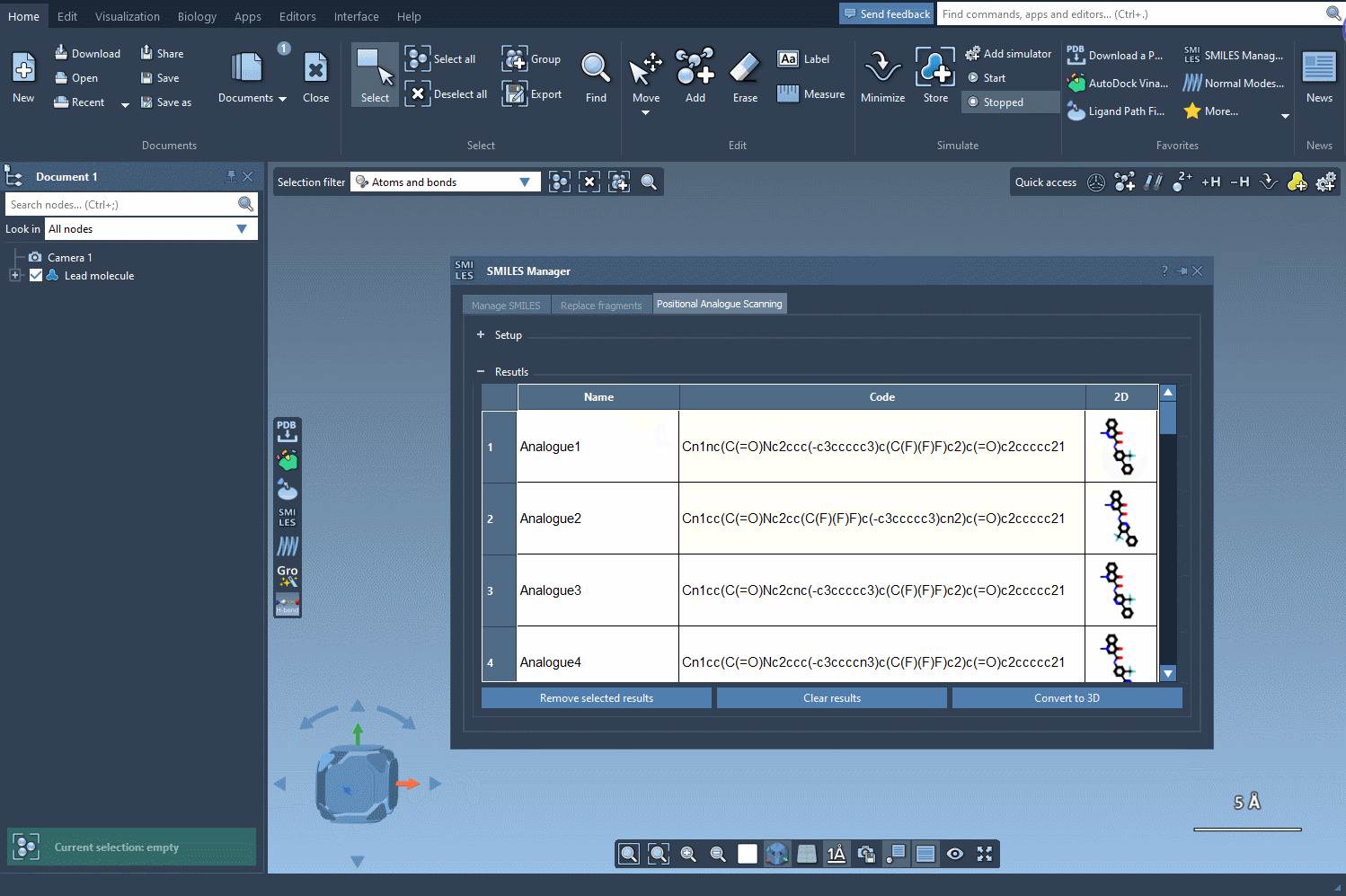
Step 4: Exploring and Refining Results
The generated analogues are displayed in a results table, where you can:
- Edit the name or SMILES code of any analogue.
- View larger 2D depictions of molecules by double-clicking on their images.
- Generate 3D structures for specific analogues for further analysis.
- Filter or remove individual or all analogues with simple actions.
Step 5: Generating 3D Structures
Once you identify promising analogues, convert them into 3D structures by pressing the Convert to 3D button. These 3D representations are immediately ready for docking studies, making this workflow an excellent choice for integrative molecular design. The following image shows the process:
Why Use this Workflow?
Positional analogue scanning in the SMILES Manager simplifies what could otherwise be a cumbersome process. From systematic substitution to quick visualization and compatibility with docking workflows, it offers a comprehensive approach to evaluating substitutions in less time. Whether you’re an expert or a newcomer to molecular modeling, this feature offers practical value for your research and development tasks.
For more information and step-by-step guidance, visit the official documentation page.
SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON today at https://www.samson-connect.net.





