Protein alignment is a critical step in molecular modeling, whether you’re analyzing evolutionary relationships, studying structural similarities, or preparing homology models. However, aligning sequences and structures can often feel tedious, especially without the right tools. This is where the Protein Aligner in SAMSON comes in, providing an efficient, all-in-one workflow for protein sequence and structure alignment.
Why Protein Alignment Matters
Protein alignment allows you to:
- Pinpoint conserved residues involved in key functional or binding roles.
- Compare conformations across species or variants.
- Create accurate homology models for downstream design tasks.
These applications are particularly valuable to molecular modelers focused on drug design, bioinformatics, or evolutionary studies. Yet, even small alignment errors can ripple into larger issues later in the workflow, making accurate tools like the Protein Aligner an important asset.
What Makes SAMSON’s Protein Aligner Stand Out?
The Protein Aligner extension simplifies the often-complex process of protein sequence and structure comparisons. Integrating seamlessly into SAMSON’s interface, it offers features such as:
- Alignment workflows for both sequences and 3D structures.
- Visualization of conserved residues with options to highlight properties like polarity and similarity.
- Region-specific alignment to focus on specific structural motifs of interest.
These features make it a versatile and user-friendly tool for aligning your target proteins quickly and with precision.
How to Launch the Protein Aligner
Starting with the Protein Aligner is as simple as following these steps:
- Prepare your proteins. Use Home > Prepare to strip out solvent, ligands, and alternate locations for a cleaner structure.
- Load your proteins. Under Home > Fetch, you can input PDB codes (e.g.,
1DLWand1RTX) and fetch them directly. - Open the Protein Aligner via Home > Align.
The interface automatically guides you to align sequences, superimpose structures, and inspect conserved residues.
Region-Specific Alignment: When Precision Matters
Sometimes, focusing on a local structural region rather than the entire protein is more relevant. For example, aligning similar alpha-helices separately can provide a clearer comparison. The Protein Aligner enables region-specific alignment by letting you:
- Select specific residues within both protein sequences.
- Run alignment based only on these selected parts, overriding global alignment results.
With just a few clicks, you can superimpose only the regions that are most crucial for your study, as shown below:

This feature is particularly useful when studying active sites, binding pockets, or structurally significant motifs.
Superimpose Structures in a Snap
Whole-protein superposition is another highlight of the Protein Aligner. Simply click Align to this on a model’s row to make it the reference, and the button for the other model will display the root-mean-square deviation (RMSD). Your structures are then instantly superimposed:
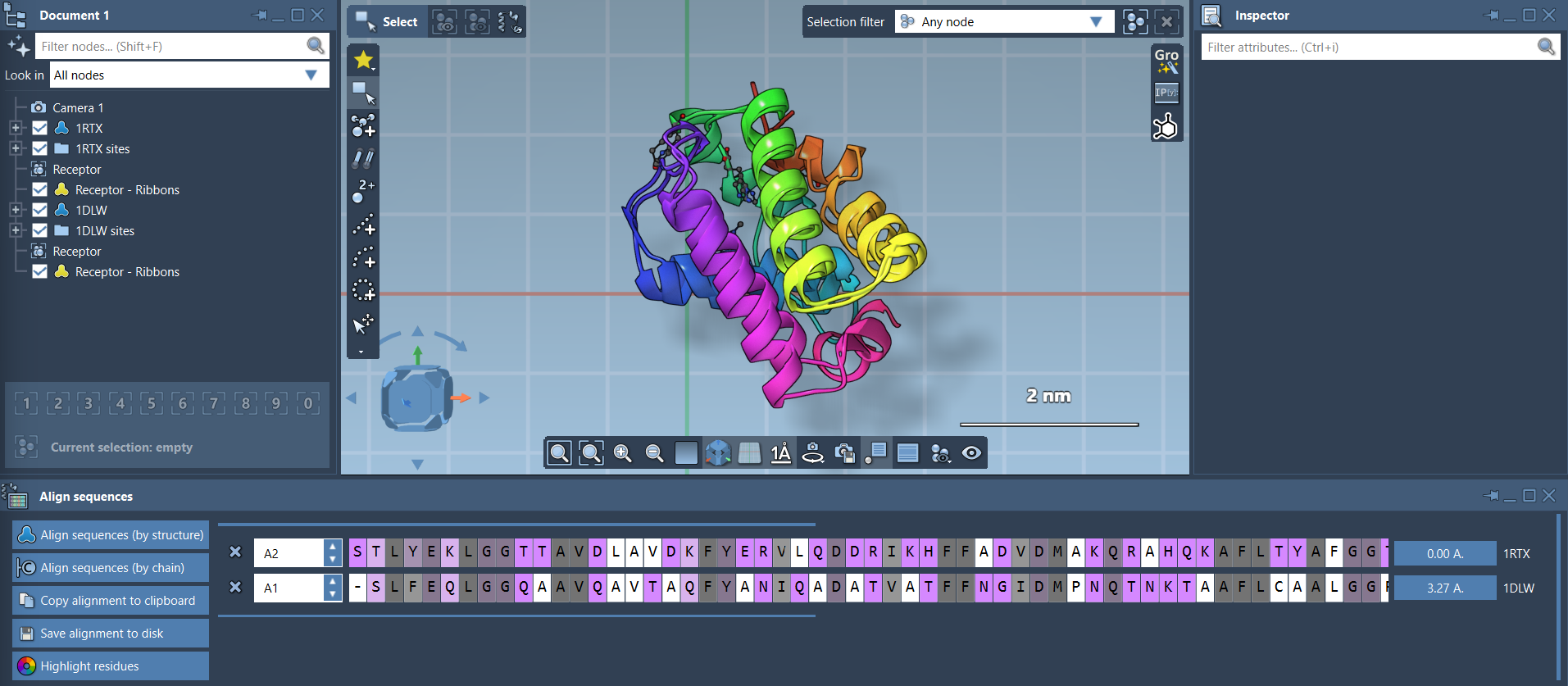
Adding secondary structure visuals (via Visualization > Visual model > Ribbons) makes interpretation easier, especially when dealing with complex assemblies.
Next Steps
Once you’ve aligned your proteins, you can:
- Export the results as inputs for homology modeling or molecular docking workflows.
- Highlight conserved residues within binding pockets for ligand design experiments.
- Iteratively align additional chains or homologous proteins for broader comparisons.
To dive deeper into how the Protein Aligner can enhance your projects, check out the complete documentation at this link.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Get started today!





