Molecular modelers often face the challenge of identifying specific substructures within large molecules or datasets. This task can become tedious and time-consuming if the right tools and features are not at hand. Thankfully, the Substructure Search functionality in the SMILES Manager extension of SAMSON makes this process much more efficient and user-friendly.
What is Substructure Search?
The substructure search feature allows users to filter molecules based on specific patterns or features using their SMILES representations. With this functionality, you can efficiently identify molecules that meet complex criteria, such as the presence of certain atoms or functional groups spatially separated in a specific manner. For molecular researchers, this can mean dramatically streamlined workflows and faster data analysis.
How It Works
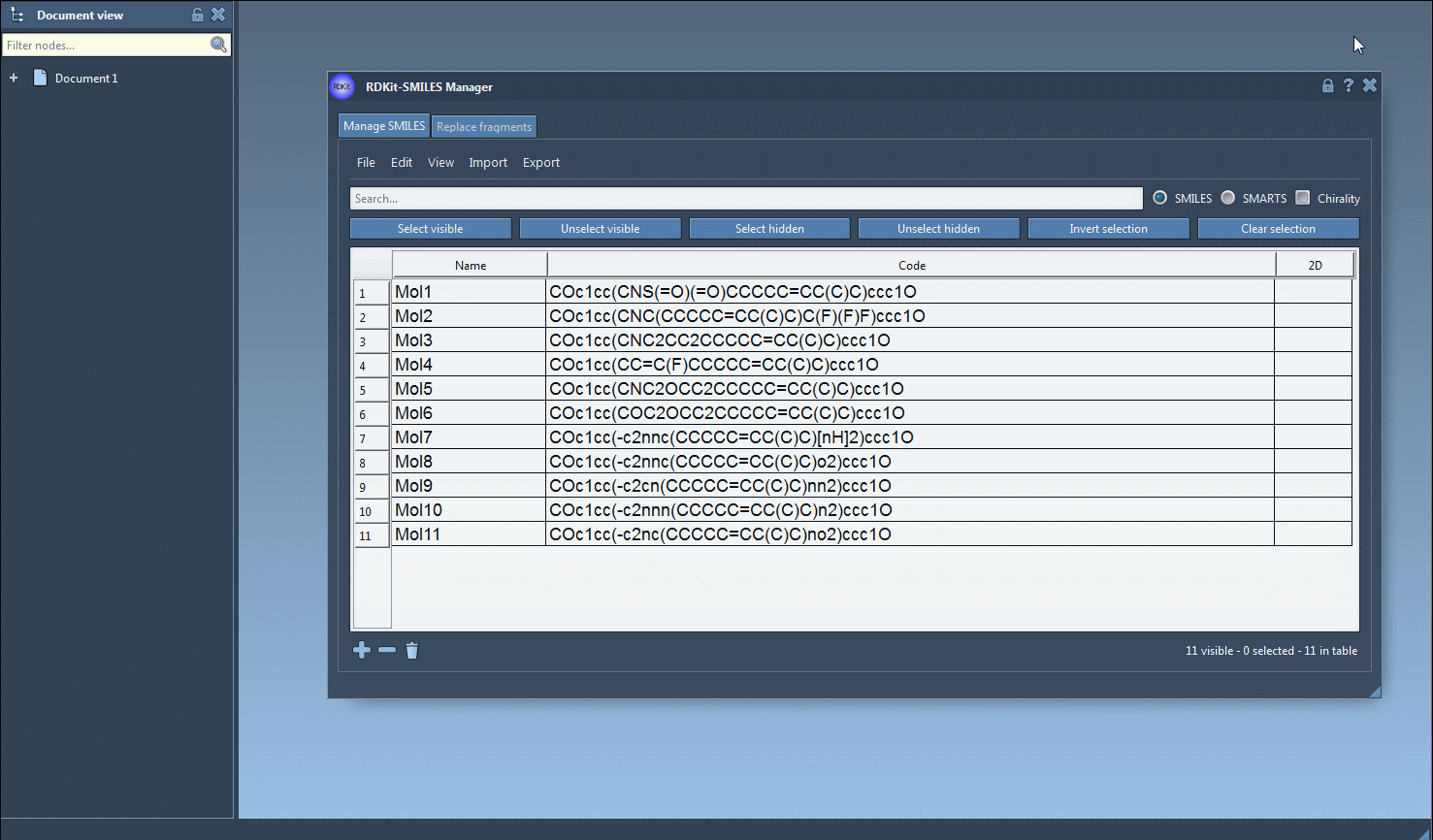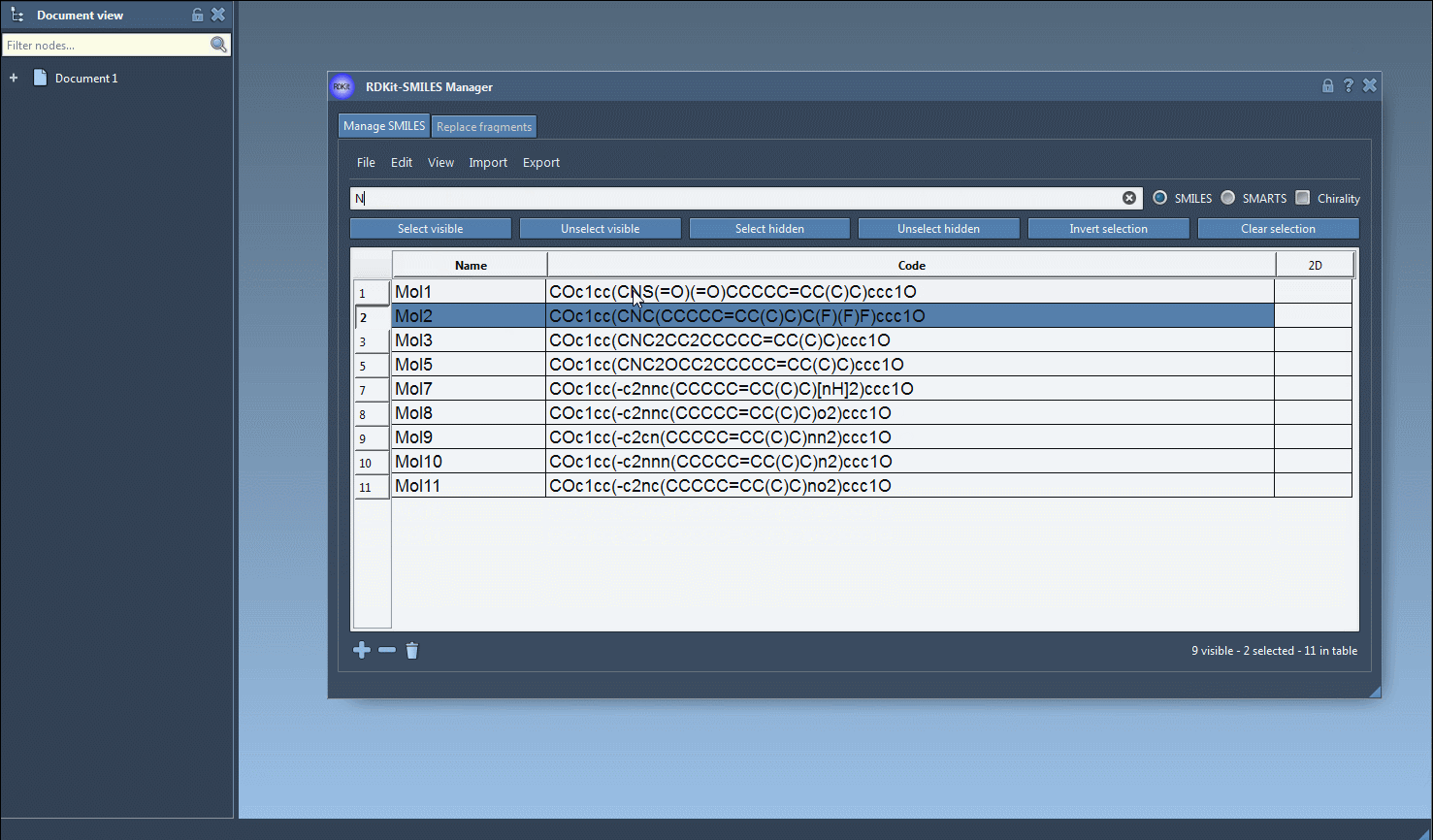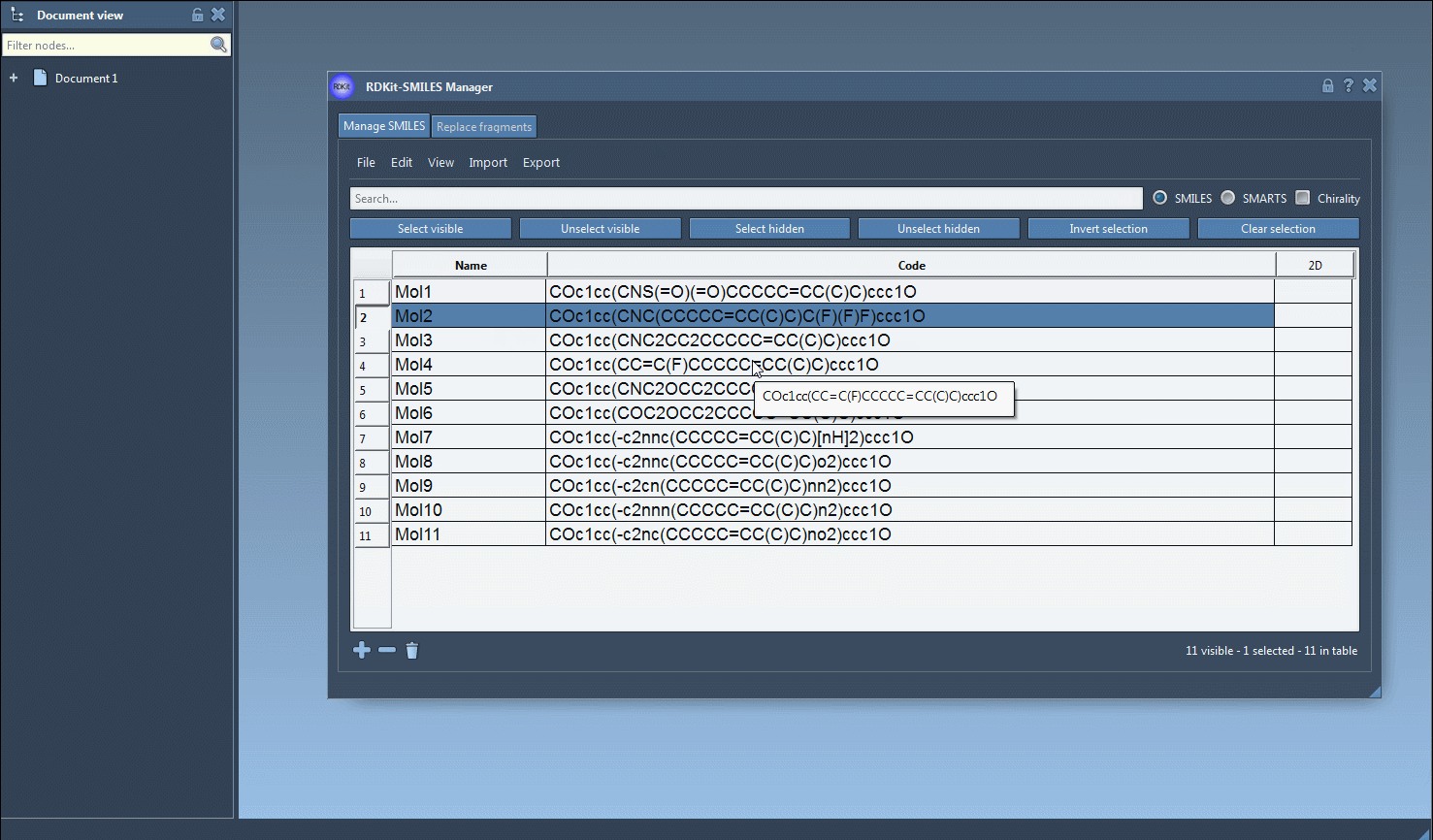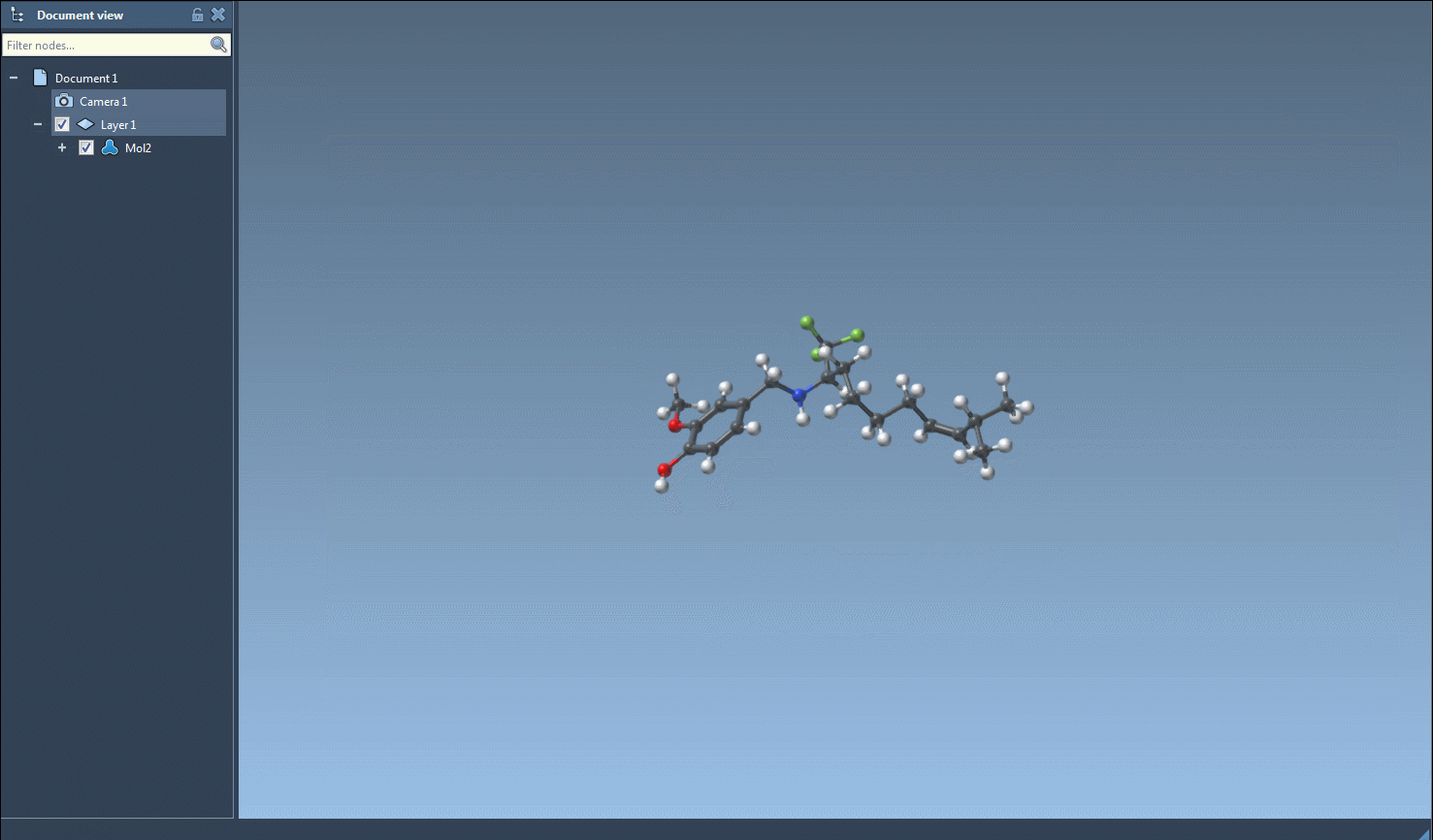
The substructure search tool is intuitive. Here’s an example of how you can perform a search for molecules containing both a fluorine (F) atom and a nitrogen (N) atom:
- Using the Search bar in the SMILES Manager interface, input your first search pattern, such as
F, to identify molecules containing a fluorine atom. Then press the Select visible button to select these results. - Clear the search bar, ensuring that your prior selections remain intact, and input a second search pattern such as
Nto locate nitrogen atoms. At this point, you’ll see overlapping results from both searches. - Click on Unselect hidden to refine your selection, keeping only the molecules that meet both criteria (in this case, containing both fluorine and nitrogen atoms).
Customization Options
This feature can handle increasingly sophisticated searches by combining multiple patterns. By default, it uses RDKit’s substructure search rules, which ignore stereochemistry. Those who wish to enable chirality considerations can adjust this setting.
Why It’s Useful
For researchers working with large molecular datasets, this targeted approach offers multiple advantages:
- Efficiency: Quickly narrow down datasets based on specific structural features.
- Accuracy: Perform advanced queries that precisely meet your requirements.
- Automation: Combine searches for more complex analyses without manually highlighting molecules.
In terms of practical applications, substructure searches can be used to identify molecules with desired functional groups for further experimental or computational studies, save results for future reference, and even export them for visualization or 3D structure generation.
Tips for Success
As a best practice, familiarize yourself with the SMILES syntax to make the most out of the substructure search feature. Additionally, ensure that your molecular dataset is clean and formatted correctly prior to importing it into the SMILES Manager.
Start Searching Smarter
The substructure search functionality in the SMILES Manager is designed to save time and improve your workflow. Whether you are analyzing small libraries or extensive datasets, this tool offers immense value for anyone working with molecular data.
Learn more about this and other functionalities of the SMILES Manager to enhance your molecular modeling projects.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON here.





