Streamlining Protein Superposition: A Guide for Molecular Modelers
If you’re a molecular modeler, chances are you’ve encountered the challenge of aligning proteins—both their sequences and structures. Whether you’re building homology models, comparing mutated proteins, or mapping conserved residues, protein alignment is at the heart of many computational workflows. But achieving precise superposition without tedious manual adjustments can be a pain.
Fortunately, SAMSON’s Protein Aligner provides a seamless tool for aligning proteins at both the sequence and structural levels, saving you time and ensuring accuracy. In this post, we’ll focus on aligning structures—a frequent but vital task for many molecular modelers looking to compare entire proteins or specific regions.
Why Structural Alignment Matters
Structural alignment allows you to find critical spatial relationships between proteins by superimposing one structure onto another. This is indispensable for:
- Understanding conservation of binding sites and functional residues.
- Analyzing conformational changes across species or mutants.
- Building accurate models for downstream molecular design workflows.
Let’s break down how you can leverage SAMSON to easily superimpose proteins.
Getting Started with Protein Alignment in SAMSON
Follow these steps to align two proteins, such as hemoglobins (1DLW and 1RTX) from different organisms:
- Fetch the proteins:
- Navigate to Home > Fetch within SAMSON.
- Enter the PDB IDs
1DLWand1RTX. - Click the Load button.

- Prepare the proteins: Clean up the proteins before alignment. Use Home > Prepare to remove water molecules, alternate locations, or ligands (when needed). You can also refer to SAMSON’s Protein Preparation guide.
- Launch the Protein Aligner: Click on Home > Align to open the Protein Aligner tool
 .
.
Performing Whole-Protein Superposition
For aligning entire proteins:
- Ensure no residues are pre-selected (click anywhere in the viewport to clear selections).
- Click Align to this on the row of the first model.
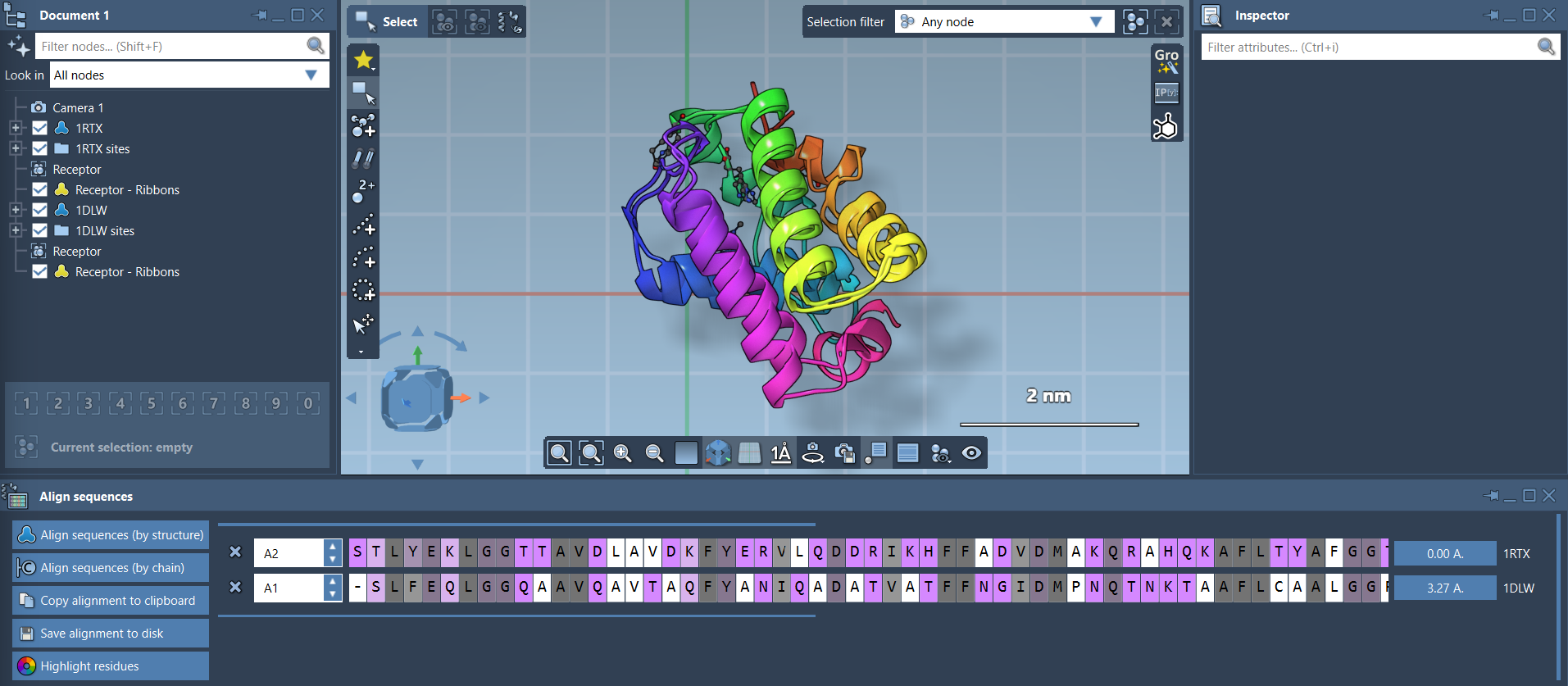
Once aligned, RMSD (Root Mean Square Deviation) is automatically calculated, as shown in the interface:
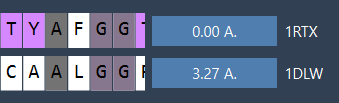

Your proteins are now superimposed! For visualization, SAMSON lets you toggle secondary structures, apply color differentiation for each protein, and adjust atomistic views:
Pro Tip
Enhance clarity by showing ribbons via Visualization > Visual model > Ribbons. To color aligned proteins differently, create separate ribbon models for each structure.
Region-Specific Structural Alignment
What if only part of the protein is relevant? SAMSON allows region-specific alignment, so you can focus on key sections like binding sites or conserved domains:
- Select residues of interest in both sequences using the Protein Aligner.
- Click the alignment button (e.g.,
0.0 Å) next to the selection to superimpose just those residues.
For example, in the image below, the alignment focuses on two pink alpha-helices from the N-terminal regions:

The final result highlights how precise this localized superposition can be:

Next Steps
After aligning your proteins:
- Export the alignment for homology modeling or structure-based analysis.
- Map conserved residues to identify potential ligand-binding hotspots.
- Repeat alignment for additional related structures or chains.
In just a few clicks, SAMSON simplifies complex alignment tasks, enabling molecular researchers to focus on their design challenges.
Learn more by exploring the official Protein Aligner documentation.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Download SAMSON for free at SAMSON Connect.





