Understanding molecular dynamics is essential for molecular modelers, especially when investigating ligand unbinding, protein conformational changes, or analyzing the behavior of molecular complexes. A common challenge for researchers lies in visually tracking the motion of specific groups of atoms or the center of mass during simulations. That’s where the Pathlines feature of SAMSON becomes a valuable asset.
What Makes Pathlines Useful?
The Pathlines visual model in SAMSON enables researchers to visualize the trajectory of the center of mass (COM) for selected atoms along predefined paths. This can help in understanding various critical phenomena in molecular systems, such as:
- Tracking the motion of ligands or residues over time.
- Analyzing displacement patterns in complex macromolecular assemblies.
- Studying processes such as unbinding, diffusion, or large-scale domain movements.
How to Get Started with Pathlines
Here’s a step-by-step guide to set up and use Pathlines within SAMSON, catering to both novice and experienced users.
1. Load a Sample System
If you’re new to this feature, SAMSON provides an excellent sample system to experiment with:
- Go to Home > Download in SAMSON.
- Paste the link below into the download bar and hit Download:
https://www.samson-connect.net/documents/046f1acd-c799-40f6-8185-cb4847eff795

This document contains the model of Lactose permease (1PV7) and its ligand, Thiodigalactosid (TDG), alongside paths demonstrating ligand unbinding. You can explore these paths in the Document view.
2. Select Atoms and Paths
To create Pathlines, you need to select atoms and the corresponding paths:
- Open the Document view and choose a group of atoms for which you want to visualize the COM’s motion.
- Select one or more paths (you can hold Ctrl / Cmd to multi-select).
If nothing is selected, Pathlines will use the entire system and all paths by default.

3. Create a Pathline Visual Model
With your atoms and paths ready, you can visualize the COM trajectory:
- Go to Visualization > Visual model > More… or use the shortcut (Ctrl/Cmd + Shift + V).
- In the dialog, choose Pathline of the center of mass and click OK.
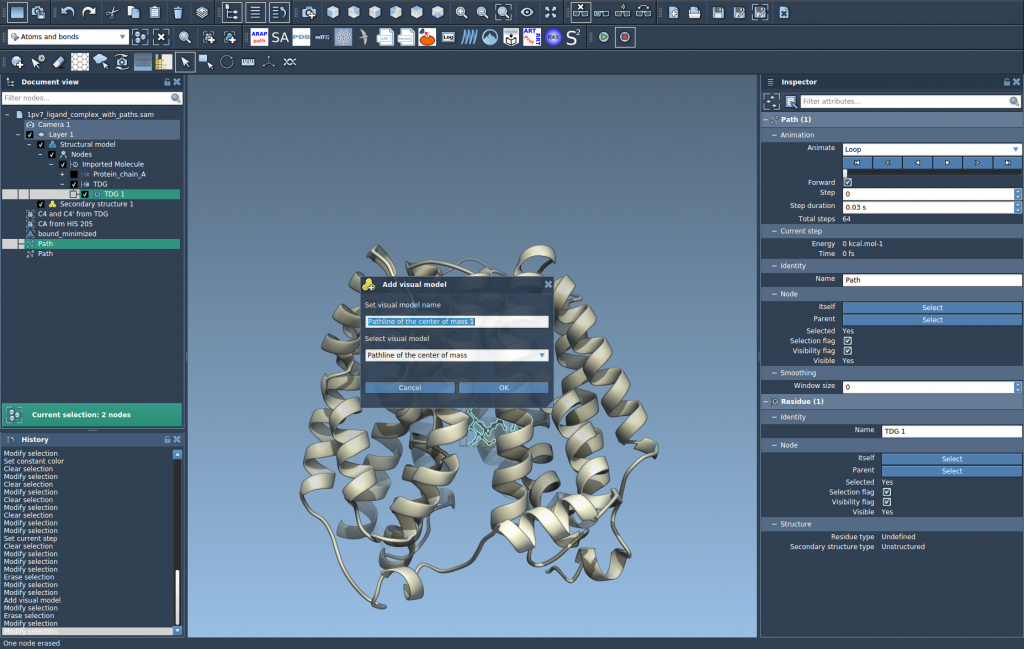
Once created, the Pathline will display the movement of the selected atoms’ COM along the defined paths.
4. Customize and Explore
SAMSON allows users to further refine and analyze their Pathline visual models:
- Double-click a path in the Document view to toggle its visibility.
- Right-click to access additional options in the context menu under Path > ….
- Open the Inspector (Ctrl/Cmd + 2) to tweak properties such as pathline thickness and color.
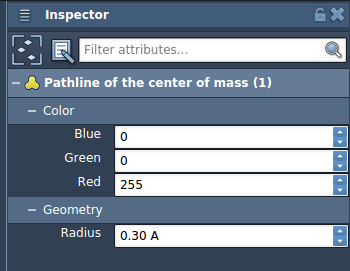
Applications of Pathlines
The Pathlines feature is ideal for investigating diverse molecular phenomena, including:
- Visualizing ligand unbinding/rebinding routes.
- Analyzing large-scale collective domain movements in protein complexes.
- Tracking COM motion in reaction coordinate workflows.
By incorporating Pathlines into your molecular modeling workflow, you can obtain a deeper understanding of dynamic processes and visualize intricate atomic motions with ease.
For a more detailed explanation of Pathlines, visit the official SAMSON documentation page: https://documentation.samson-connect.net/tutorials/pathlines/pathlines/.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON at https://www.samson-connect.net.





