For molecular modelers, one common challenge is efficiently building, editing, and simulating molecular structures that may undergo significant topological changes. SAMSON’s Interactive Modeling Universal Force Field (IM-UFF) offers a solution tailored to this need, seamlessly blending interactive molecular modeling with physically-based inter-atomic forces. If you’re unfamiliar with IM-UFF, here’s how it can transform your workflow and save you headaches during molecular modeling.
Why IM-UFF Matters
Modeling dynamic molecular systems can be complex. Standard force fields often fall short when it comes to addressing real-time changes like the creation or breaking of covalent bonds, variations in bond orders, or modifications in atom classifications. IM-UFF bridges this gap by extending the Universal Force Field (UFF) for interactive modeling. Built into SAMSON, it allows you to visually and interactively simulate molecular systems, adjusting their topology as needed to reflect the physical realities of atomic interactions.
IM-UFF is particularly valuable for tasks like:
- Adding or deleting atoms directly within molecular models while maintaining accurate simulations.
- Adjusting bond orders dynamically, based on atomic positions.
- Facilitating smooth transitions between different molecular topologies.
How to Set Up IM-UFF
Getting started with IM-UFF in SAMSON is straightforward. Follow these steps:
- Open a document with a molecular system you’d like to simulate.
- Add a simulator by navigating to Edit > Simulate > Add Simulator or using the keyboard shortcut
Ctrl + Shift + M(orCmd + Shift + Mfor macOS). - From the list of interaction models, select Interactive Modeling Universal Force Field.
- Choose your preferred state updater, such as FIRE (Fast Inertial Relaxation Engine).
- Click the OK button to finalize your setup.
Unlike the standard UFF, IM-UFF does not require a setup window, making the process faster. Once enabled, you’ll see the IM-UFF parameter window, which provides additional options for interactive modeling, such as:
- Static topology (UFF only): Switch between standard UFF and IM-UFF with this setting. Note that the bond energy reference is different in IM-UFF—it is zero at infinite atomic distance rather than equilibrium distance, ensuring consistency with interactive adjustments.
- Keep vdW for manipulated: Decide whether to include Van der Waals interactions for atoms manipulated with the mouse. This feature helps when connecting or adjusting atoms, as disabling vdW forces simplifies movement and reduces interference.
Running and Customizing IM-UFF Simulations
Once your system is set up, running a simulation is easy. Simply ensure the Static topology (UFF only) and Keep vdW for manipulated options are unchecked, then select Edit > Simulate > Start. During the simulation, you can:
- Visualize total system energy as well as individual energy components in real time.
- Interactively adjust atom positions to observe how molecular topologies adapt.
- Form or break covalent bonds as necessary, empowering you to refine structures dynamically.
IM-UFF also lets you customize parameters like the Van der Waals cutoff, switching distance, and neighbor list periodicity for fine-tuned simulations. Additionally, users can update atomic typizations on the fly, ensuring accurate models even as molecules evolve during editing or simulation.
For a closer look at how IM-UFF handles dynamic topology changes, check out the animation below:
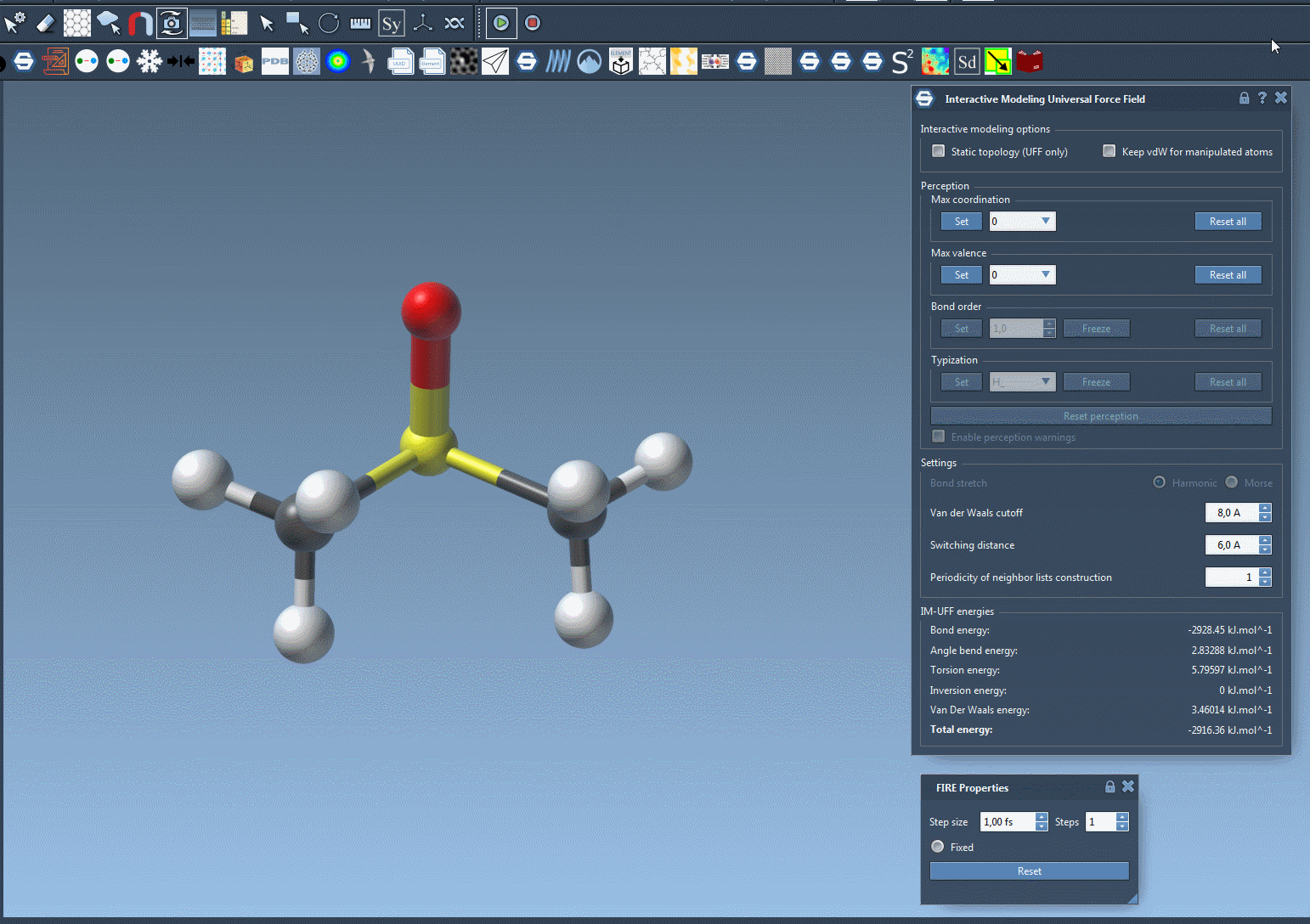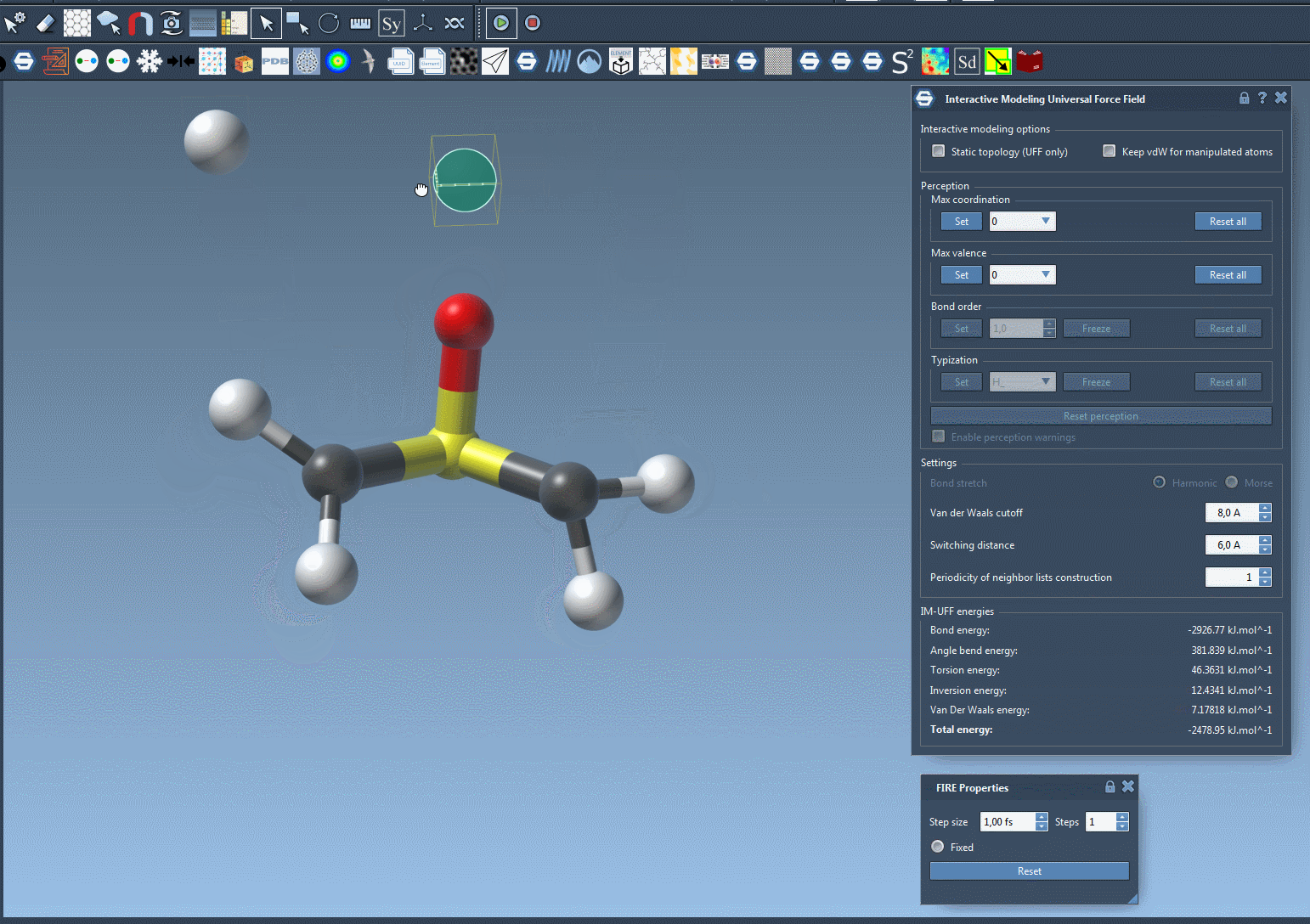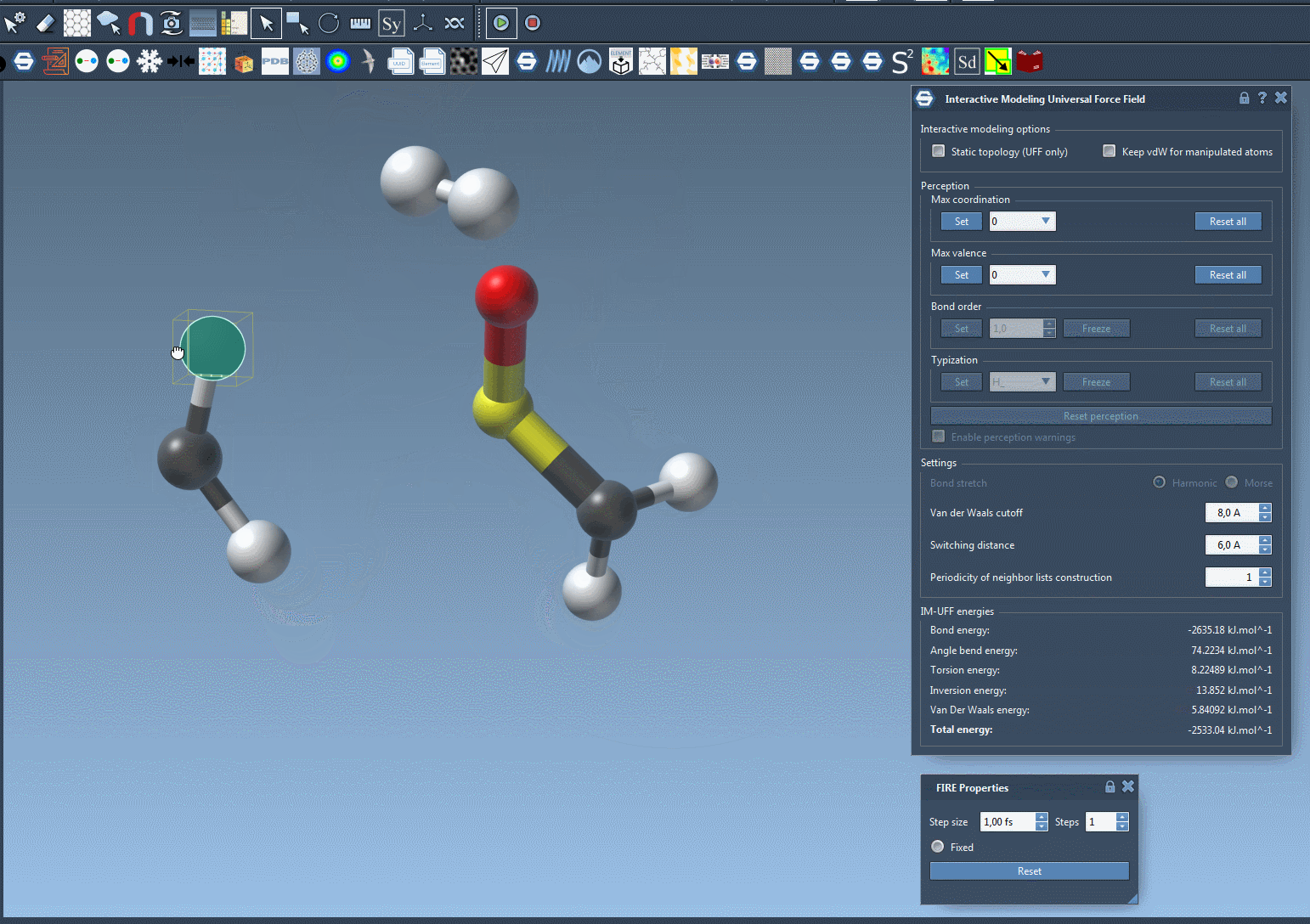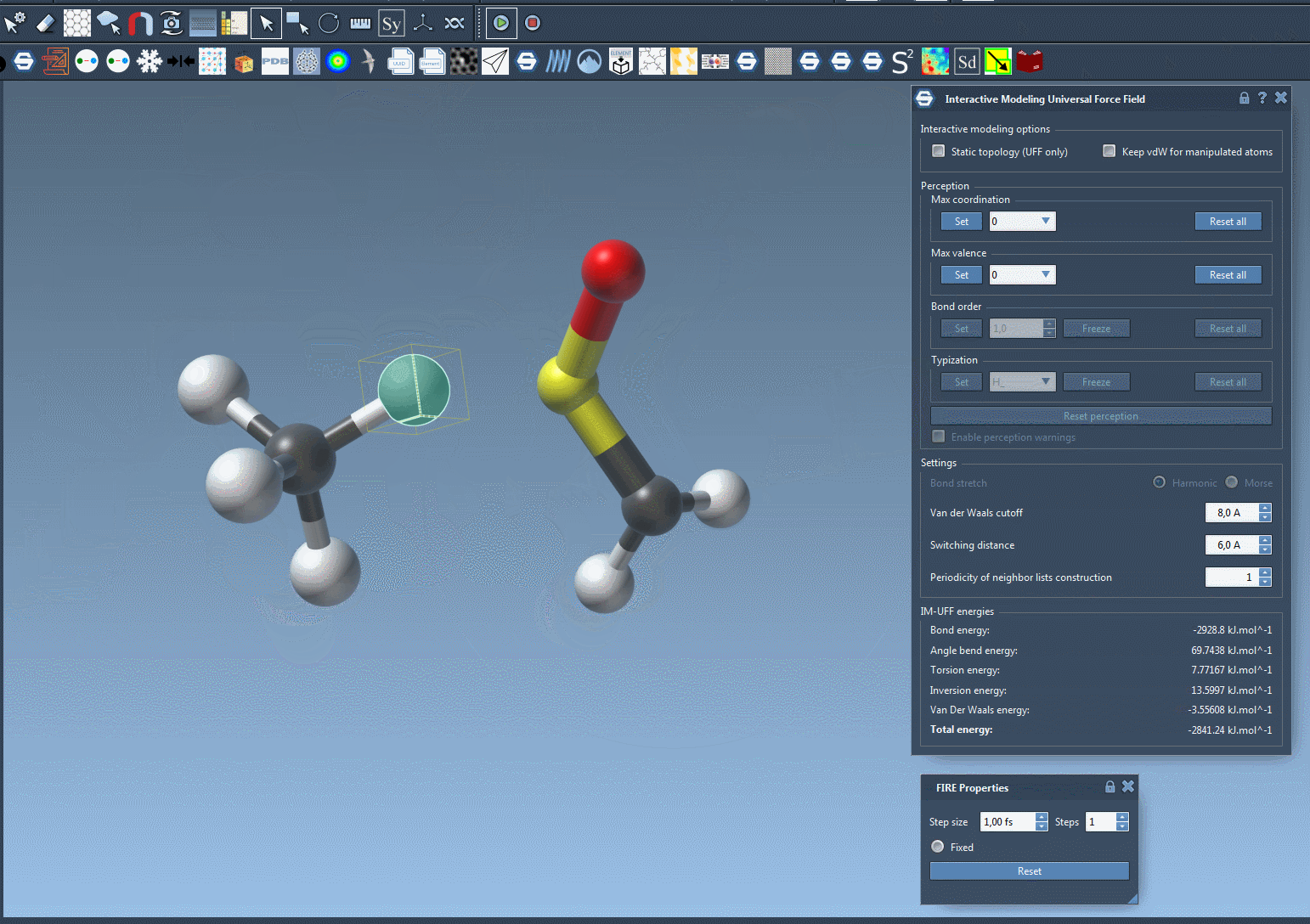
Learn More
IM-UFF simplifies molecular modeling by combining user interactivity with robust force-field calculations, enabling you to tackle complex restructuring tasks effortlessly. Whether you’re designing molecules, studying system behavior, or refining structures, this tool can make the process more intuitive and flexible.
To dive deeper into IM-UFF’s capabilities or explore advanced features, visit the full official documentation.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Download SAMSON today at https://www.samson-connect.net.





