Molecular modelers often face the challenge of visualizing and understanding the intricate movements of biological structures under varying conditions. A pressing topic in recent molecular research has been the motion of the SARS-CoV-2 spike protein, a key player in the virus’s entry into human cells. For researchers and enthusiasts, detailed insights into this motion can provide crucial understanding for vaccine development and treatment strategies. Here, we explore how the spike transitions between its closed and open states using computational modeling tools.
Why Focus on the Spike Protein?
The SARS-CoV-2 spike protein plays a central role in viral infection by recognizing and binding to a host receptor known as ACE2 (Angiotensin-Converting Enzyme 2). This process initiates the entry of the virus into human cells, making the spike a critical target for neutralizing antibodies. A deep understanding of its motion offers valuable insights into how the virus operates and how treatments can be designed to block this mechanism.
The Motion: Transitioning Between Closed and Open States
A fascinating feature of the spike protein is its ability to switch between two major states: a closed state that conceals the receptor-binding domain and an open state that exposes the domain to bind ACE2. This dynamic behavior is essential for its function and opens doors for in-depth computational exploration.
The motion of the spike protein between these states has been computed using SAMSON, an integrative molecular design platform. Using known structural data—closed-state structure PDB 6VXX and open-state structure PDB 6VYB—the pathway of this transition was simulated, providing key insights into the intermediate conformations and energy landscapes involved in this motion.
Visualizing the Motion and Trajectories
Thanks to computational methods, various visualizations can showcase the intricate dance of the spike protein during its transition. Below are examples of how the spike appears at different stages:
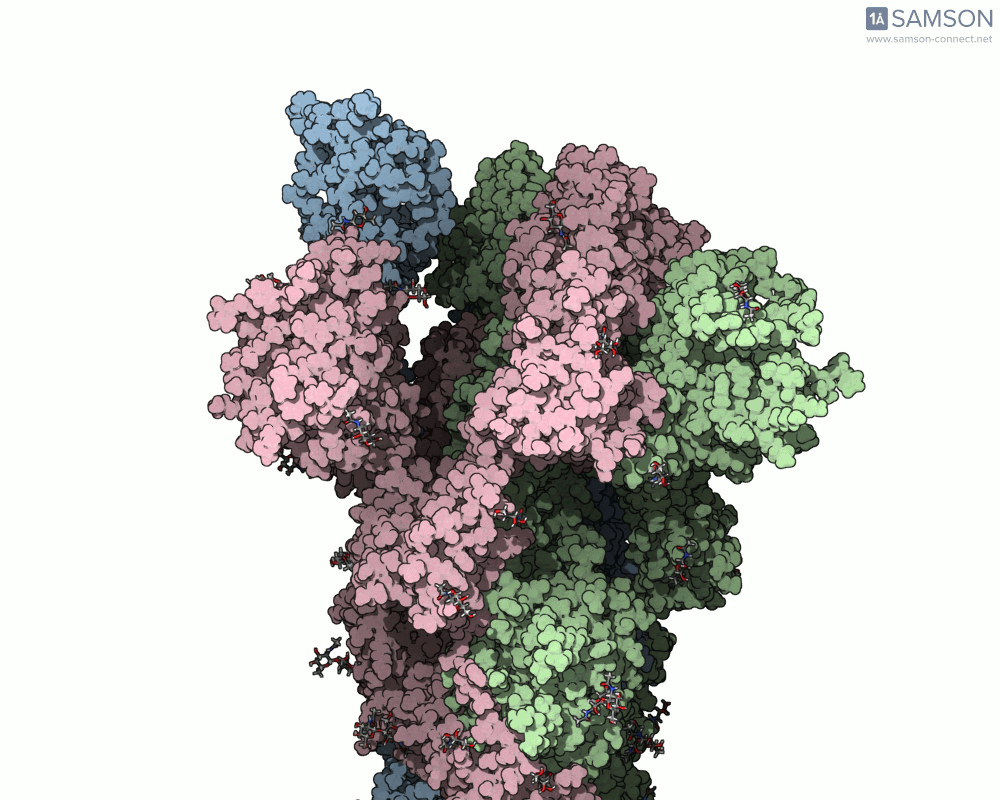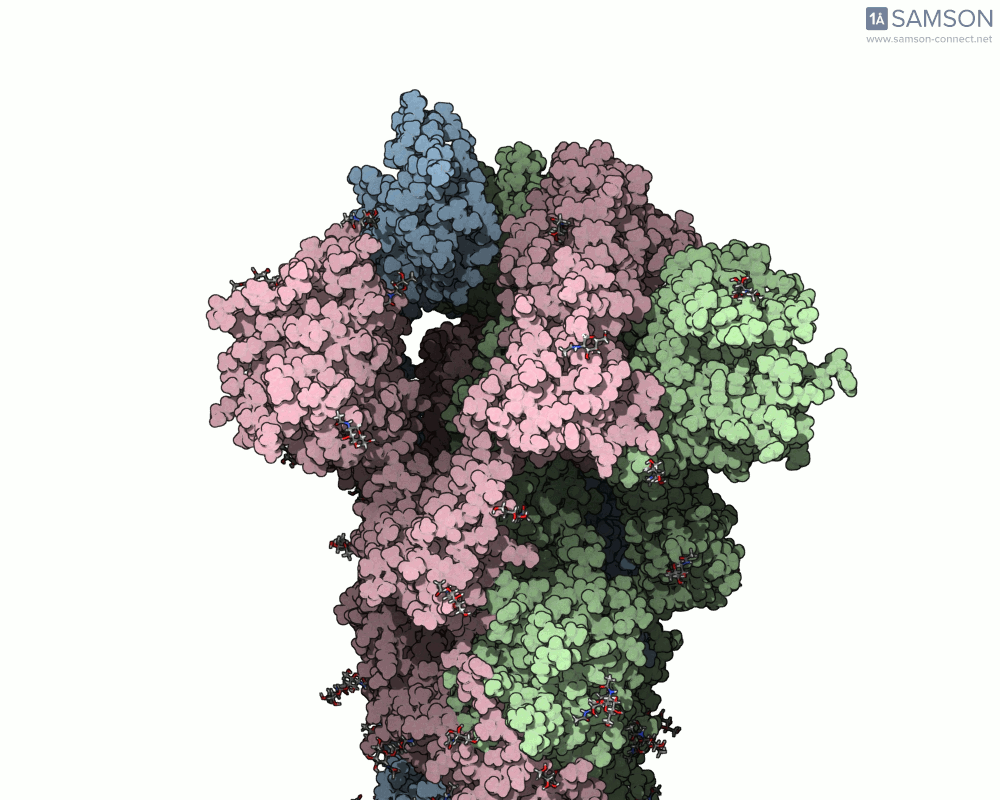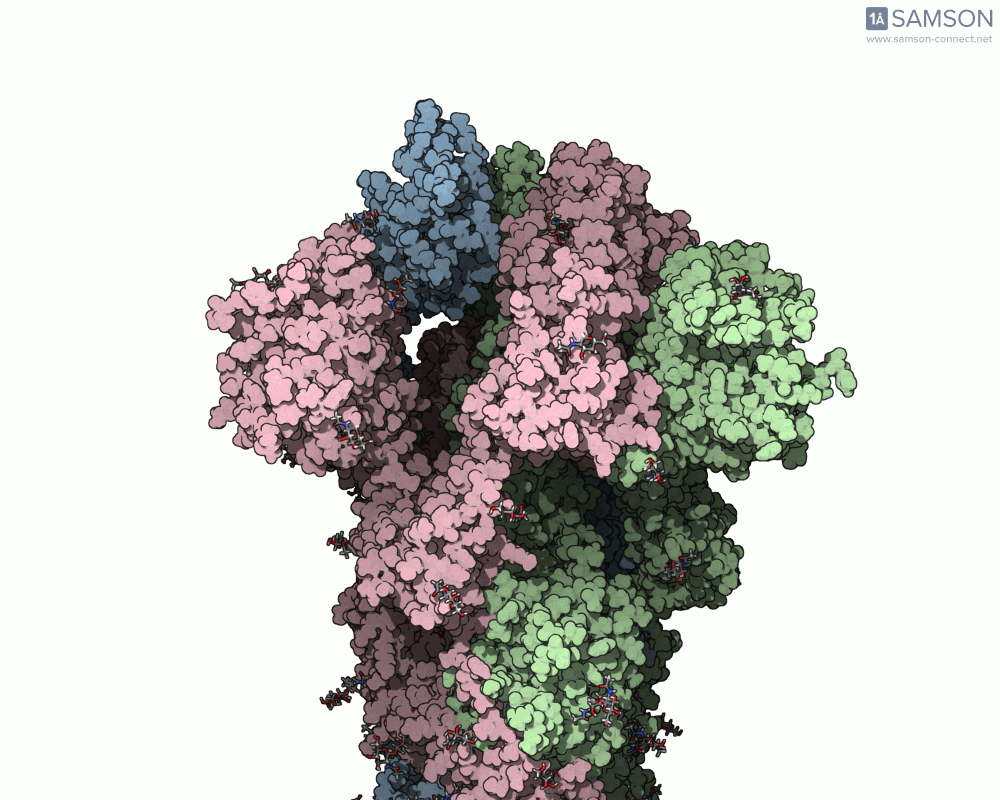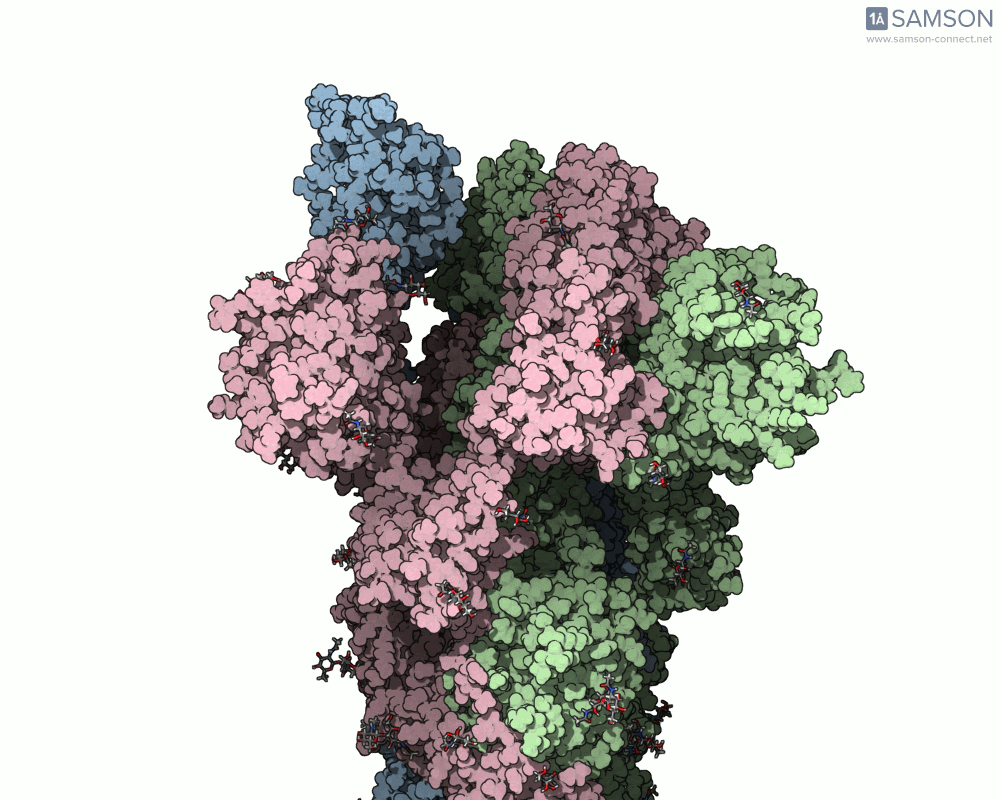
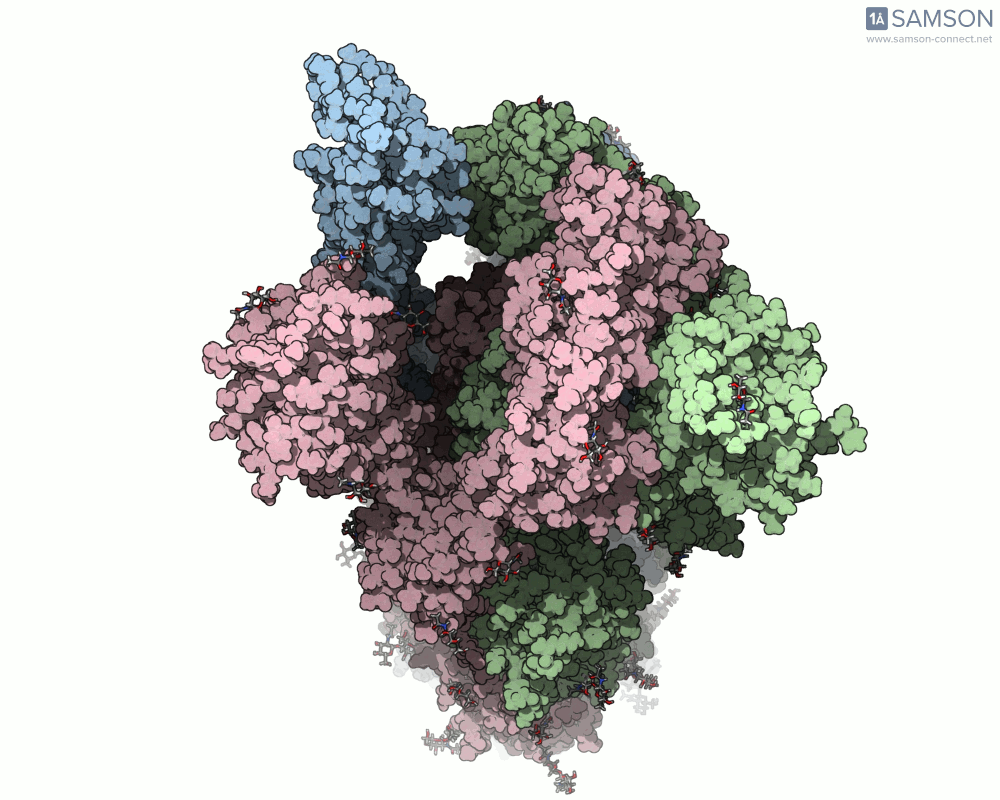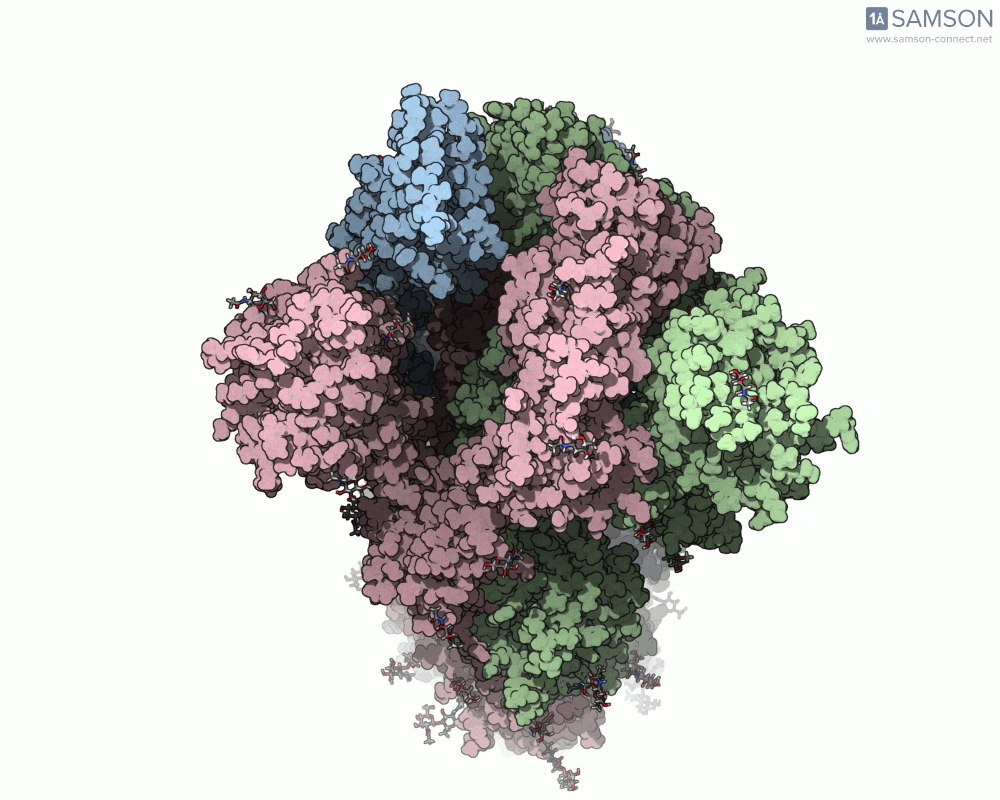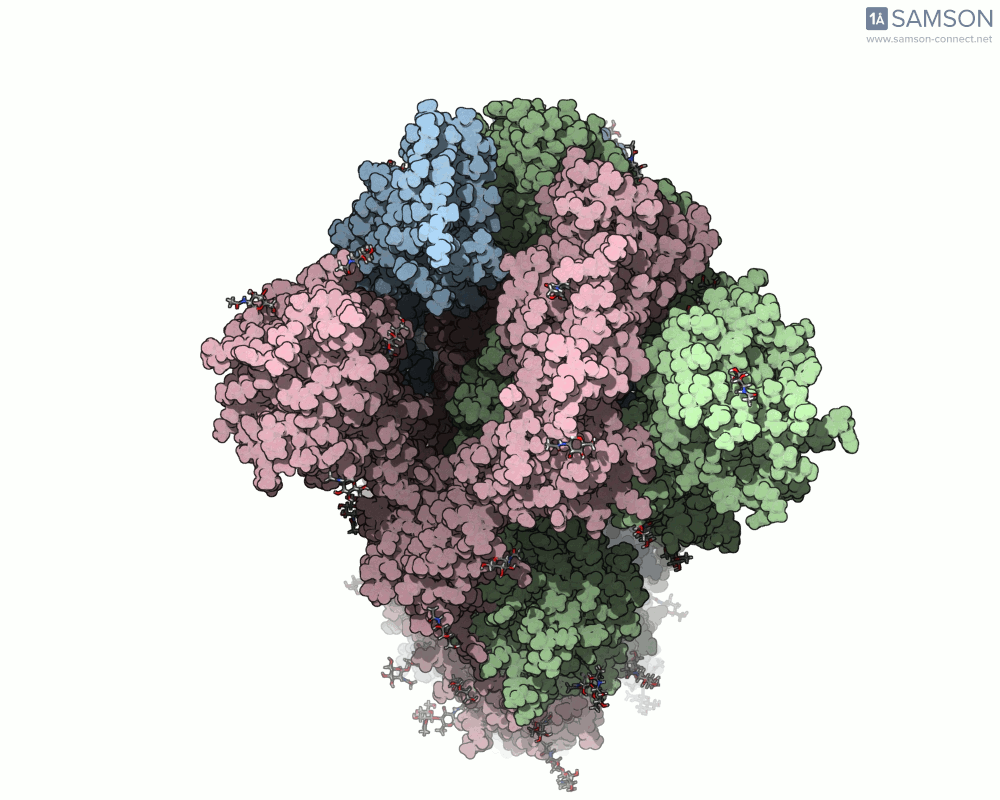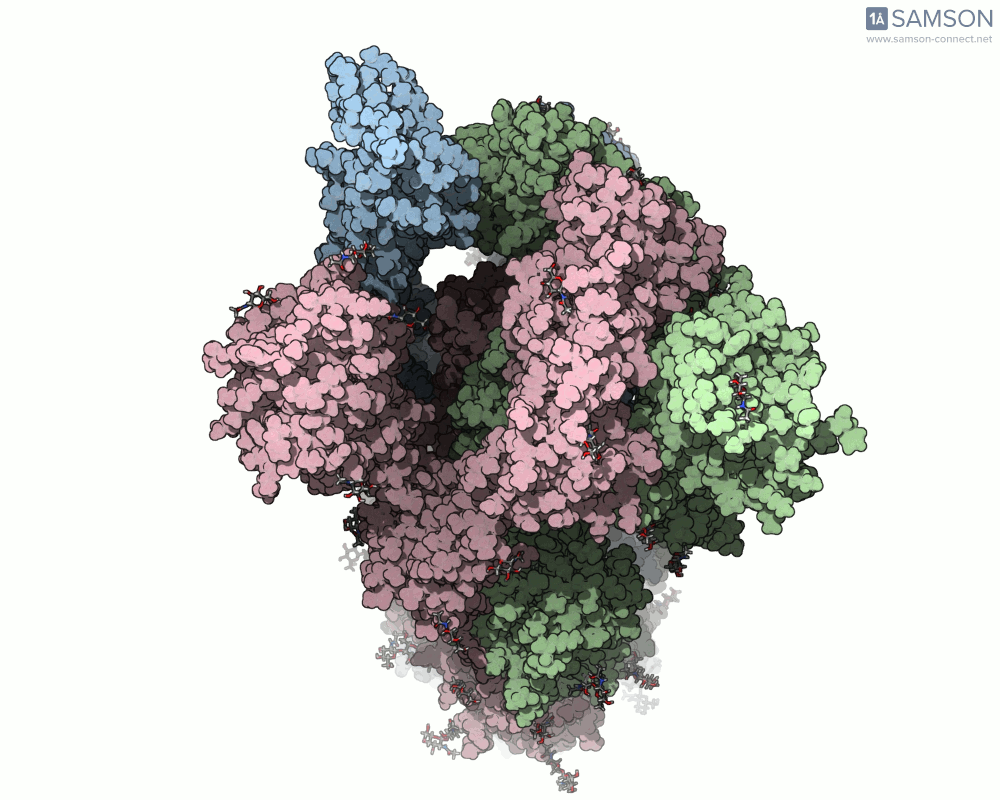
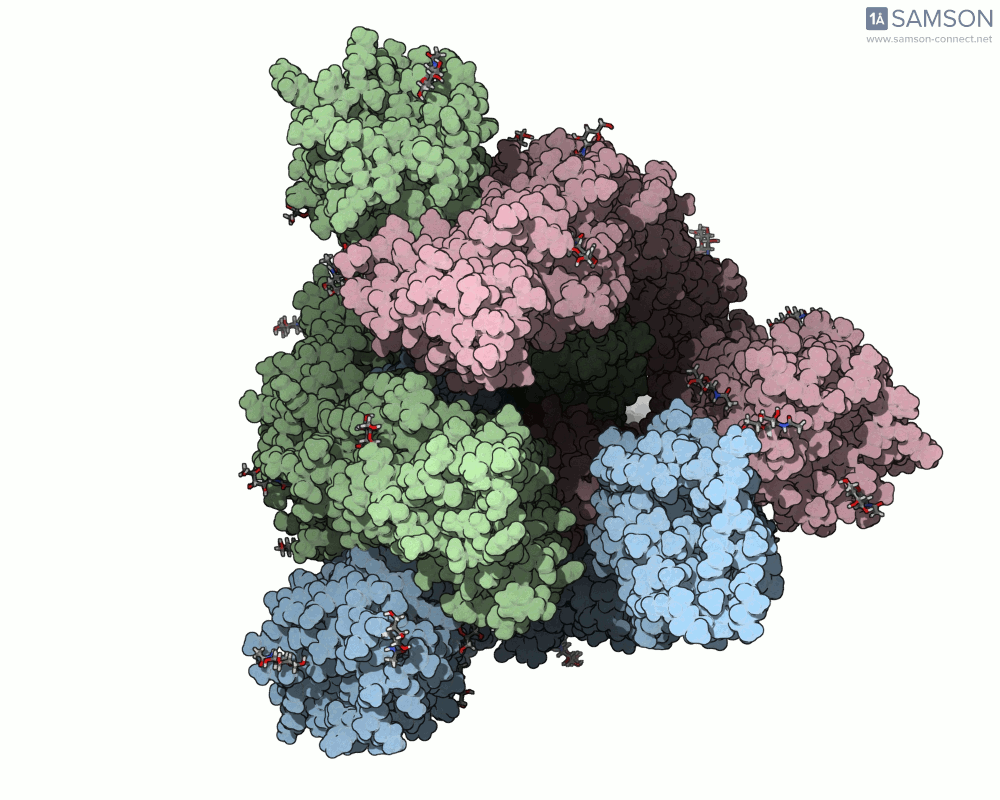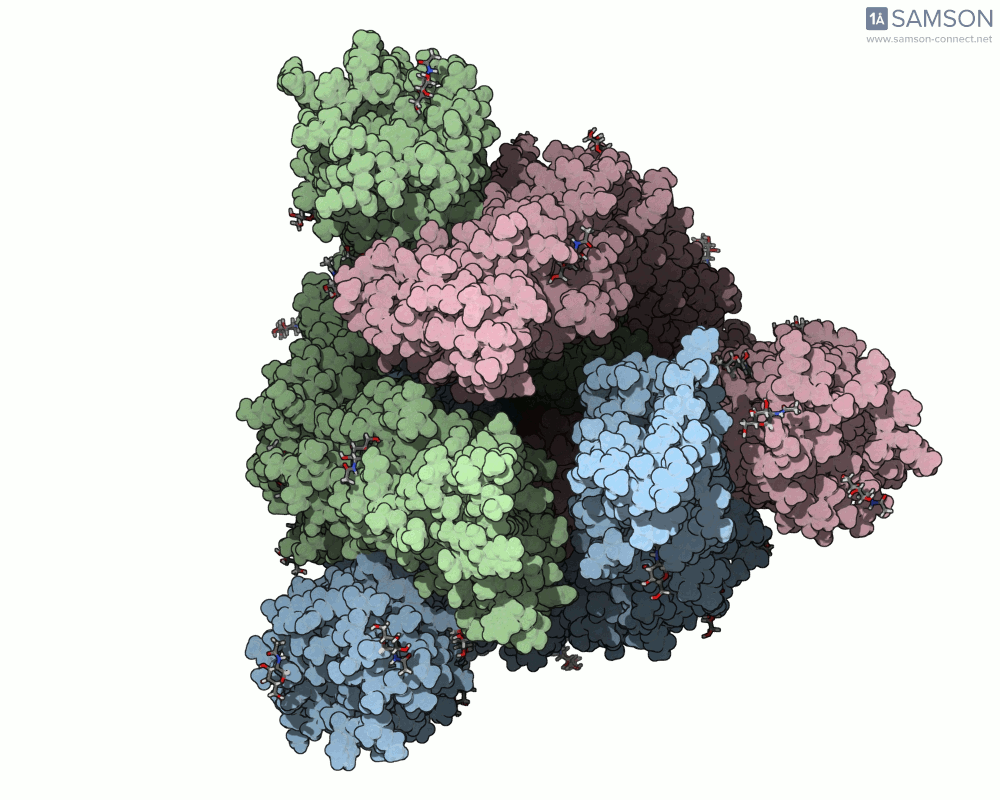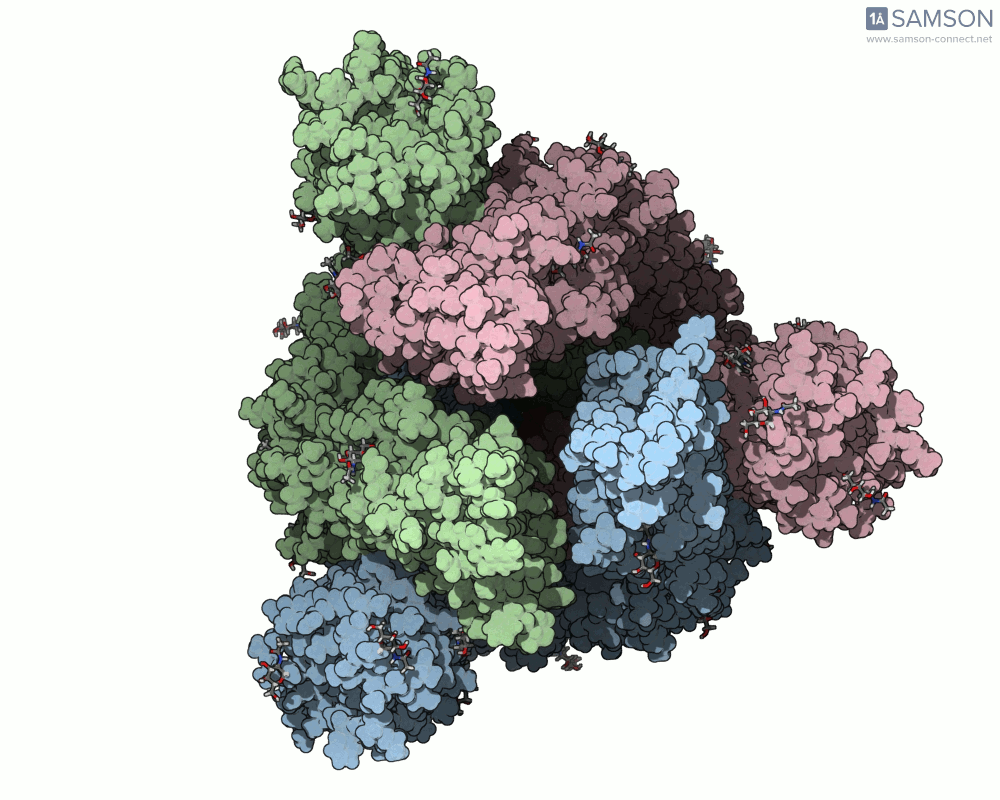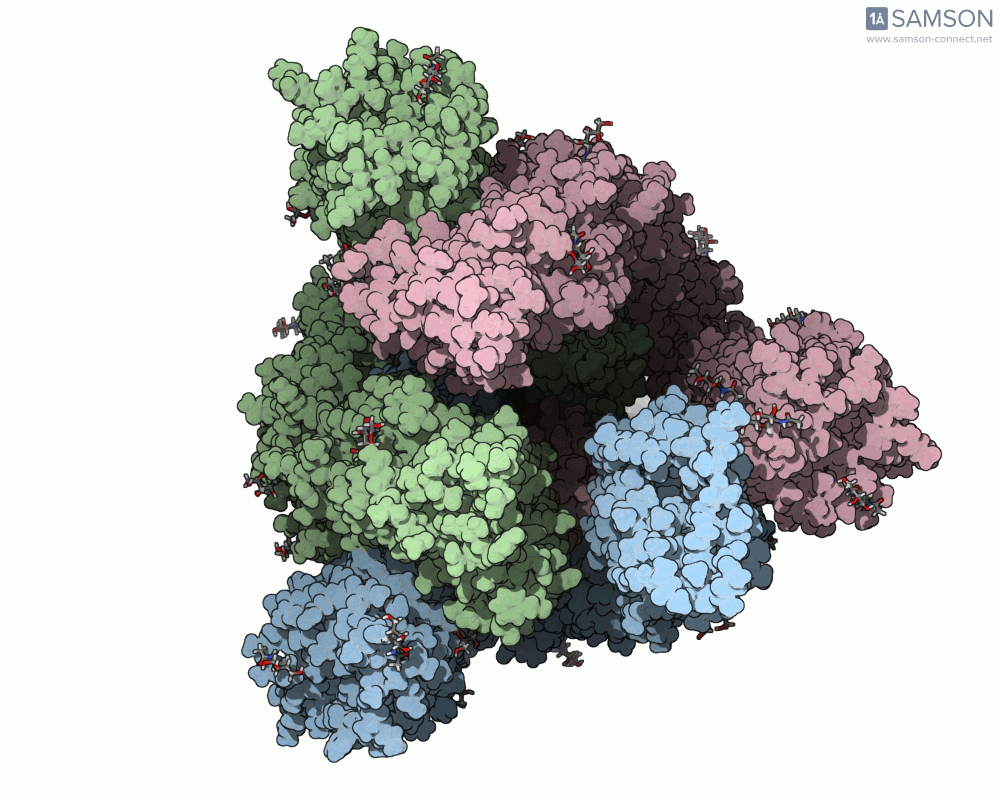
These animations highlight the protein’s flexibility, offering vital clues to researchers studying its interaction with the ACE2 receptor.
How This was Computed
Using SAMSON’s advanced modules, the following pipeline was employed:
- A combination of the ARAP Interpolation Path module and the P-NEB (Parallel Nudged Elastic Band) module was used to simulate and refine the intermediate states of the spike protein.
- Modifications, minimizations, and interpolation steps prepared the structures for these simulations, ensuring accuracy even when the structures differed in residue counts.
- The results provided both a qualitative visualization and a quantitative understanding of the pathway between the closed and open states.
Such tools not only visualize molecular flexibility but also serve as an initial step for further simulations, such as drug binding or antibody interaction studies.
Applications and Outlook
The ability to computationally reproduce and visualize spike protein motion provides opportunities for drug and vaccine development, offering a detailed understanding of its receptor-binding behavior. Whether you are computationally designing molecules or assessing drug efficacy, understanding large protein motions is increasingly important in molecular modeling workflows.
Would you like to delve deeper into how SAMSON computed this motion? Visit the original documentation page here.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Download SAMSON at https://www.samson-connect.net.





