Molecular modelers and researchers working to understand the SARS-CoV-2 virus often encounter a specific challenge: analyzing and simulating the complex opening motion of the virus’s spike proteins. This motion is crucial for studying how the virus interacts with human cells, but modeling it accurately can be daunting. If you’re working in this domain, SAMSON, the integrative molecular design platform, provides a guided solution to explore and compute the spike protein’s opening motion. Here, we delve into the importance of this spike motion, describe how it can be analyzed, and highlight the tools that make this possible.
The Significance of Spike Motion
The spike of the SARS-CoV-2 virus is composed of glycoproteins and facilitates the virus’s entry into human cells. It achieves this by transitioning from a closed state to an open state, enabling it to bind to the human ACE2 receptor molecule. This receptor-binding domain is strategically located at the top of the spike, a region exposed due to functional constraints, making it a key target for neutralizing antibodies.
The spike’s structure has unique properties—its C3 symmetry, three protein subunits, and its sugar-coated defense mechanism. These characteristics allow it to evade the immune system effectively while leaving critical regions exposed for cell recognition. Understanding, visualizing, and simulating this “open-close” motion can help develop insights into neutralizing strategies or drug designs targeting COVID-19.
Visualizing the Spike and its Movement
Using SAMSON, you can examine detailed visualizations of the spike protein’s closed and open conformations. For example:
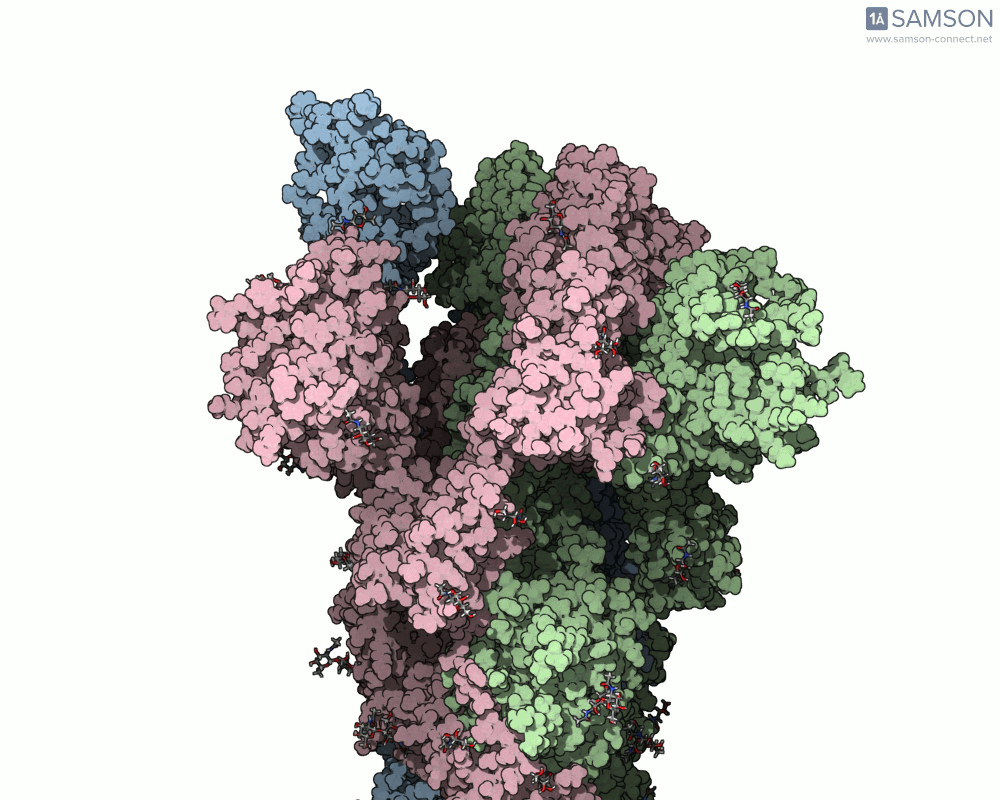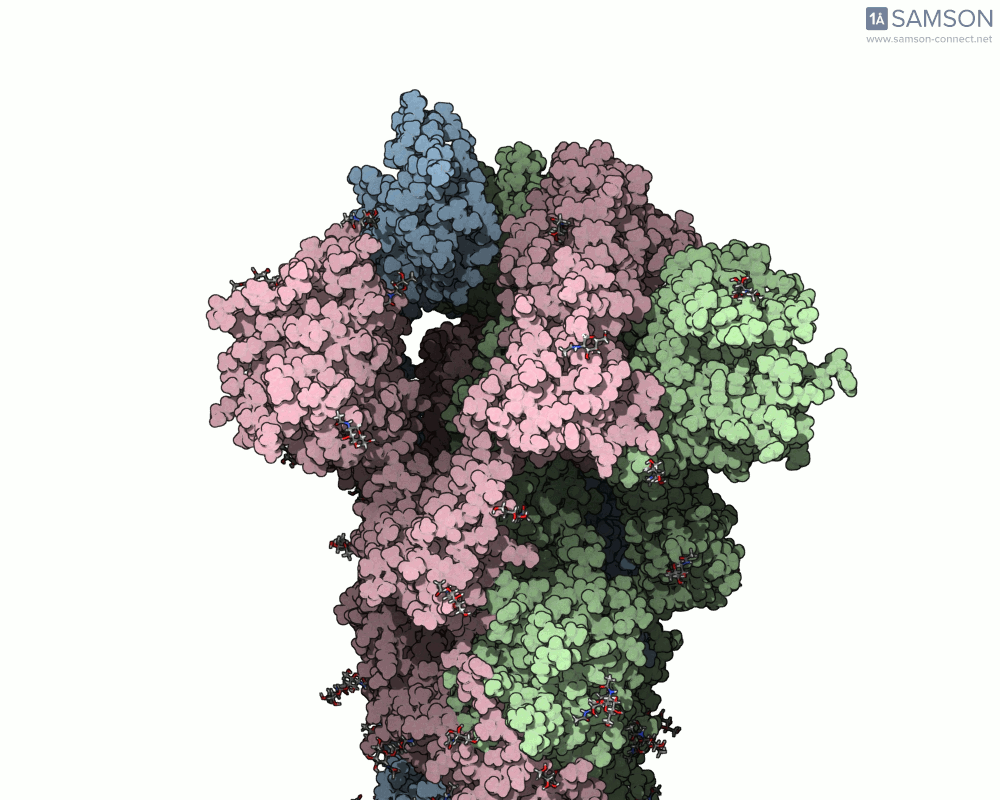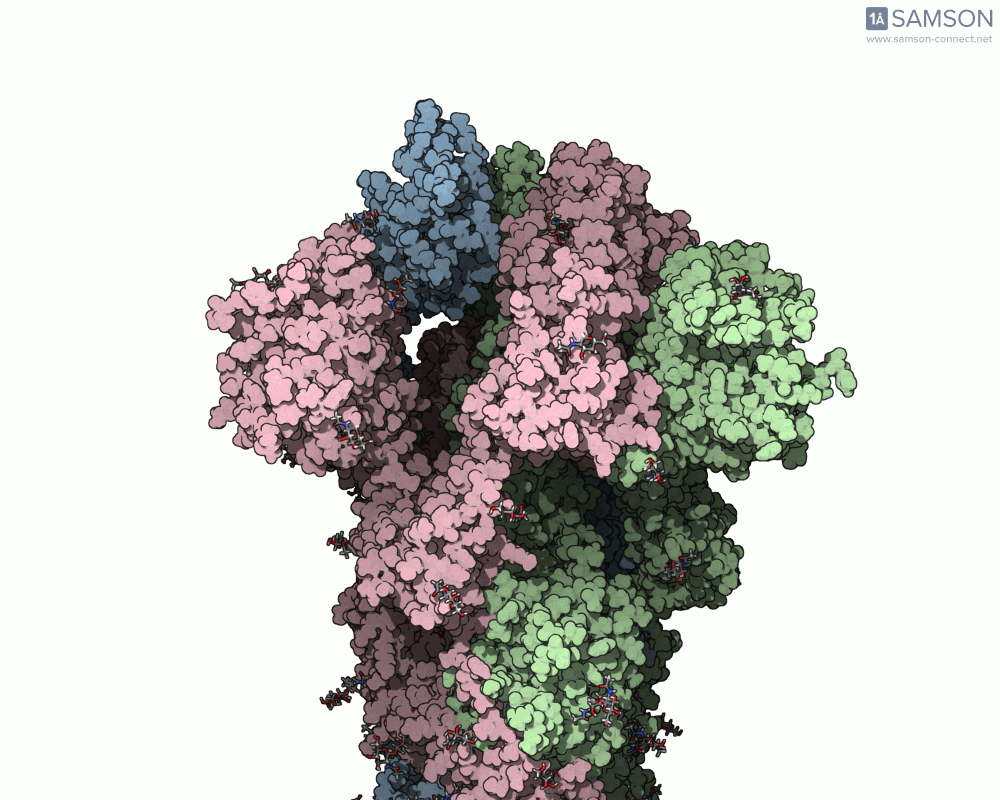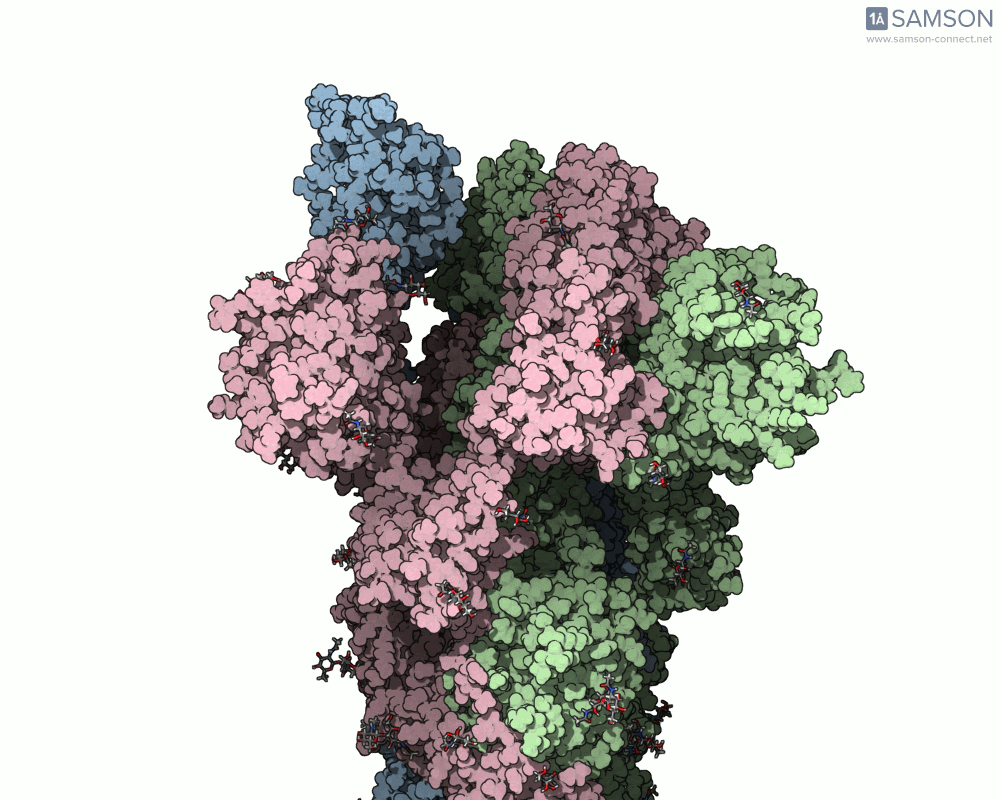
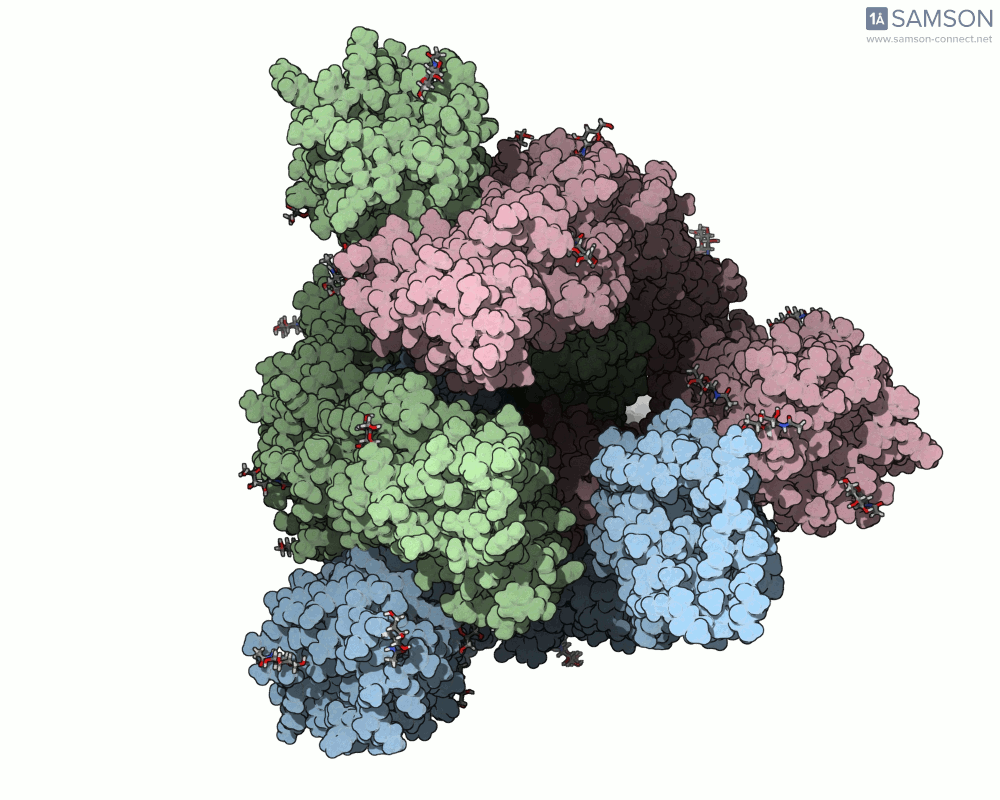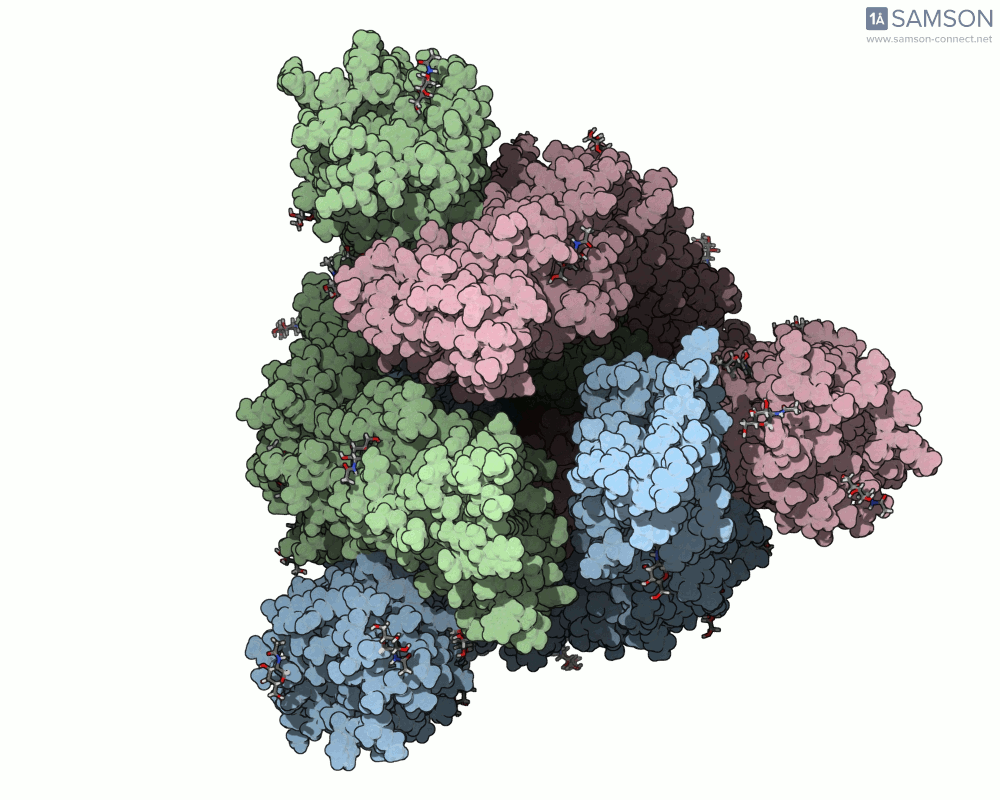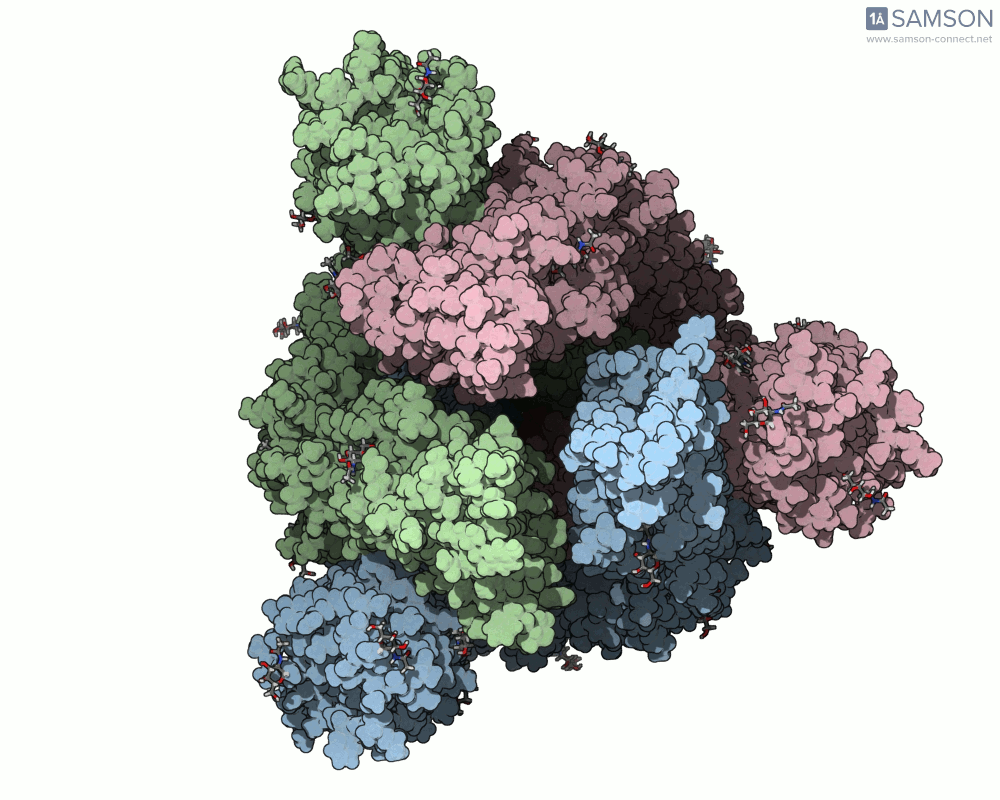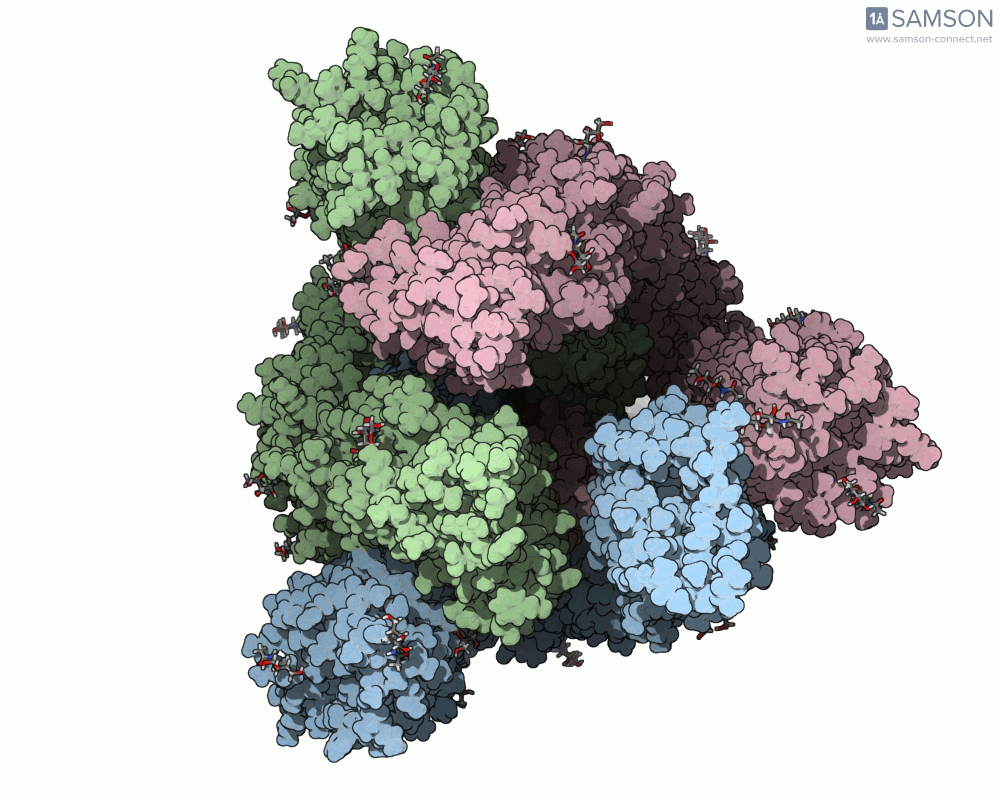
These animations showcase how the spike transitions from its down state (closed) to its up state (open), the latter being capable of recognizing the ACE2 receptor. The downloadable computed trajectory, provided in multiple formats (e.g., PDB), allows users to directly observe and analyze these pathways.
Explore downloadable trajectory files
- Set of PDB trajectory files: Download here.
- Single PDB file: Download here.
- SAMSON format file: Download here.
The SAMSON format file is particularly useful as it integrates the spike structure, animations of conformational changes (accessible with double-clicks), and trajectory computation results using specific modules. These files, while not experimentally verified, serve as valuable illustrations of the spike’s behavior and can be integrated into further computations and workflows.
How to Recreate the Motion
SAMSON provides a detailed step-by-step pipeline for generating the spike’s motion between its open and closed states:
- Prepare PDB structures for the open (PDB: 6VYB) and closed (PDB: 6VXX) states.
- Use the “As-Rigid-As-Possible (ARAP) Interpolation Path” module to simulate the interpolated motion connecting the two states.
- Refine and optimize the path using the “Parallel Nudged Elastic Band (P-NEB)” module to achieve a more accurate trajectory.
Both the ARAP and P-NEB modules are powerful tools that allow for seamless trajectory analysis. Setup and use are straightforward, and their computations are efficient even on standard laptops.
Accessing SAMSON Tools
The ARAP and P-NEB modules, along with many other SAMSON tools, are currently free for everyone during the COVID-19 pandemic. This ensures that researchers can easily explore and simulate processes critical to SARS-CoV-2 research. Simply sign up for free on SAMSON Connect, download SAMSON, and get started.
To dive deeper into the SARS-CoV-2 spike motion and explore available tools, visit the full documentation page here.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Begin your journey into molecular design today by downloading it at SAMSON Connect.





