When working on molecular design and protein modeling, understanding the arrangement of proteins in crystal structures or biological assemblies can often be a crucial step. These arrangements provide insights into protein-protein interfaces, binding sites, and overall structural symmetry. This is where SAMSON’s Symmetry Mate Editor becomes indispensable, allowing you to generate symmetric replicas of proteins directly from transformation data stored in PDB files.
Why Symmetry Mates Matter
Reconstructing symmetry mates goes beyond visual appeal. Here are some practical benefits:
- Visualizing biological assemblies: It allows you to see how proteins interact within their crystallographic units.
- Exploring protein interfaces: You can identify potential binding sites or study protein-protein interactions.
- Reconstructing quaternary structures: This is essential for molecular dynamics simulations and molecular design workflows.
- Designing symmetric nanostructures: Create symmetric protein complexes for advanced applications in nanotechnology or drug delivery.
Getting Started
- Install the Symmetry Mate Editor extension in SAMSON if you haven’t already.
- Open a PDB file containing symmetry information. This metadata is often included in CRYST1 or BIOMT records within PDB files.
If you’re new to this, CRYST1 describes symmetry from the crystal lattice, while BIOMT represents annotations for biological assemblies. The Symmetry Mate Editor supports toggling between these two types, helping you to choose which representation suits your objectives better.
Interactive Tools for Visualization
The Symmetry Mate Editor is designed for user interactivity, making the workflow efficient and intuitive:
- Activate the Editor: Use the Find everything command (Shift + E) or access it via the viewport menu (… > General > Symmetry Mate Editor).
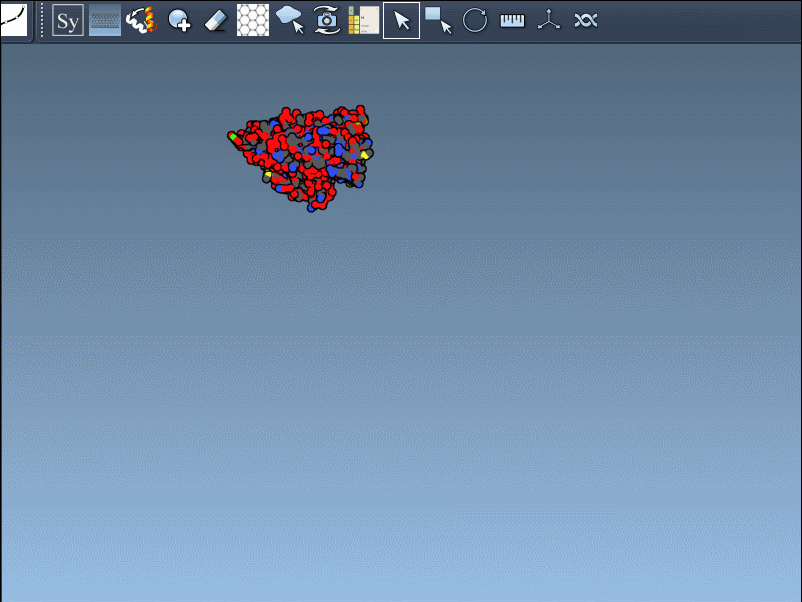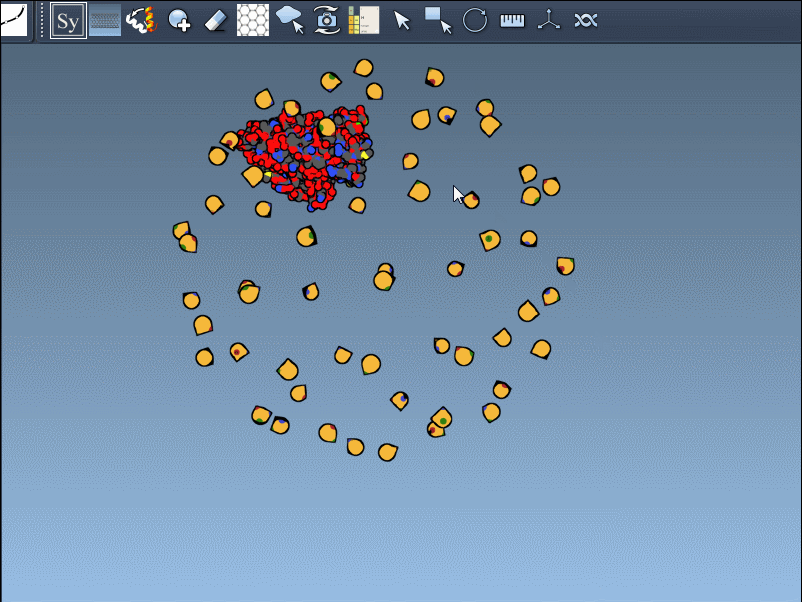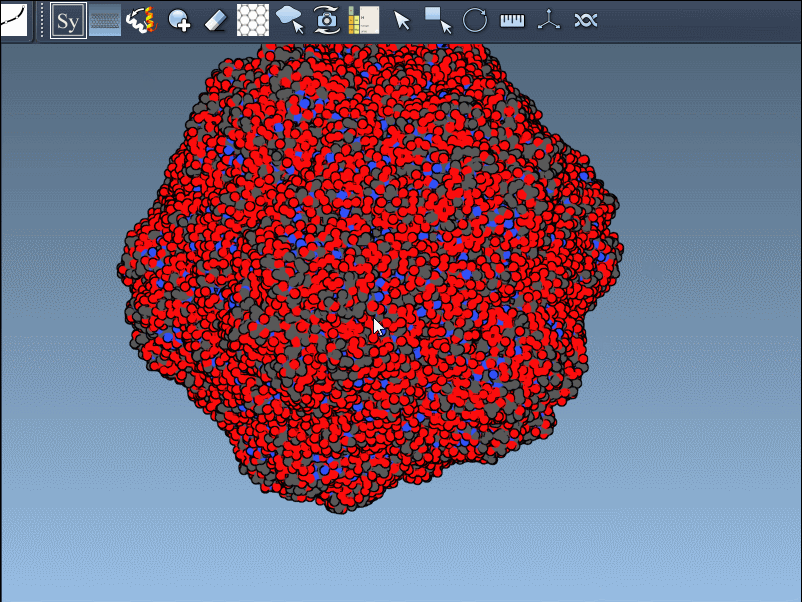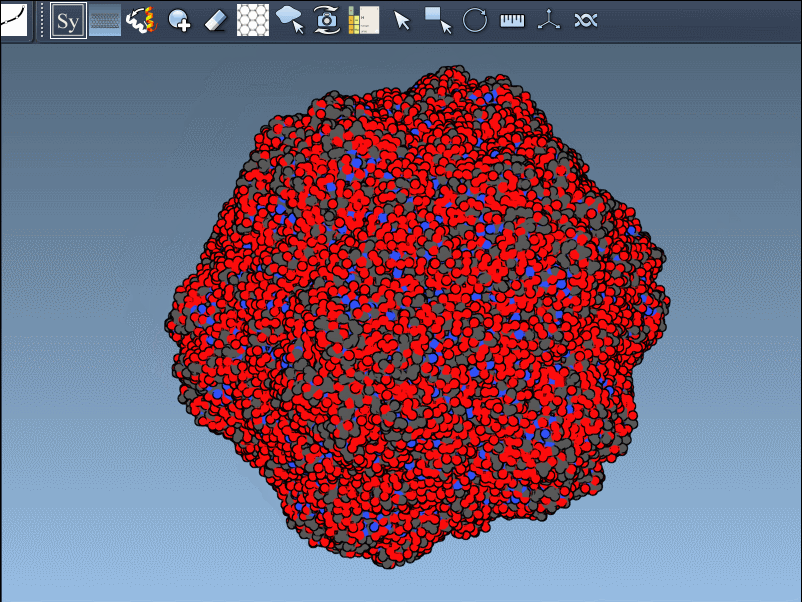
- Scaling replicas: Adjust the number of visible control nodes by holding Ctrl/Cmd and scrolling your mouse wheel.
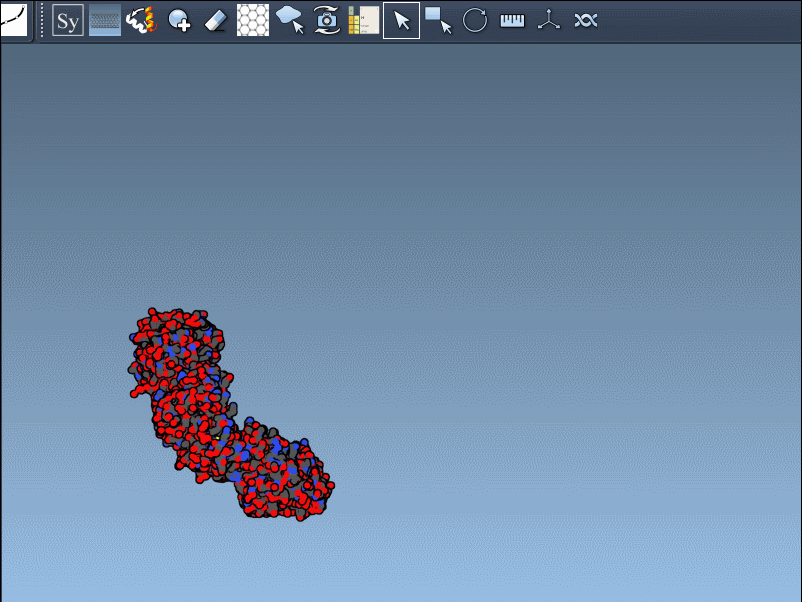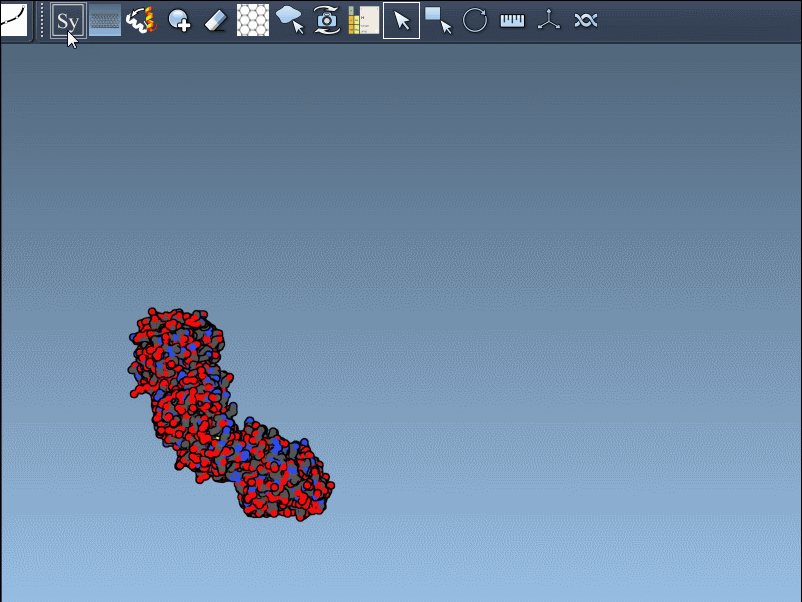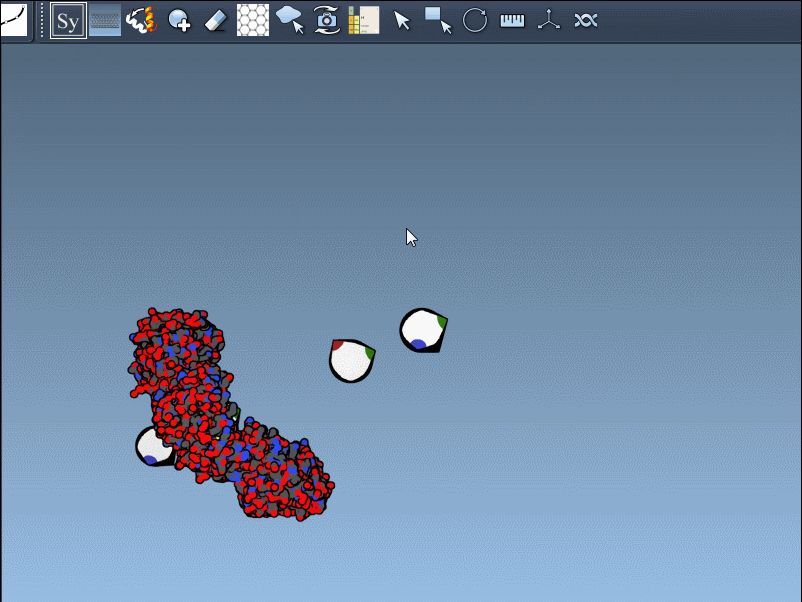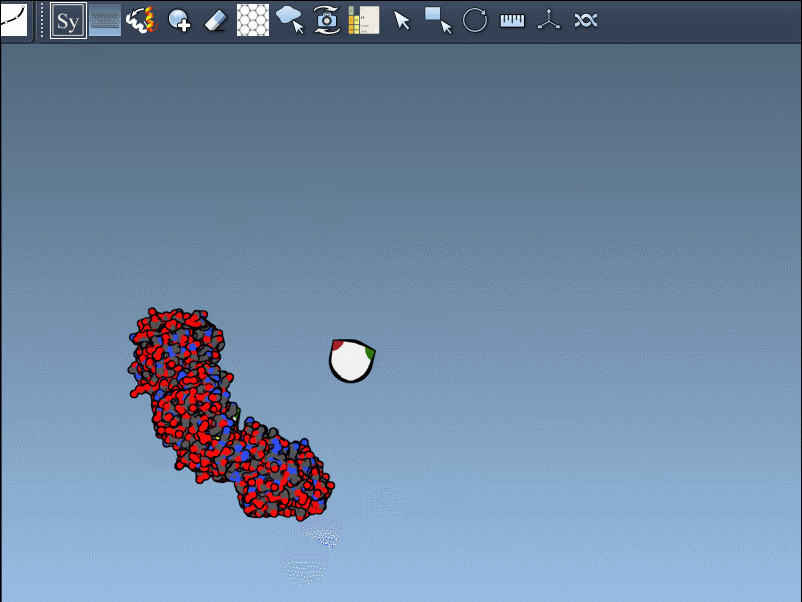
- Real-time preview: Hover over a control node to preview replicas instantaneously. Left-clicking a node generates the replica permanently.
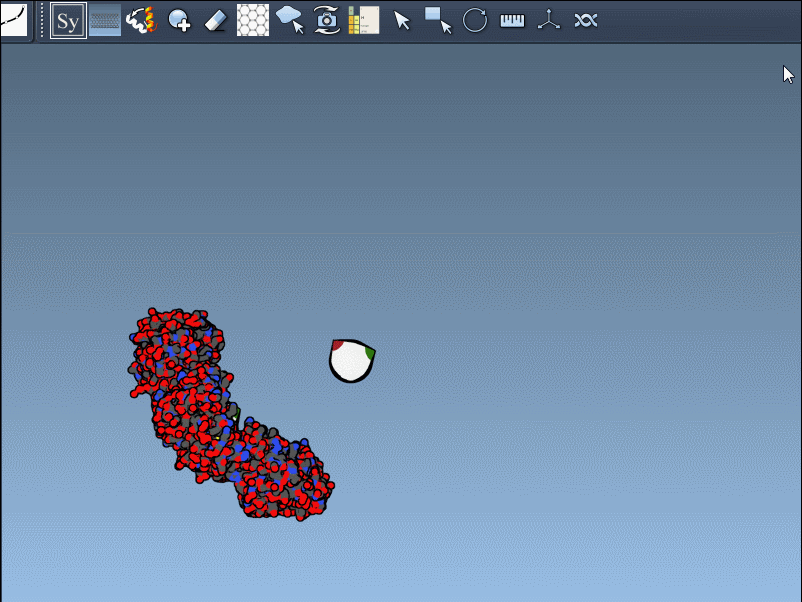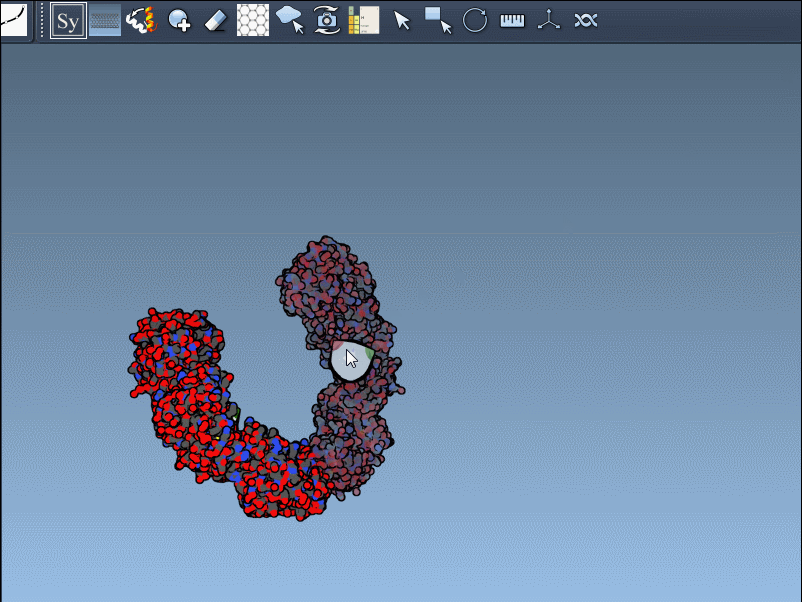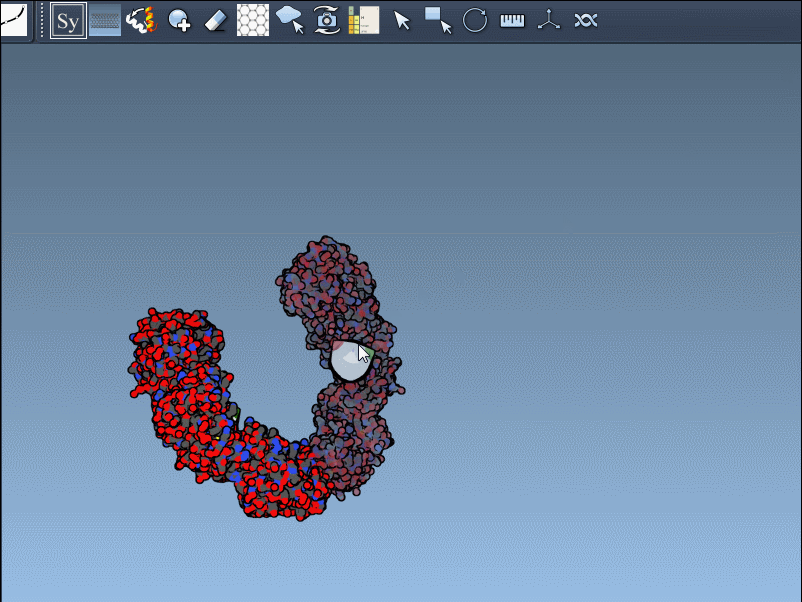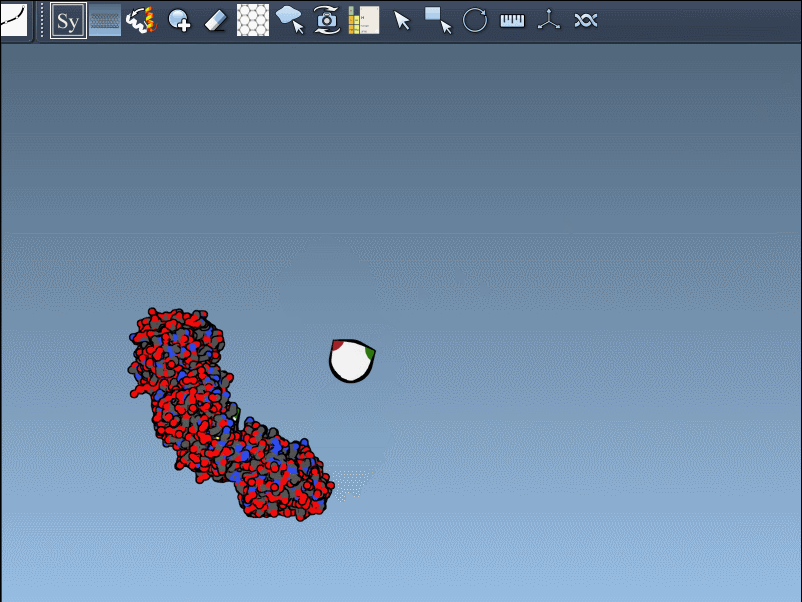
- Generate all replicas at once: To create an entire symmetric structure in one action, hold Ctrl/Cmd while clicking a control node.
Applications in Molecular Design
By generating symmetry mates, researchers and designers unlock several capabilities:
- Build complete oligomeric assemblies from asymmetric units.
- Simulate the interactions between symmetric chains during molecular dynamics studies.
- Design protein-based scaffolds or cages for structural biology experiments.
- Test binding site repeats across symmetric mates for drug discovery applications.
Tips to Enhance Your Workflow
- Use the Ribbons visual model to distinguish chains by coloring each uniquely.
- Combine with other extensions like Symmetry Detection to analyze axes and structural alignments.
- Don’t worry about mistakes—use Ctrl/Cmd + Z to undo unwanted replicas.
Recreating symmetric protein structures is a powerful tool for molecular modeling and design. To access more details and further guidance, visit the full documentation here.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON at https://www.samson-connect.net.





