If you’re generating protein transition paths and encountering the dreaded error message:
“Cannot proceed because the structure does not make one connected component”
you’re not alone. This issue can be frustrating, especially when everything else seems correctly set up. But the good news is: it’s easy to prevent – if you know what’s causing it.
What’s really going on?
When SAMSON’s As-Rigid-As-Possible (ARAP) Interpolator runs, it needs to work with clean, connected protein structures. That means the system it’s processing must form a single connected molecular graph (i.e., one component). If there are disconnected parts—like water molecules, ions, or alternative residue locations—it can confuse the algorithm and prevent interpolation.
This is particularly common when fetching structures from the Protein Data Bank (PDB), as downloaded entries often include:
- Water molecules
- Ions like Na+, Cl–
- Ligands or drugs bound in the crystal structure
- Alternate atom positions (especially in side chains)
These seemingly minor details end up breaking the assumption that your protein is one cohesive, connected object.
The simple fix: Prepare first
Before using the ARAP extension, run the Home > Prepare step. This built-in feature in SAMSON automatically cleans up both input structures and ensures the system is in a state that ARAP can handle.
It performs several essential tasks:
- Removes alternative locations
- Deletes water molecules and solvents
- Removes ligands and ions
By doing this early, you avoid possible failures later in your workflow.
A helpful tip: Know what’s inside your structure
When you’re visualizing fetched structures, it’s good practice to browse the Document view in SAMSON and manually inspect what’s inside each system:
- Are there multiple chains?
- Are there ligands or water molecules?
- Is everything you see relevant to your motion of interest?
If not, don’t hesitate to delete them manually or rerun the prepare step. SAMSON even provides visual tools to make inspecting and editing structures intuitive. You can use Del or click the erase button to remove chains or molecules you don’t need.
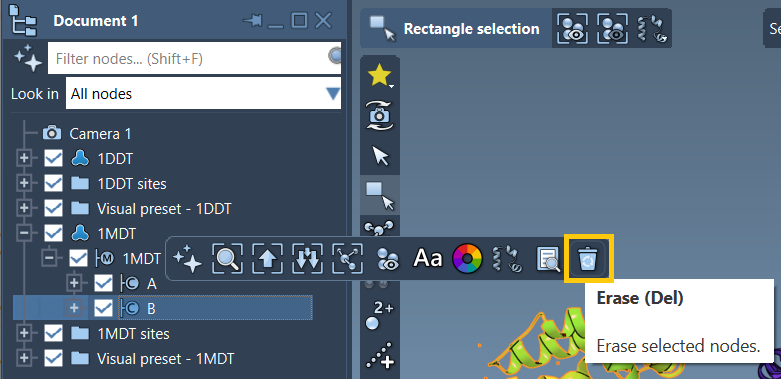
This figure shows an example of removing chain B from one of the structures (1MDT). In this workflow, only chain A is needed for the transition path.
Prevention = Smooth interpolation later
Taking a few seconds at the beginning to prepare your structures pays off. It ensures that ARAP interpolation runs on a clean, connected system—and that results are reliable. Without this, downstream steps like simulating transition states or performing umbrella sampling can be compromised.
If you’re experiencing issues, start by double-checking your structures. Are there leftover components? Did you skip preparation? If yes, that’s likely your answer.
You can consult the full tutorial for ARAP-based interpolation here: https://documentation.samson-connect.net/tutorials/arap/arap-interpolation-for-protein-structures/
SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON at https://www.samson-connect.net.





