One of the challenges in molecular modeling lies in identifying the most energy-efficient pathways a system might follow when transitioning between two states. Whether you’re studying a ligand unbinding trajectory or the conformational changes of a protein, finding accurate transition paths is crucial for understanding these complex biomolecular processes. This is where the Parallel Nudged Elastic Band (P-NEB) method can make a notable difference.
The P-NEB Extension in the SAMSON integrative molecular design platform is a remarkable tool for improving transition paths. It uses the widely recognized Nudged Elastic Band (NEB) method to compute Minimum Energy Paths (MEPs) between two conformational states. With the P-NEB Extension, users can not only optimize pre-existing paths but also optimize a set of intermediate conformations, increasing the reliability of the results.
How Does It Work?
The P-NEB method relies on optimizing a series of “images” (intermediate states) between the two conformations of interest. By applying constraints such as spring forces, it ensures these images are spaced evenly along the path while simultaneously minimizing their energy. This dual-optimization workflow is further enhanced with parallel execution, which speeds up computational times by running multiple processes simultaneously.
Applying P-NEB to a Path
If you already have an initial path loaded in your SAMSON workspace, applying P-NEB is straightforward:
- Open the P-NEB app from Home > Apps > All > P-NEB.
- In the Document view, select your path node.
- Set the desired parameters in the P-NEB interface:
- Spring constant: Enter 1.00.
- Number of loops: Choose 100 optimizations.
- Interaction model: Use “Universal Force Field” for energy and force calculations.
- Optimizer: Select “FIRE” for optimization.
- Parallel execution: Enable this setting to speed up the calculations.
- Click on the Run button to start the computation.
The app will display the computational progress in the status bar. Once the optimization is complete, a new path will appear in the document view, representing the optimized transition path.
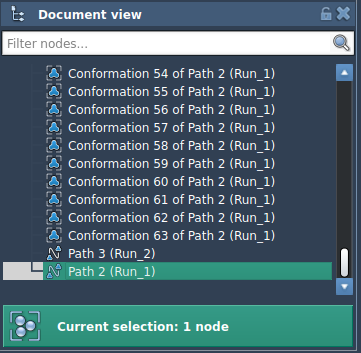
Optimizing Conformations
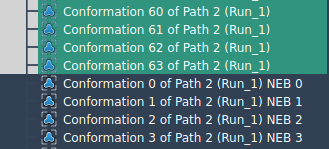
For situations where paths are unavailable and you only have a set of conformations, P-NEB can also optimize transitions based on these conformations. To do so, select the conformations in the Document view, then click “Run” in the P-NEB app. After computation, a new set of optimized conformations will appear, giving you valuable insights into the most likely transition mechanisms.
While optimizing conformations can take longer than optimizing an already-defined path, SAMSON allows you to combine conformations into a path for more efficient optimization through the context menu (Conformation > Create path from conformations).
Conclusion
Whether you’re working on understanding molecular mechanisms or exploring pathways for drug discovery, the P-NEB Extension in SAMSON offers a structured and powerful way to optimize transition paths. Its integration with SAMSON makes it highly accessible and customizable for researchers in molecular modeling.
If you’re ready to dive deeper, visit the official documentation page for a step-by-step guide: Optimize Transition Paths with the Parallel Nudged Elastic Band Method.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON at https://www.samson-connect.net.





