For molecular modelers, efficiently exporting atomic trajectories along predefined paths can greatly enhance workflows such as reaction coordinate studies, free energy calculations, and ligand pathway analysis. This guide focuses on how SAMSON’s Export Along Paths extension simplifies this process, saving you both time and effort.
Why Export Atom Trajectories?
Whether you’re studying ligand binding/unbinding, creating reaction coordinate files, or visualizing intermediate states, exporting the atomic coordinates along a path is an essential step for many projects. However, tracking specific atoms or exporting data in a structured way can often feel like a daunting task. The Export Along Paths extension within SAMSON resolves this by providing a user-friendly yet powerful toolset designed to streamline this process.
Step-by-Step Guide: Exporting Atom Trajectories Along Paths
Here’s a straightforward breakdown of how to use this feature in SAMSON.
Step 1: Open the Export Along Paths App
Start by opening the app via Home > Apps > All > Export Along Paths. Alternatively, you can use the search bar with the shortcut Shift + E to locate it quickly.
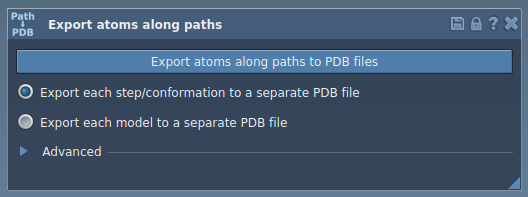
Step 2: Export All Atoms Along a Path
If you want to export all the atoms along a selected path, follow these steps:
- Select one or more paths in the Document view.
- Choose the export mode—either all frames contained in a single PDB file or each frame saved as an individual PDB file.
- Click the Export atoms along paths to PDB files button. You will be prompted to choose a destination folder and a file prefix for organized data handling.
Optional: You can define frame export intervals in the Advanced section to customize how much data you export.
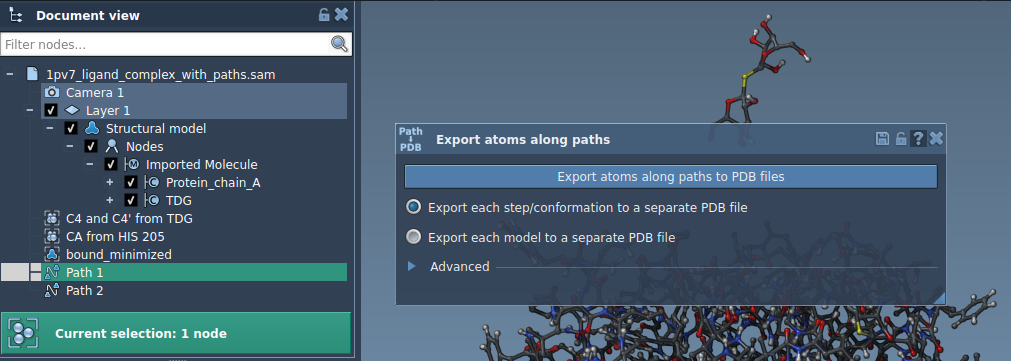
Step 3: Export a Subset of Atoms
Sometimes, you might only need the coordinates of specific atoms—such as a ligand or particular residues of a protein. To export a subset:
- Expand the Advanced panel in the app.
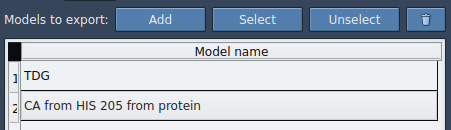
- Select your target atoms, for example, the ligand
TDG, from the Document view. - Click Add to define this group as a model for export. You can rename the model and even include multiple models in the export list.
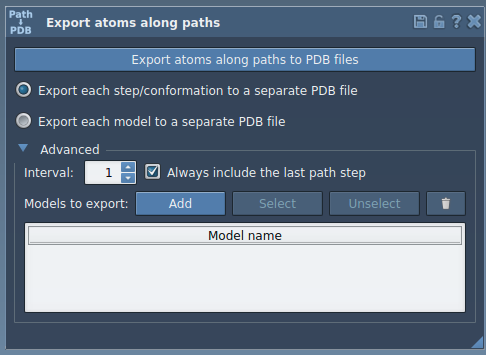
Once you’ve defined your model:
- Select the relevant path(s).
- Set the export format—single or multiple PDB files.
- Click Export atoms along paths to PDB files.
And that’s it—your subset of atom coordinates will now be exported precisely as needed.
Practical Applications
Some practical use cases of the Export Along Paths feature include:
- Generating reaction coordinate files for free energy profiling.
- Exporting ligand exit or entry trajectories to facilitate enhanced sampling.
- Tracking specific sets of atoms such as ligands, active sites, or protein backbones.
Wrapping Up
The Export Along Paths extension is a valuable tool for molecular modelers aiming to streamline trajectory analysis and exportation. By enabling users to work efficiently with both entire systems and specific subsets of atoms, it addresses a common challenge in molecular modeling workflows.
To learn more, visit the official SAMSON documentation page.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON here.





