If you are involved in molecular modeling or structural biology, understanding protein dynamics is key to your work. One critical motion that has captured global scientific attention is the opening and closing of the SARS-CoV-2 spike protein. This motion is not only fundamental to how the virus infects human cells but also a promising target for therapeutic and vaccine development. In this blog post, we’ll take a practical look at this motion and how SAMSON can help you explore and compute it.
Why the Spike Opening Motion Matters
The spike protein of SARS-CoV-2—responsible for the COVID-19 pandemic—plays a vital role in the virus’ ability to infect human cells. These spikes facilitate the virus’ entry by binding to a receptor called Angiotensin-Converting Enzyme 2 (ACE2) found on human cells. This binding leads to the viral RNA being inserted into the host cell. The exposed top region of the spike protein, which recognizes the ACE2 receptor, also becomes a critical target for neutralizing antibodies, making the spike’s dynamics central to vaccine and therapeutic design.
What Does the Motion Look Like?
Visualizing the spike’s transition between its closed (inactive) and open (ACE2-binding) states can provide incredible insight. Below, animations generated using SAMSON showcase these transitions:
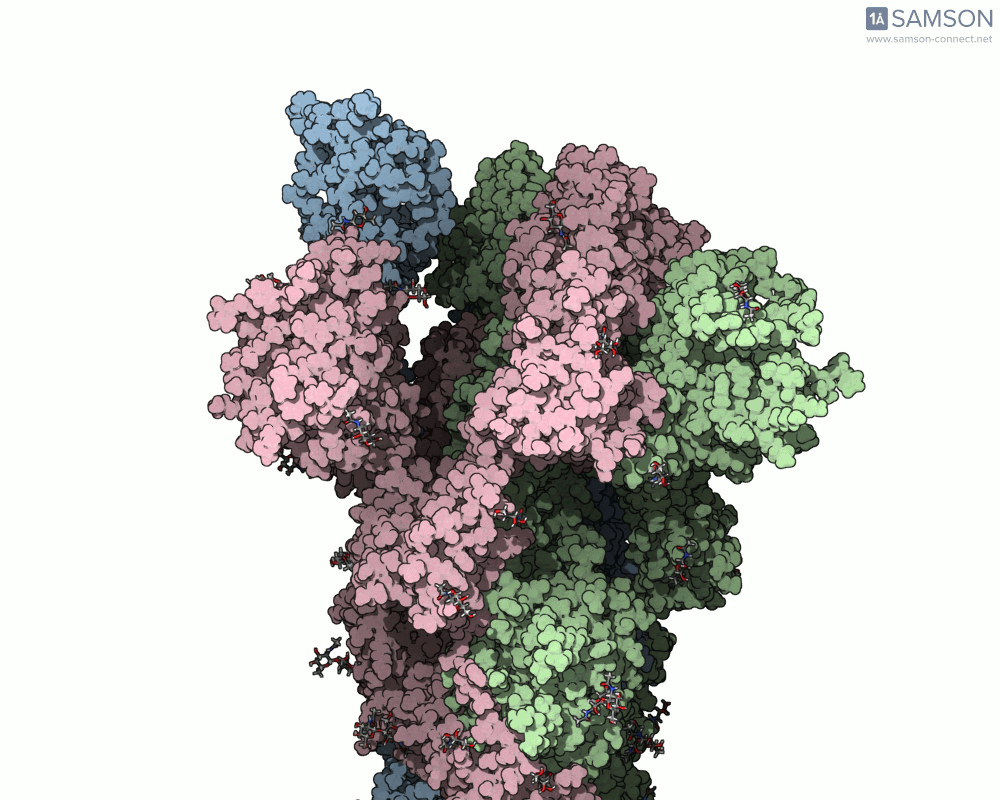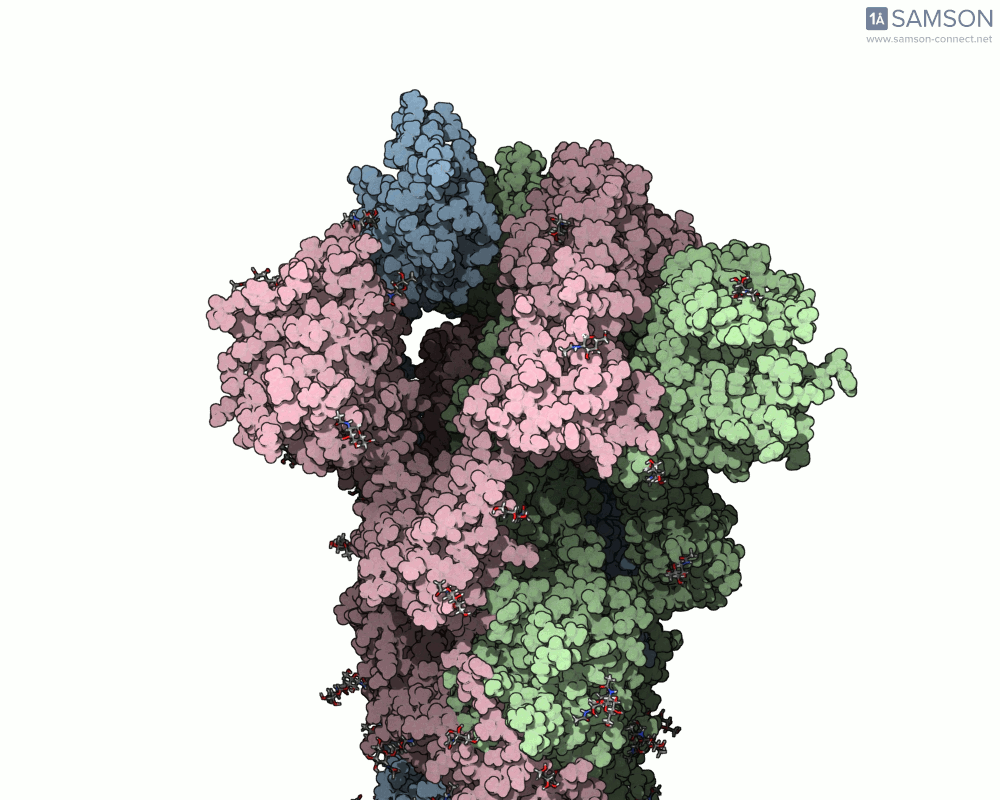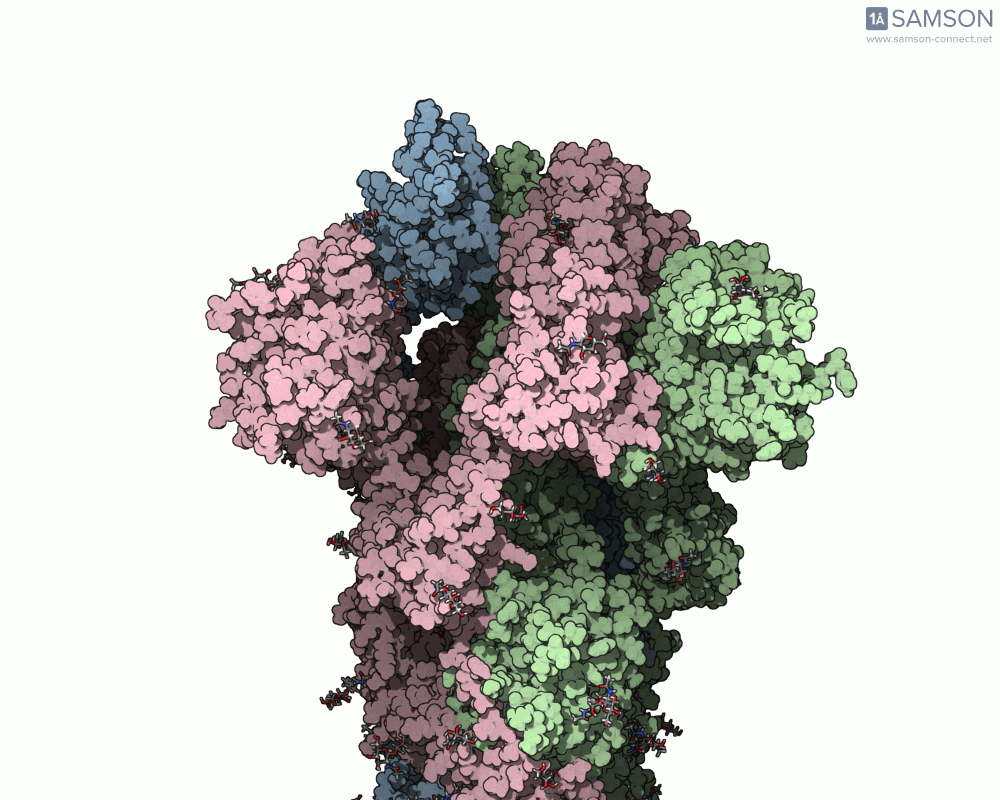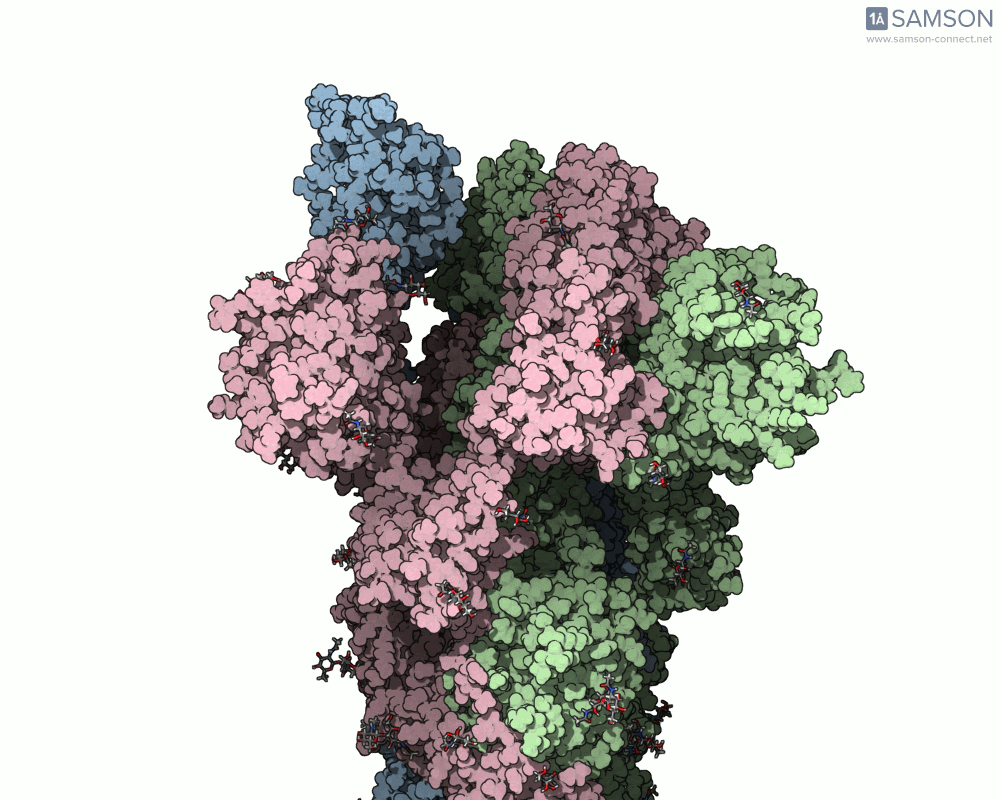
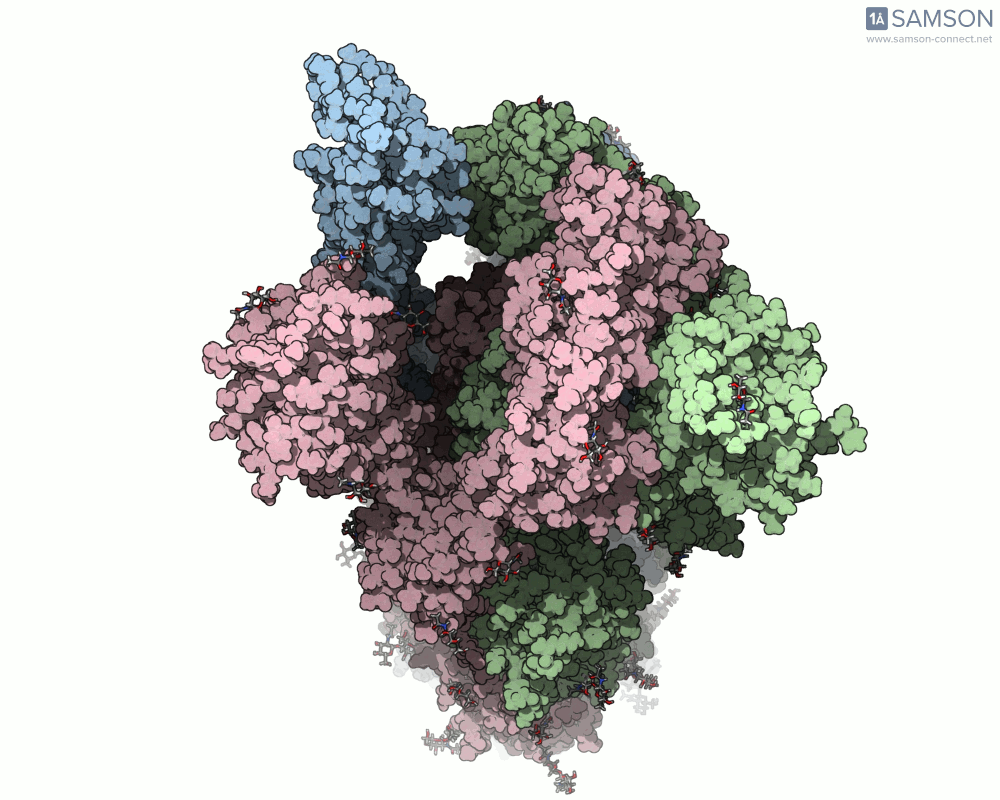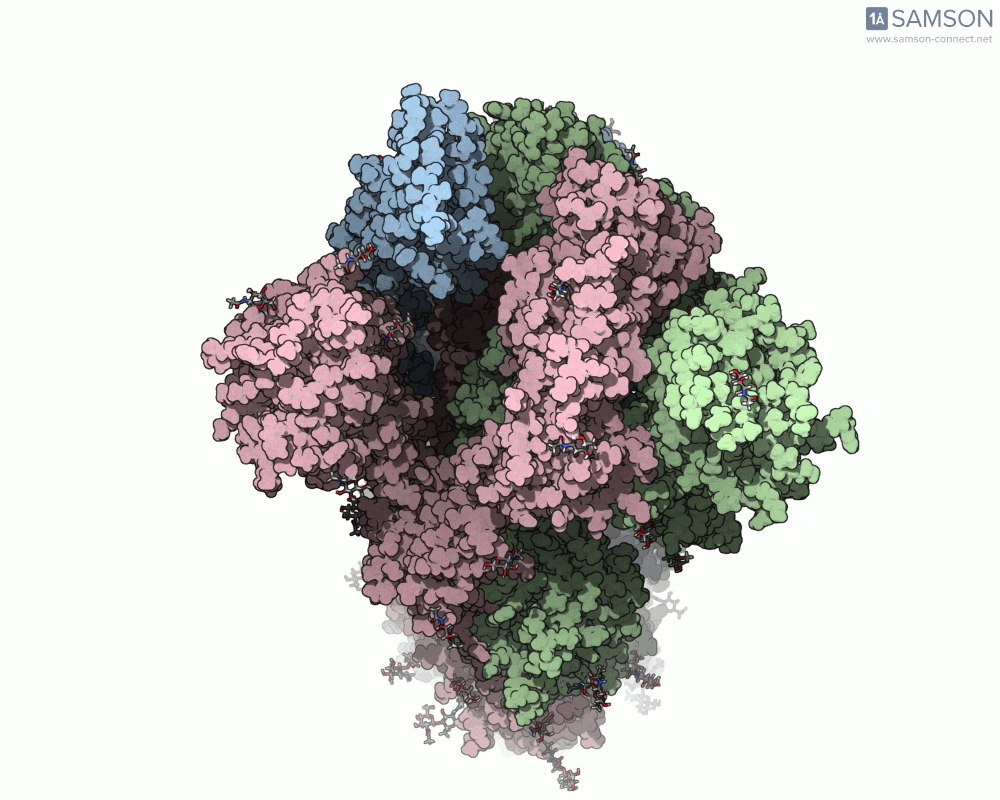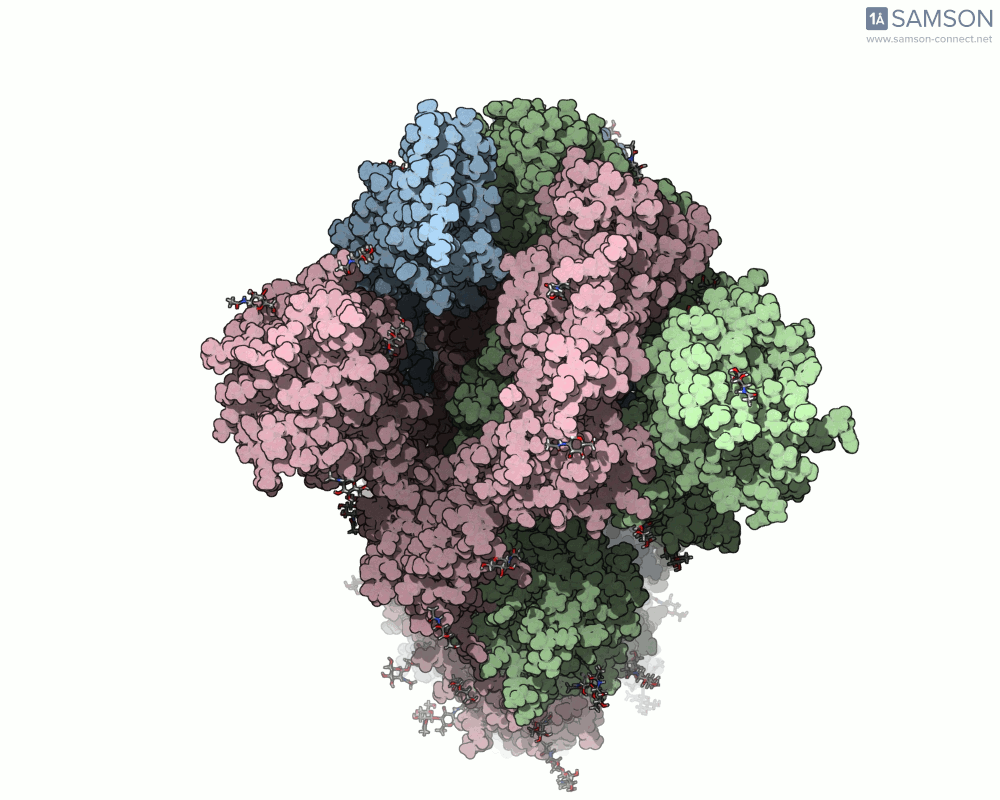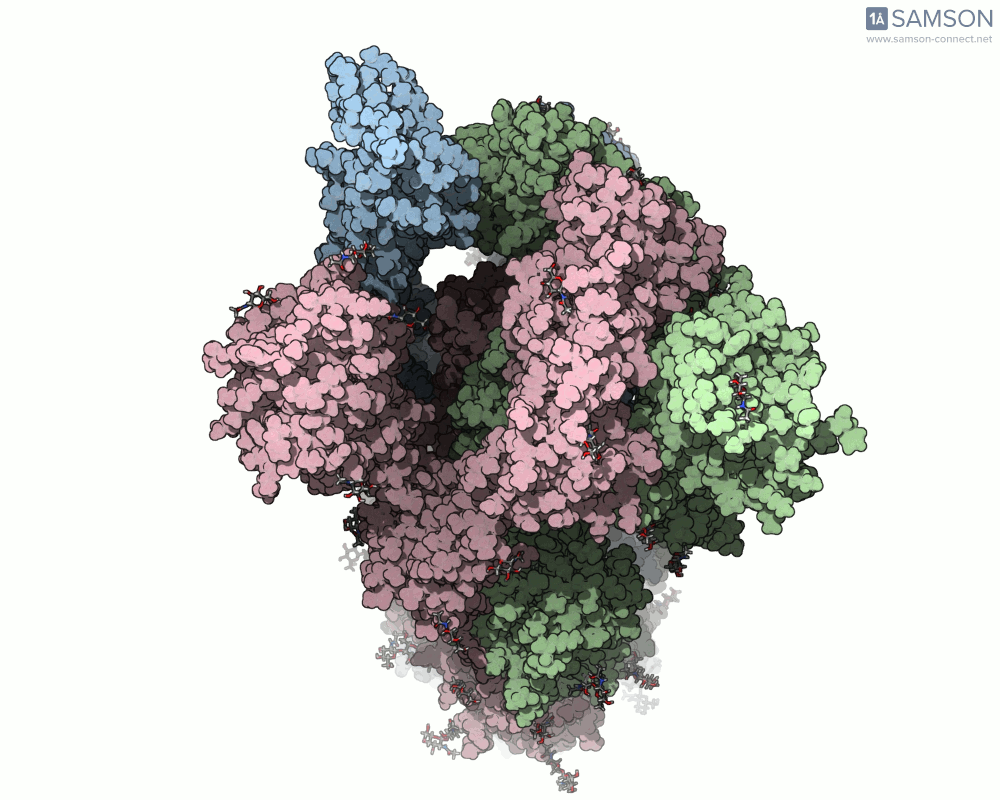
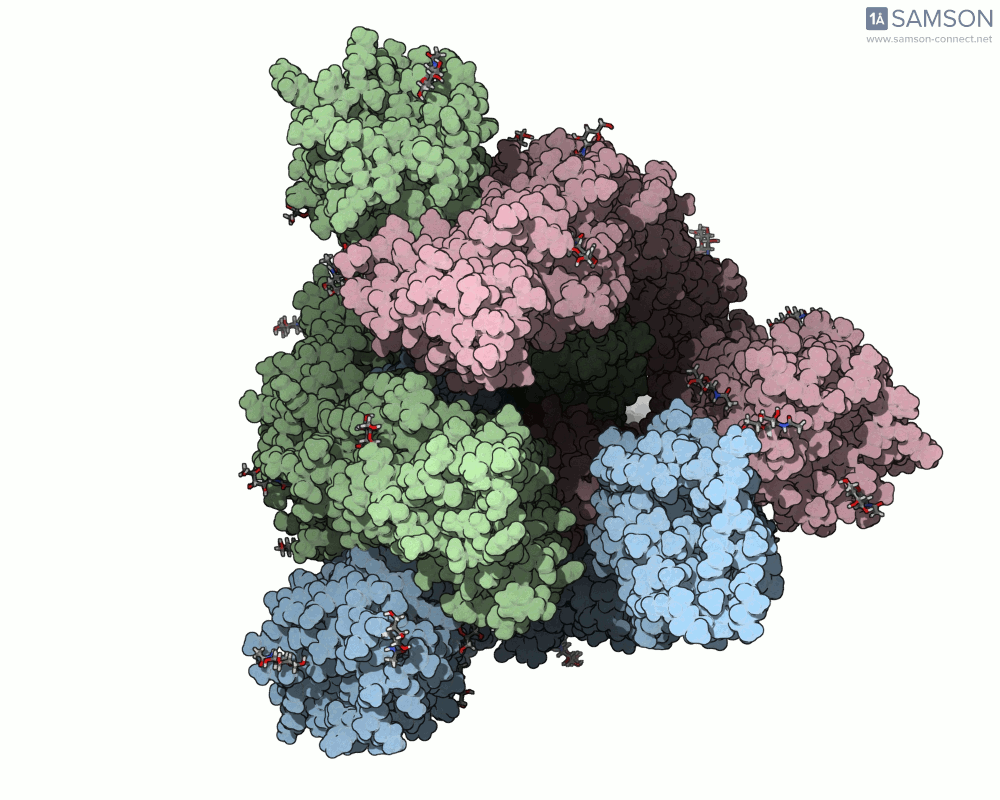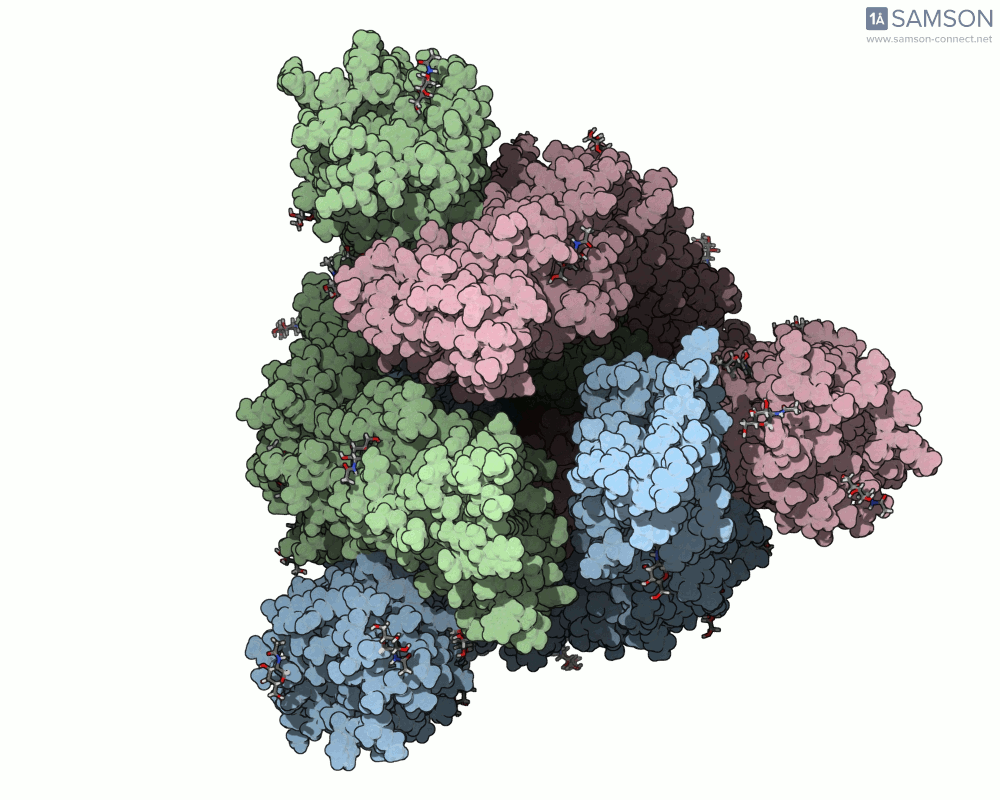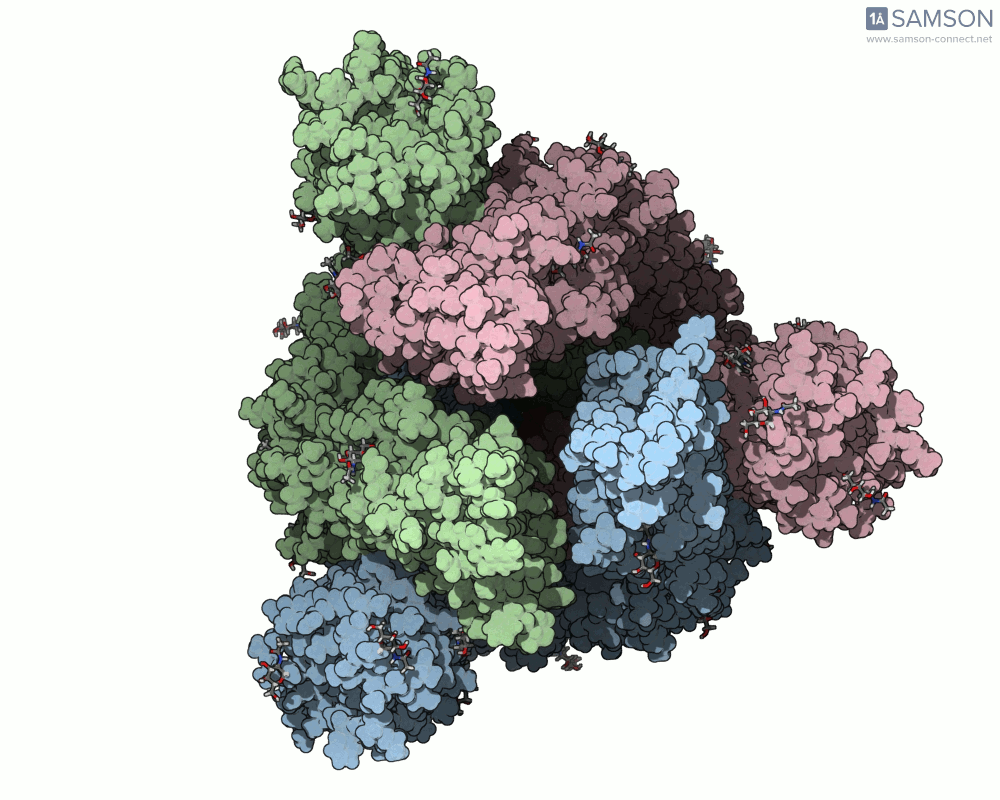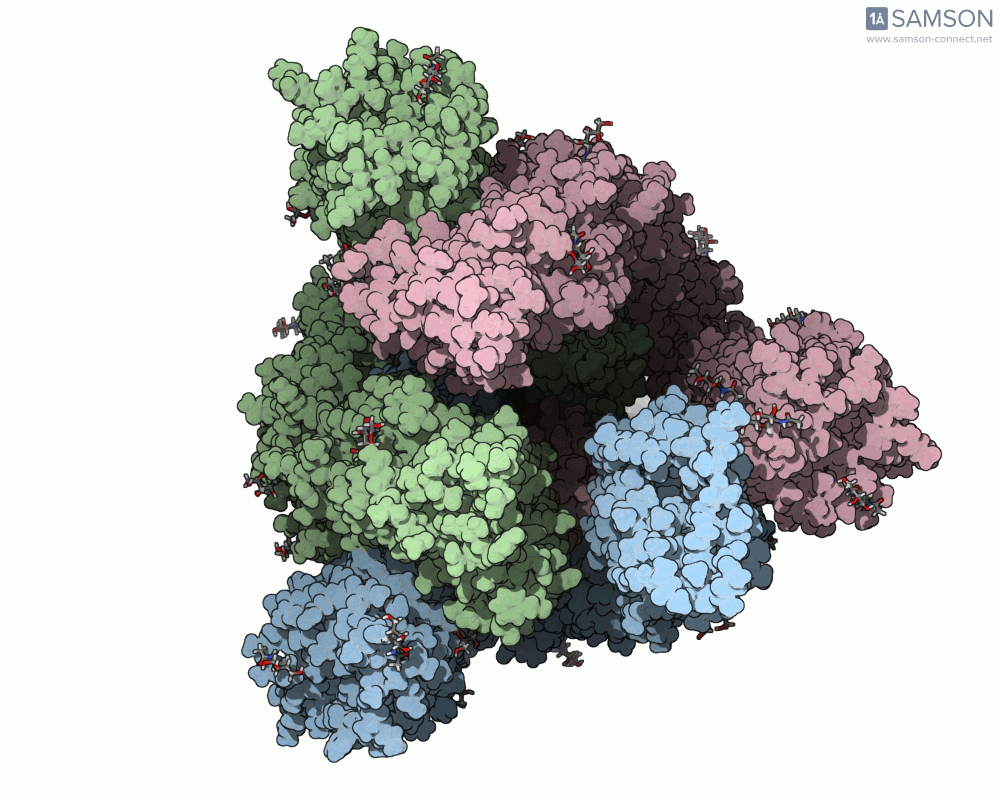
These animations highlight how the spike moves from a closed conformation to an open conformation where it can interact with ACE2. The open state is critical for the virus to bond with a host cell, leading to infection.
How Can You Study This?
SAMSON provides several computational tools to model and analyze protein motion, even for complex systems like the SARS-CoV-2 spike. You can explore the motion in detail by downloading the pre-calculated trajectory files available in different formats:
These files include the entire computed motion sequence, which you can visualize or leverage for simulations and further calculations.
How Does SAMSON Compute This Motion?
To compute the spike’s motion, SAMSON uses a precise pipeline involving two key modules: the ARAP (As-Rigid-As-Possible) Interpolation Path module and the P-NEB (Parallel Nudged Elastic Band) module. These modules allow you to:
- Generate an interpolated path between two known protein states, such as the spike’s open (PDB 6VYB) and closed (PDB 6VXX) conformations.
- Refine this path for improved accuracy using energy minimization techniques.
Using the ARAP and P-NEB modules, researchers can generate a high-quality trajectory capturing the motion from the closed to the open state. This process only requires minutes on a standard laptop, making it accessible and highly efficient.
The pipeline also includes pre-processing steps like modifying bond orders, adding hydrogens, and performing energy minimization, all seamlessly integrated into SAMSON’s environment.
Conclusion
Understanding the SARS-CoV-2 spike protein’s opening motion is crucial for anyone working in structural biology or molecular modeling. SAMSON provides powerful tools to visualize and compute these dynamics, enabling researchers to study key biological processes with greater precision.
To dive deeper into the workflow or download the trajectory files, visit the original documentation page.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON for free at https://www.samson-connect.net.





