When working with molecular simulations, selecting the most appropriate unit cell shape and dimensions is crucial for achieving efficient and accurate results. This blog post will guide you through the options available for space-filling unit cells supported by the GROMACS Wizard, helping you make informed decisions tailored to your molecular system’s needs.
What are unit cells and why do they matter?
In simulations involving periodic boundary conditions, unit cells represent the finite-volume building block replicated across space to create an infinite-like environment. Choosing an appropriate unit cell minimizes computational resources and ensures boundary effects don’t interfere with your system. The GROMACS Wizard supports a variety of shapes, each offering unique advantages.
Available shapes for unit cells
The GROMACS Wizard supports the following unit cell geometries:
- Cubic: A perfect cube, simple and often used for benchmark simulations.
- Orthorhombic: A rectangular box, useful for cases where the system extends further in one or more directions.
- Triclinic: A box with parallel or slanted faces, offering maximum flexibility for adjusting to specific geometries.
- Rhombic Dodecahedron: A shape closer to a sphere, highly efficient for simulating spherical macromolecules while saving solvent molecules and computational effort.
- Truncated Octahedron: Another spherical-like shape ensuring reduced computational overhead for globular molecules.
Each of these geometries is visually represented below to help you understand their differences:





How to choose the right unit cell?
The choice of a unit cell depends on the type of molecular system and simulation you are running:
- For approximately spherical systems like proteins in solution, consider a rhombic dodecahedron or truncated octahedron. These shapes require fewer solvent molecules compared to a cubic box due to their volume efficiency. For instance, a rhombic dodecahedron uses about 29% less volume than an equivalent cube, significantly reducing computational time.
- If your system has pronounced directional extensions, an orthorhombic or triclinic cell may fit better. These shapes offer more flexibility in adjusting box dimensions.
- For general-purpose simulations or where computational resources are not tightly constrained, a cubic cell remains a straightforward choice.
How to set the size?
Once you have selected a shape, it’s crucial to ensure that the unit cell is sized correctly:
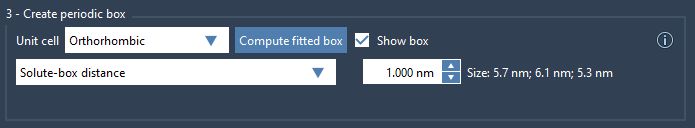
- Solute-box distance: Ensure a minimum of 1.0 nm between your solute (e.g., protein) and the box edges. This guarantees at least 2.0 nm between periodic copies of the solute, which satisfies the minimum image convention.
- Box lengths: Alternatively, you can adjust the box lengths directly for tighter control. For batch simulations, this ensures uniform box sizing across conformations or frames.
Final thoughts
Efficient molecular simulations start with careful unit cell selection and setup. Whether you are simulating a spherical protein or a large anisotropic molecule, tailoring the unit cell to your system’s needs can greatly enhance computational efficiency and accuracy. To dive deeper into the details, consult the official documentation page on periodic boundary conditions.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON by visiting this page.





