One of the central challenges in molecular modeling and simulation lies in handling dynamic and complex topologies. As molecular systems undergo significant structural changes—such as the breaking and reforming of covalent bonds—traditional interaction models often struggle to provide accuracy and user control during real-time simulations. This is where the Interactive Modeling Universal Force Field (IM-UFF) steps in to make a molecular modeler’s life significantly easier.
IM-UFF extends the Universal Force Field (UFF) by enabling smooth handling of topological changes while maintaining interactive control. This system allows for real-time edits and simulations, all while being guided by physically accurate inter-atomic forces. In this post, we’ll explore the key capabilities of IM-UFF and how you can use it to intuitively modify molecular structures and topologies with ease.
Breaking and Creating Bonds Dynamically
What sets IM-UFF apart is its ability to handle topological updates seamlessly. As you move atoms with your mouse, the model dynamically adjusts:
- Breaking bonds: When an atom is moved far enough from its neighbors, its bonds break automatically, updating the system’s topology.
- Forming new bonds: Conversely, when an atom is moved closer to other atoms, IM-UFF can dynamically form new bonds to create a new topology.
These smooth adjustments allow you to test different molecular configurations, like docking or bond rearrangements, without manually setting atom types and bond orders upfront.
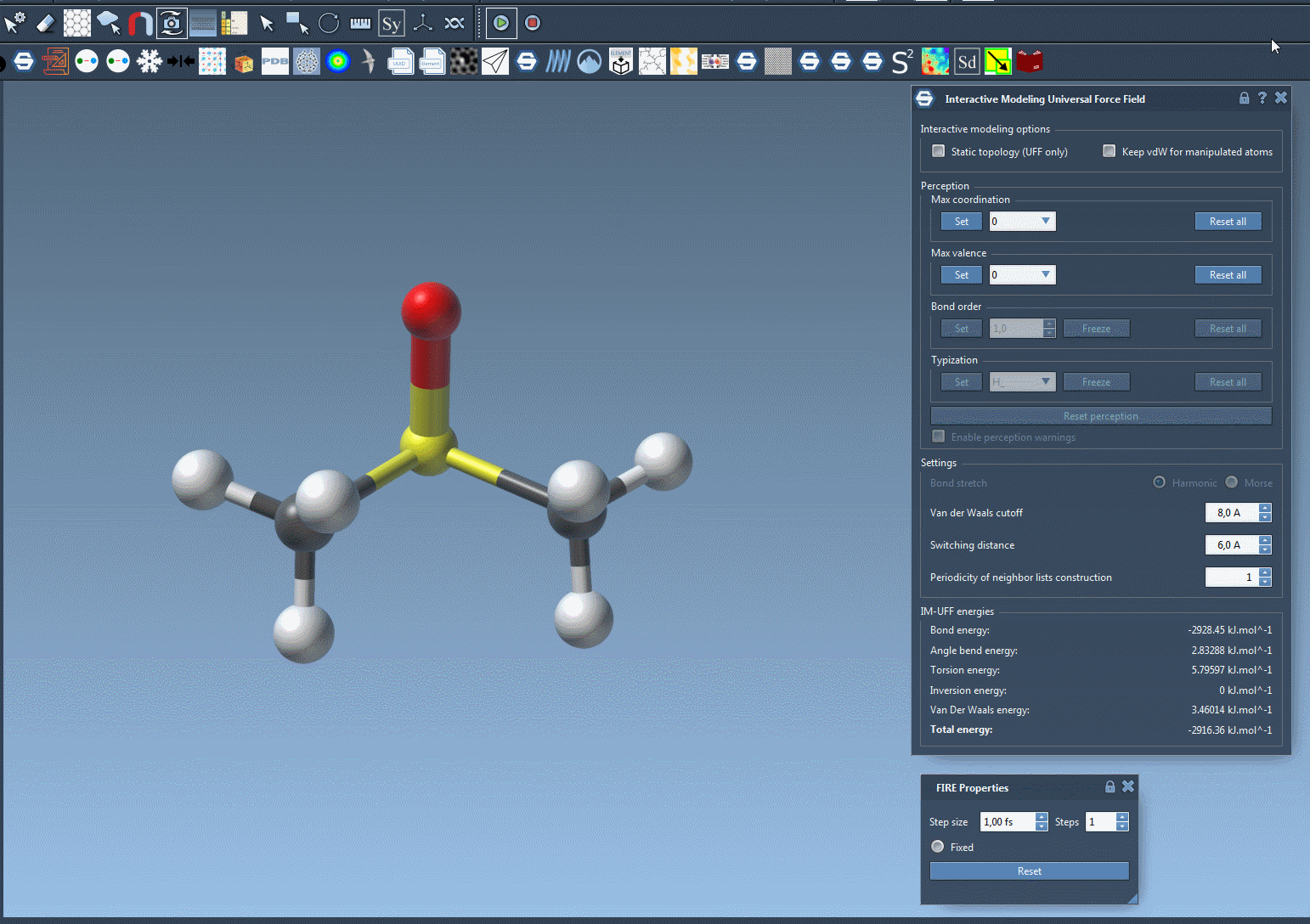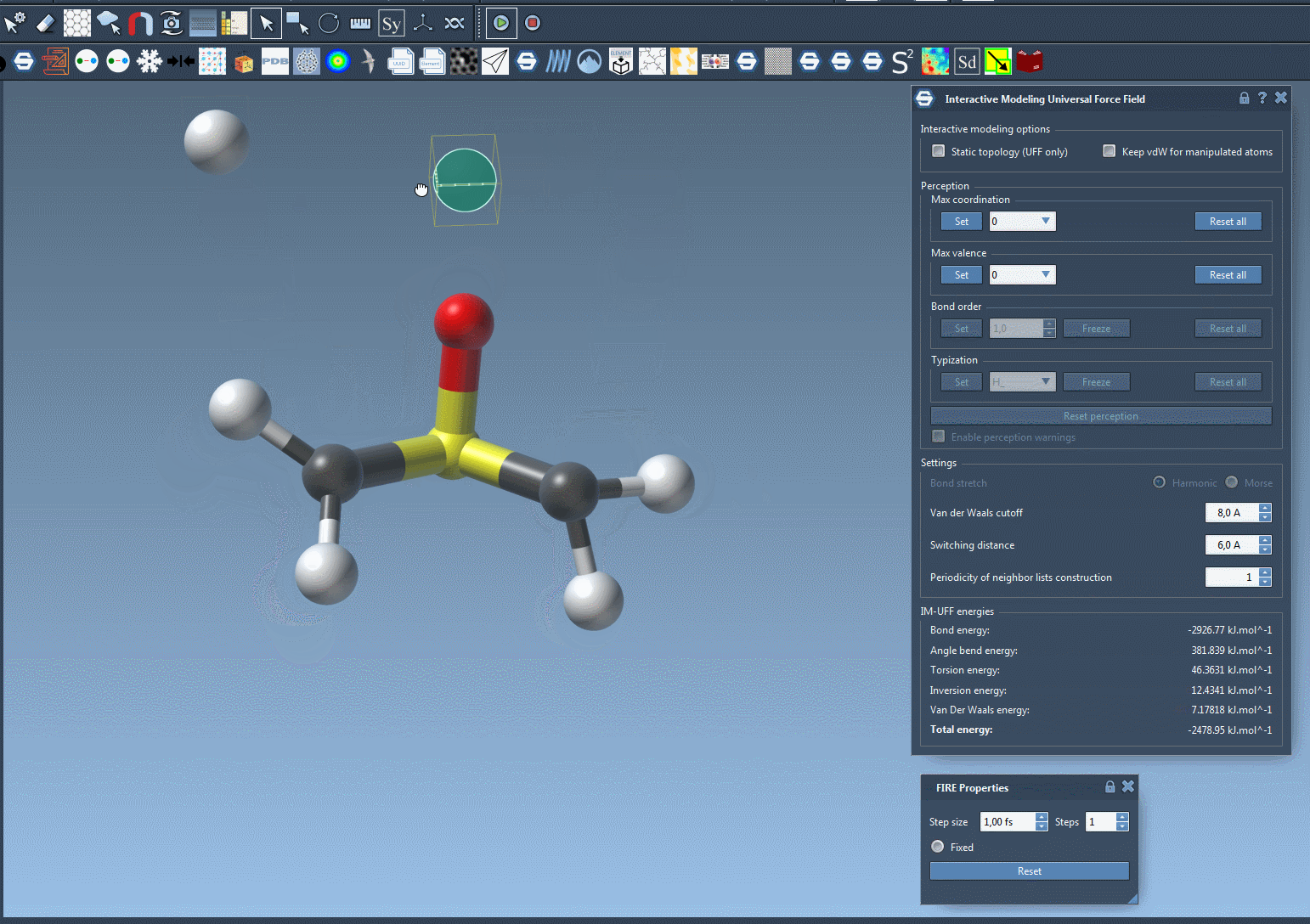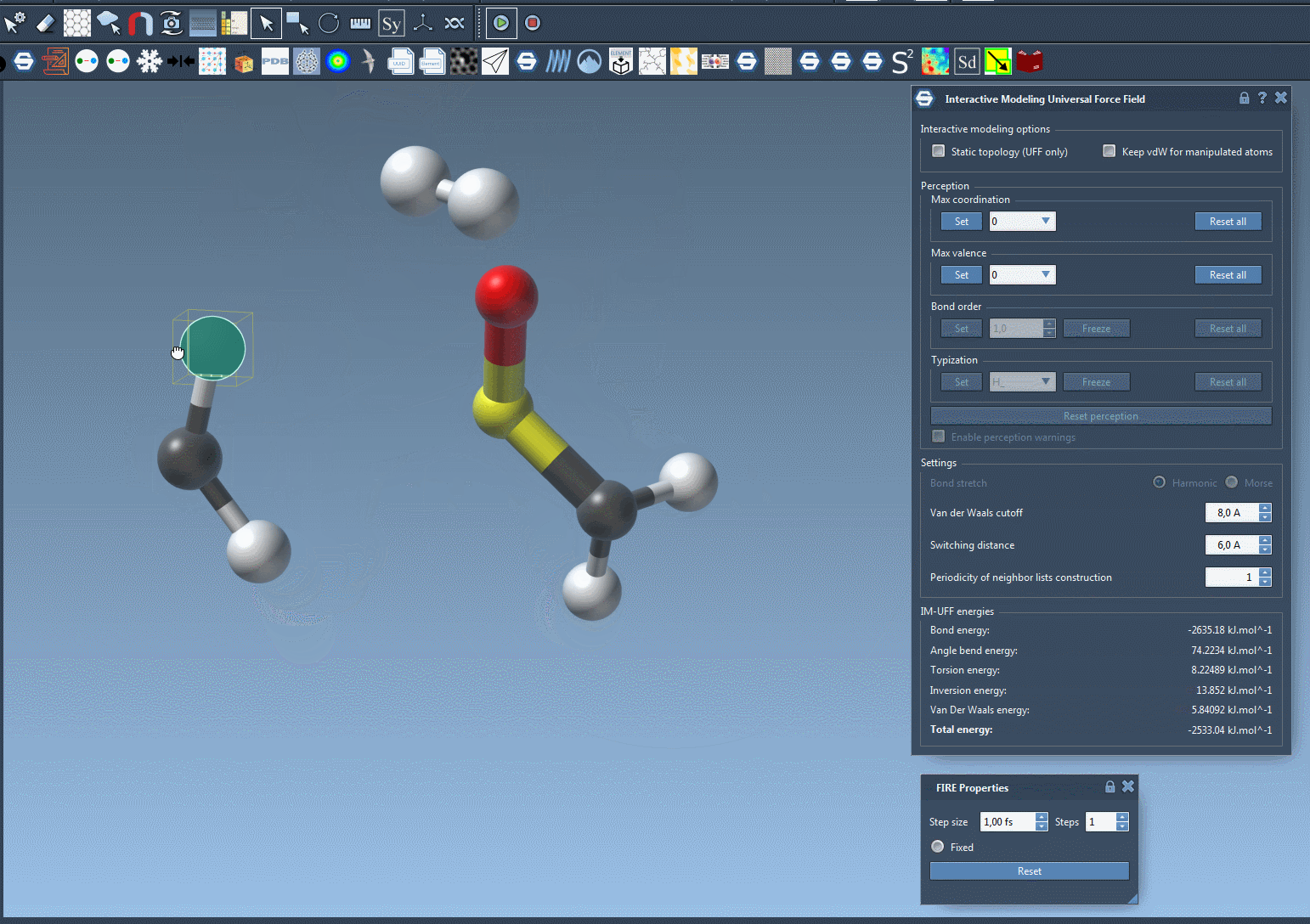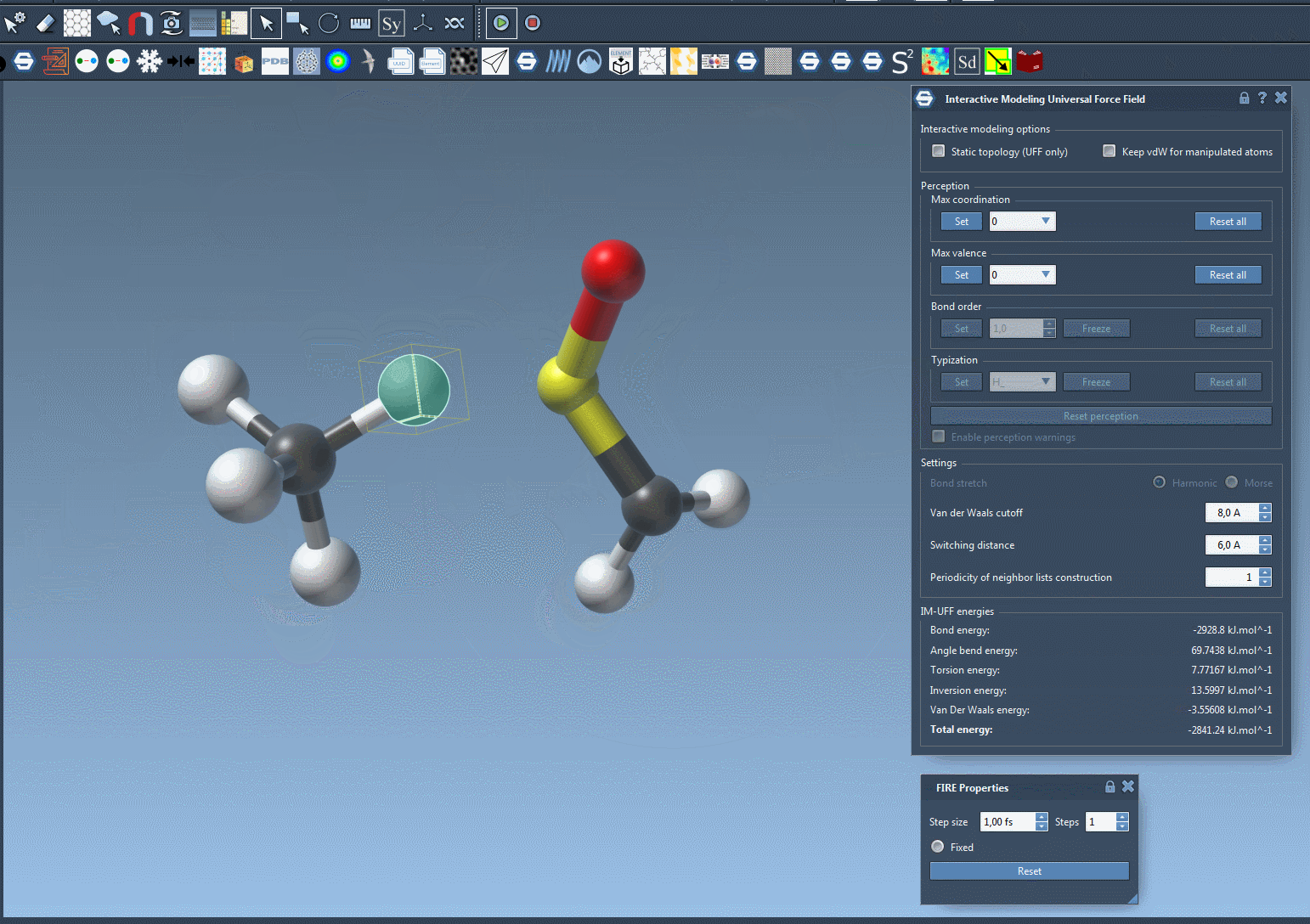
Interactive Modeling Features
IM-UFF includes two unique settings designed for maximum flexibility during interactive molecular modeling:
- Static topology (UFF only): This option lets users toggle between traditional static UFF simulations and interactive IM-UFF simulations. When enabled, the energy configurations in both models align, ensuring consistent results.
- Keep vdW for manipulated: This feature allows users to decide whether van der Waals (vdW) interactions are calculated for user-manipulated atoms. Disabling vdW for manipulated atoms makes molecular editing more intuitive, as repulsive vdW forces won’t hinder bond formations.
Such granular control during simulation makes IM-UFF an ideal tool for experiments requiring rapid prototyping of molecular configurations.
Customizable Parameters
IM-UFF is also fully customizable to suit specific simulation needs. You can configure:
- van der Waals cutoff and switching distance
- The periodic reconstruction of the neighbor list
- Coordination and valence limits during dynamic topology updates
These options are designed to give modelers precise control over their simulations, especially when working with dynamic molecular systems.
Why IM-UFF Matters
For molecular modelers working on interactive systems—such as creating novel molecular designs, simulating chemical reactions, or exploring docking scenarios—IM-UFF offers a more natural, flexible, and robust workflow. With its ability to seamlessly handle dynamic topology changes, IM-UFF bridges the gap between interactivity and physical accuracy in molecular modeling.
To dive deeper into the features and finer details of IM-UFF, check out the full IM-UFF tutorial.
SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON here.





