Exploring molecular analogs is a common challenge in molecular modeling. Whether you’re designing compounds for drug discovery, studying structure-activity relationships, or preparing structures for docking, generating analog series efficiently is essential. Fortunately, the SMILES Manager in SAMSON provides a straightforward method for performing positional analogue scanning with SMARTS patterns, saving you significant time and effort.
This step-by-step guide will walk you through using SMARTS codes to search for and replace chemical patterns in your molecules, enabling you to generate analog series and evaluate new substitutions.
Why SMARTS Patterns Matter
SMARTS (SMiles ARbitrary Target Specification) is a language used for specifying molecular substructures. By focusing on specific patterns, you can strategically substitute or attach groups to your molecule to better understand how those changes affect properties, binding affinities, and other characteristics. This approach is particularly valuable for researchers aiming to quickly test hypotheses about molecular modifications.
Using SMARTS to Define Search Patterns
To get started with positional analogue scanning in the SMILES Manager:
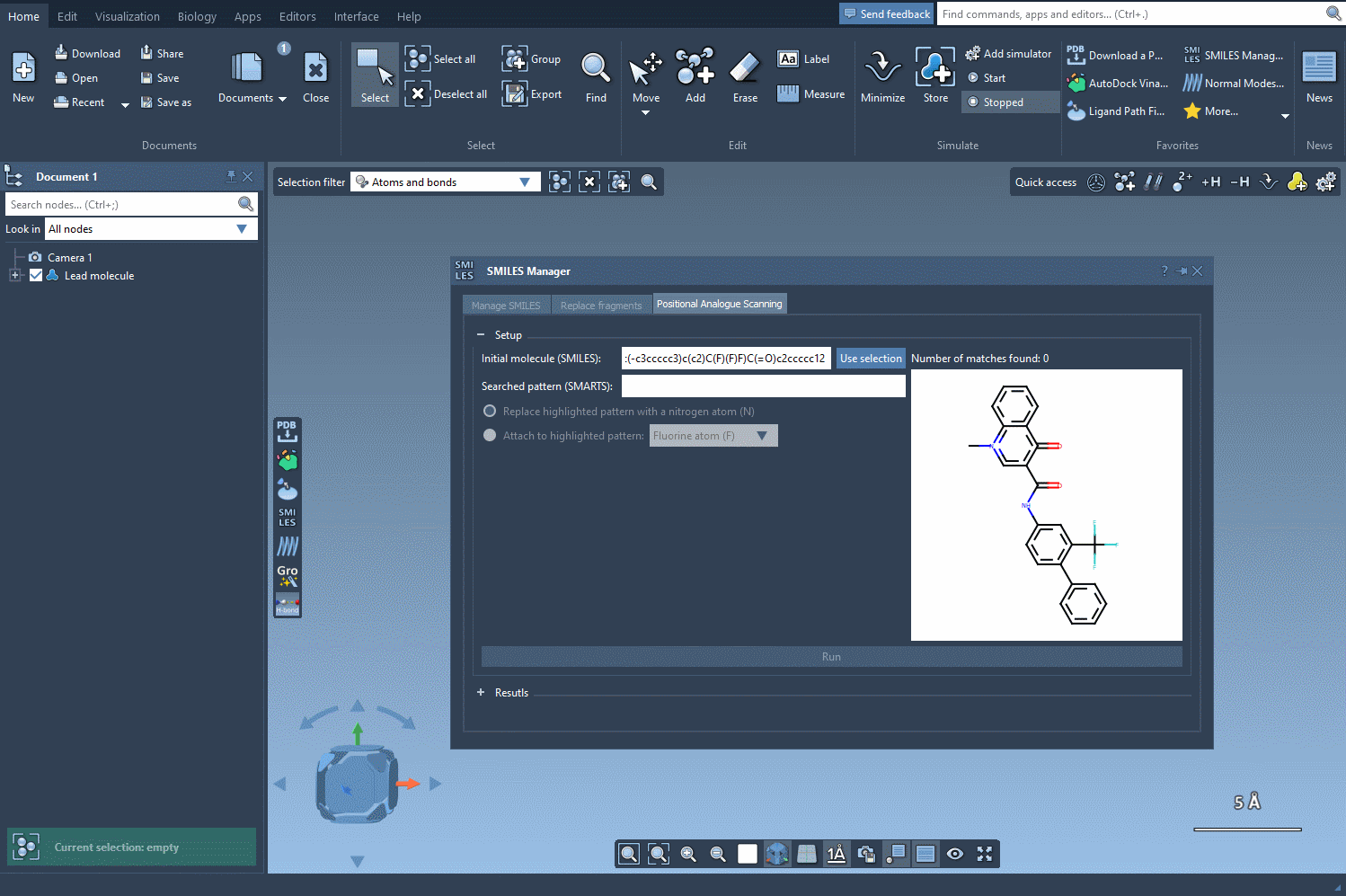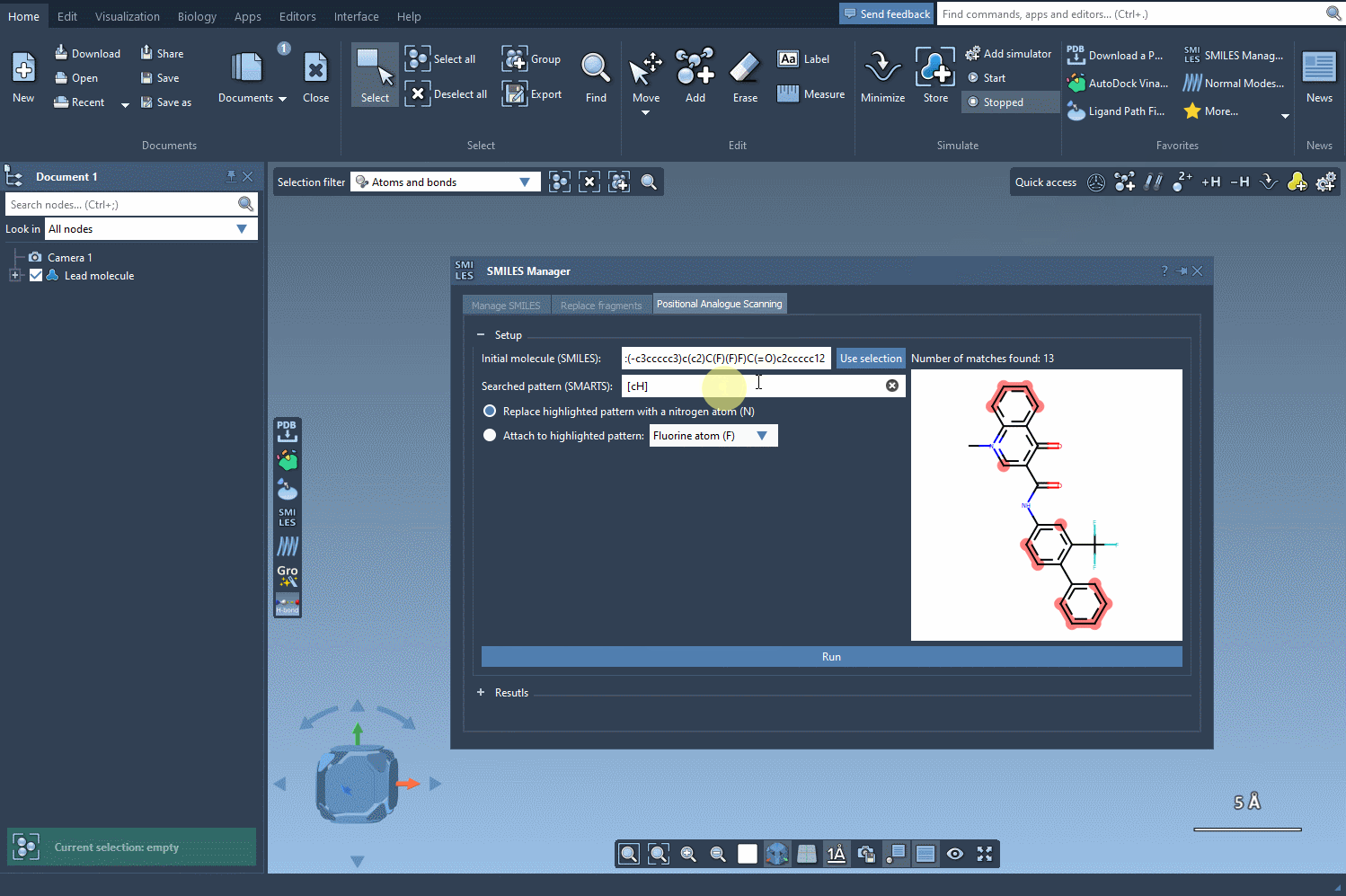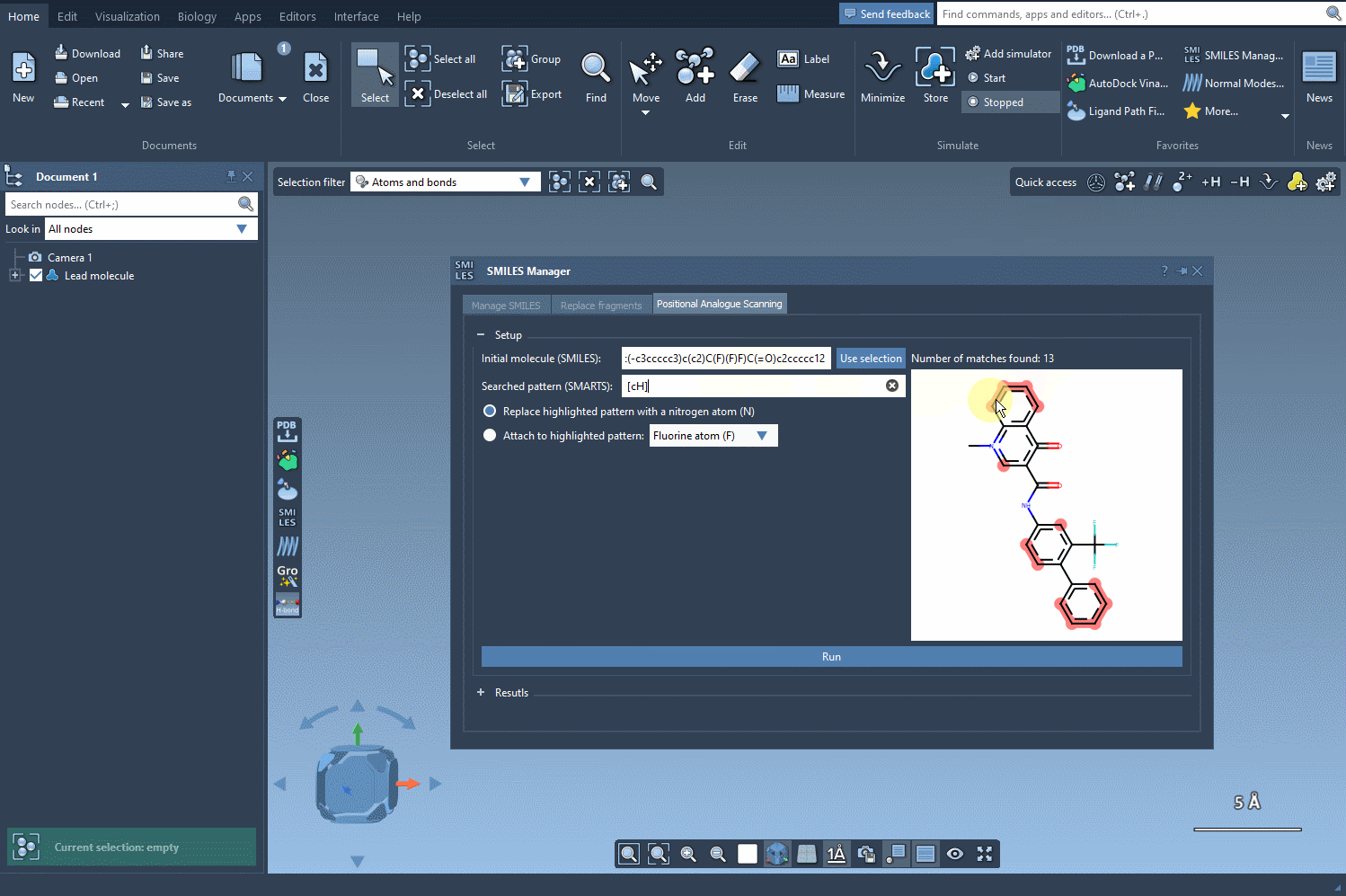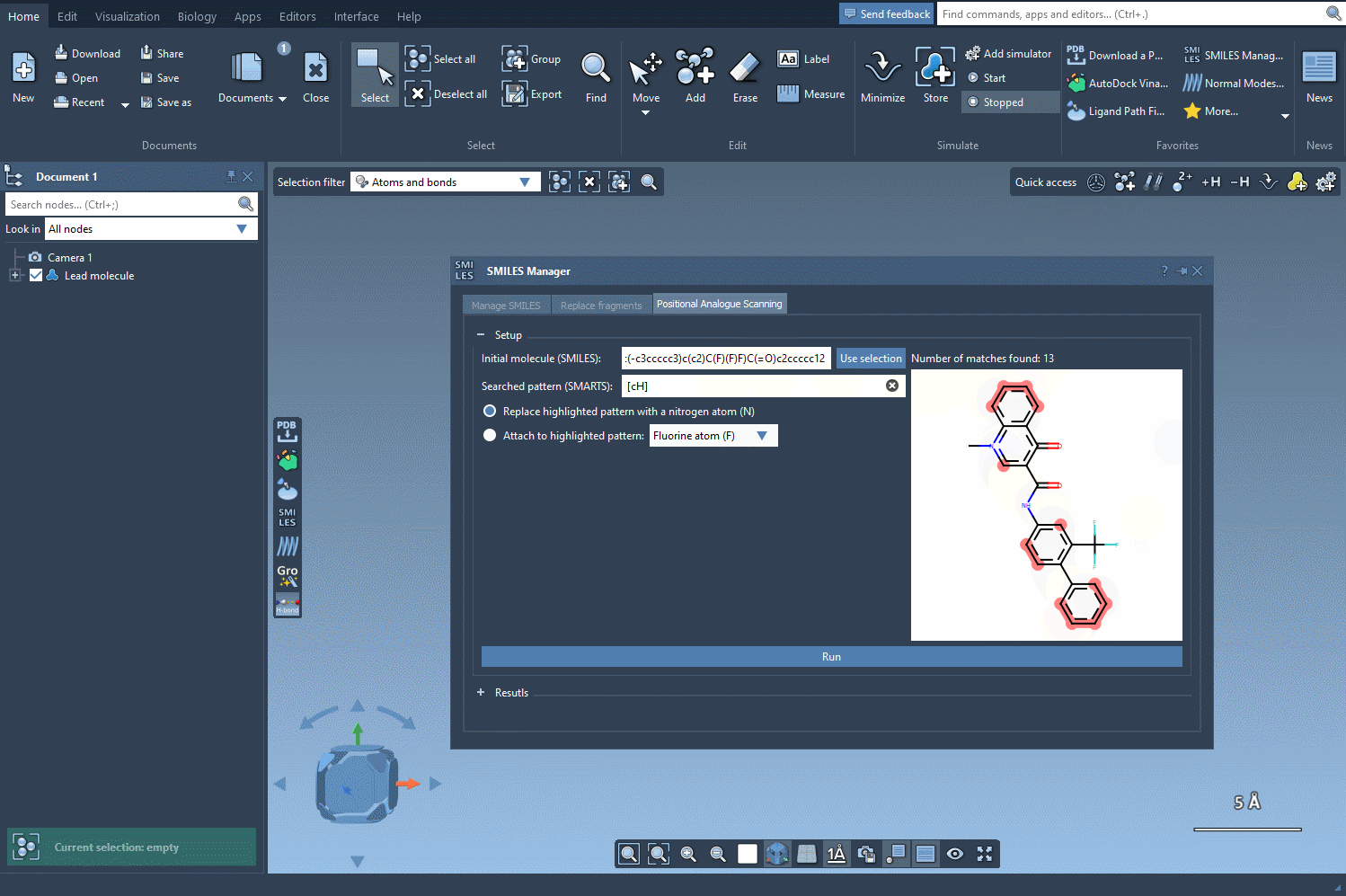
- Launch the SMILES Manager within SAMSON and switch to the Positional Analogue Scanning tab.
- Select a molecule to analyze either by entering its SMILES code directly or choosing it from your SAMSON document (using the Use selection button).
The next step involves defining the pattern within the molecule that you want to replace or modify. For instance, let’s say you want to identify all aromatic carbons in your molecule. You can use the SMARTS code [cH]. As shown in the following gif, the tool highlights the locations in your molecule matching the specified pattern:
Substituting or Attaching Groups
After identifying your desired patterns using SMARTS, it’s time to modify the molecule:
- You can replace the pattern (e.g., substitute aromatic carbons with nitrogen atoms).
- Alternatively, you can attach a functional group such as a fluorine atom (F) or a methyl group (CH3).
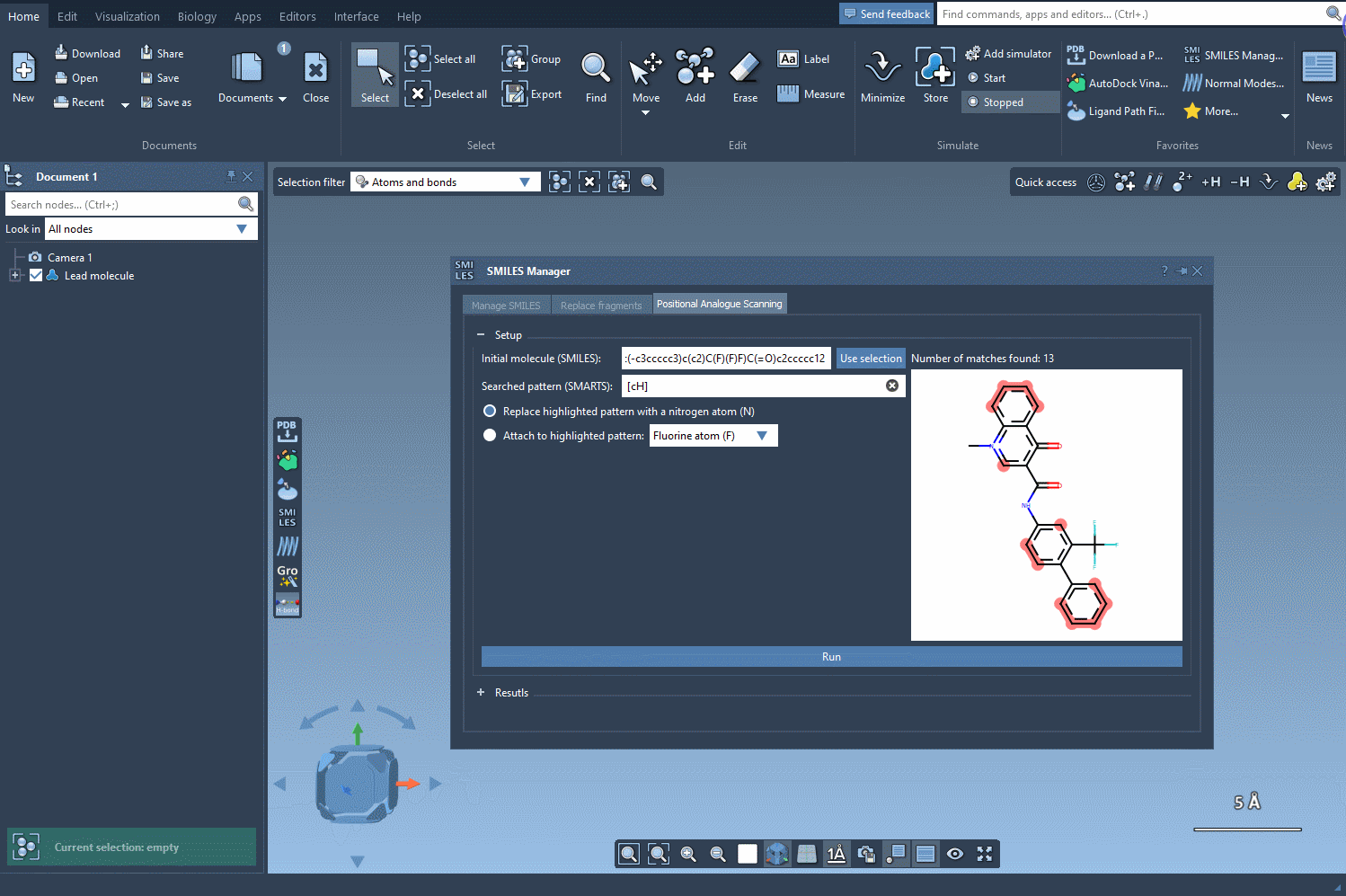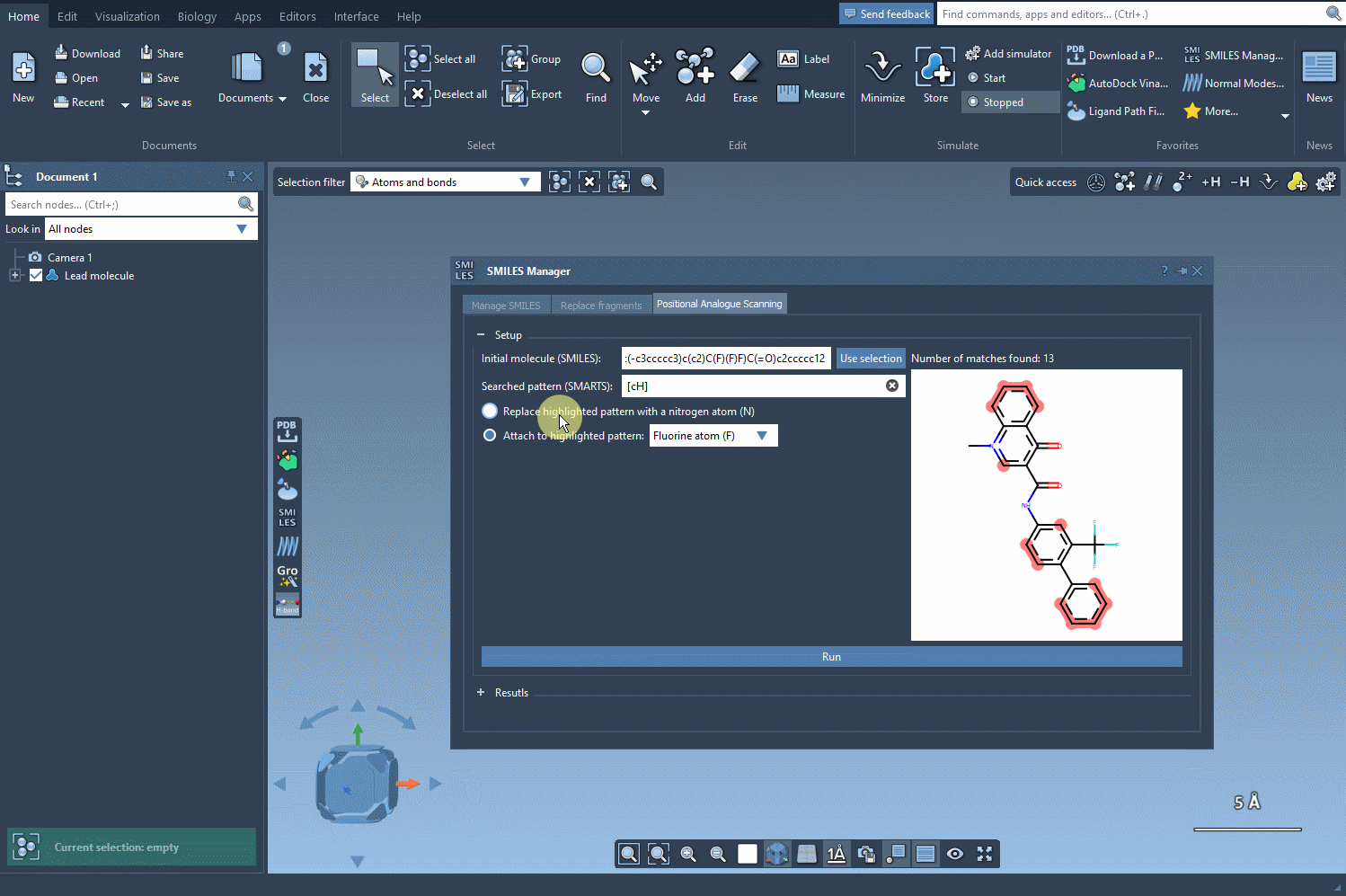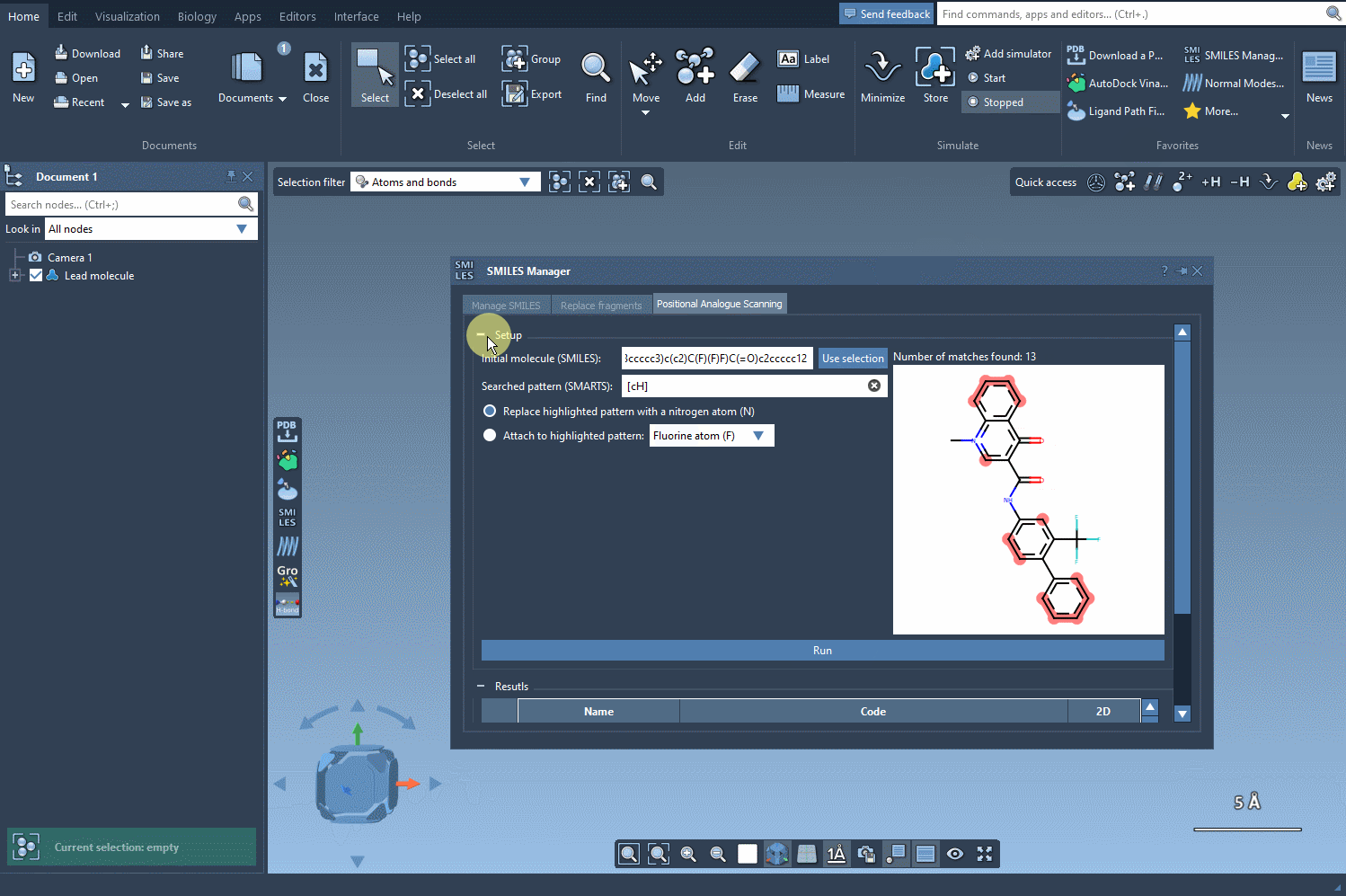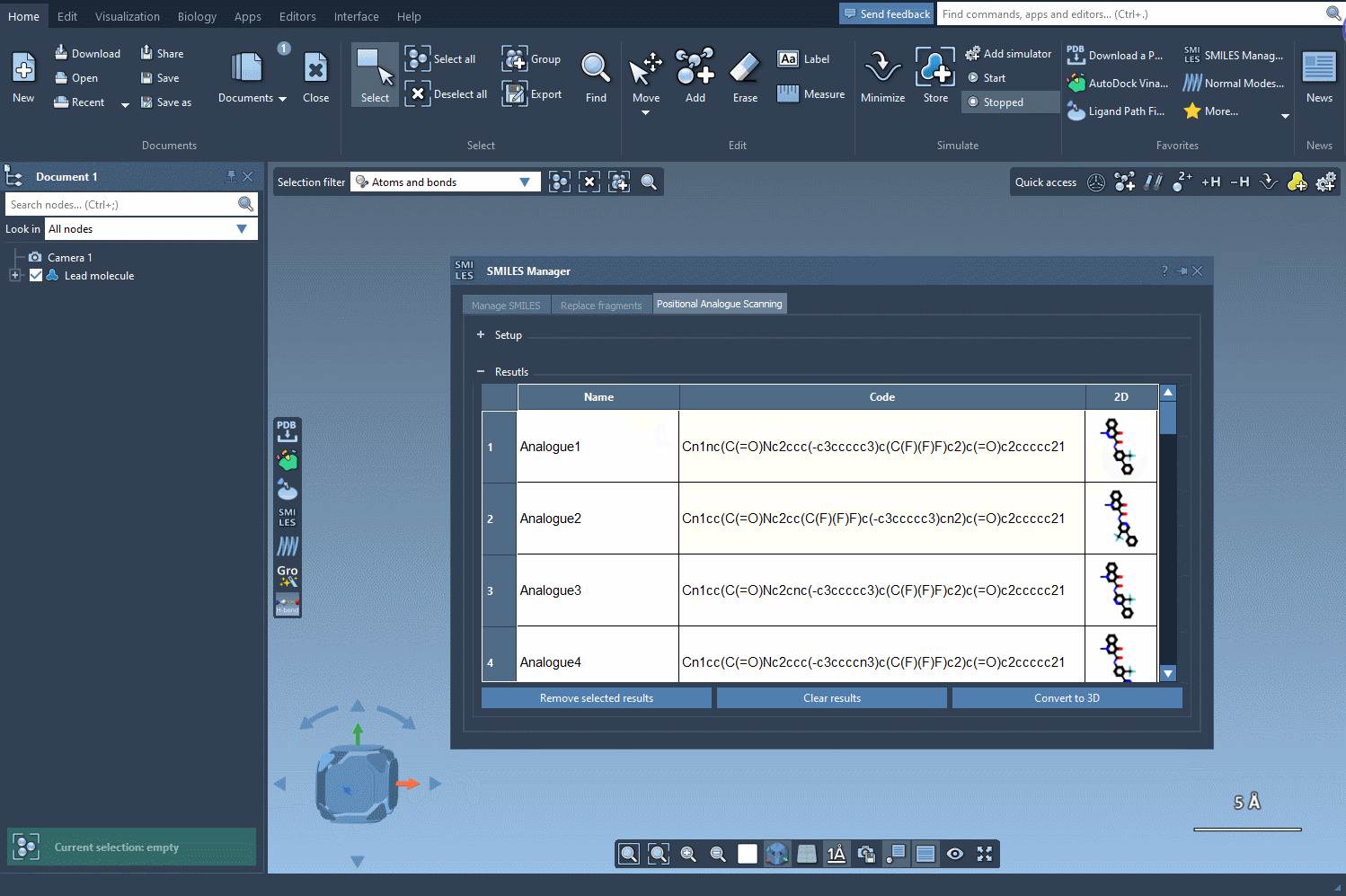
Simply choose your modification and click Run. The SMILES Manager swiftly generates a series of analogs along with their corresponding SMILES codes and 2D depictions. Here’s an example:
Taking It Further
Once you’ve generated your analogs, you can take additional steps to refine or evaluate them:
- Edit individual analogs by modifying their SMILES codes or adjusting their names directly in the results table.
- Visualize 2D representations in an expanded view or toggle their display in the results table.
- Generate 3D structures for deeper analyses, such as docking studies or interaction analyses.
This workflow not only simplifies the process of generating analog series but also provides immediate tools for screening your substitutions, making it highly effective when designing structures for downstream molecular modeling tasks.
Learn More
To dive deeper into the specifics of positional analogue scanning using SMARTS in SAMSON, please visit the official documentation page here.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can download SAMSON at https://www.samson-connect.net.





