Molecular modelers often face the challenge of effectively visualizing and analyzing the motion of molecules during simulations, particularly when it comes to tracking center-of-mass (COM) trajectories. Whether it’s studying ligand unbinding pathways, protein conformational changes, or atomic motion across simulations, having an intuitive way to observe and analyze such behavior is essential. This blog post introduces a solution: the Pathlines visual model in SAMSON, a powerful tool designed to help you do precisely this.
Why Use Pathlines?
Pathlines visual models offer unique capabilities that address common challenges faced during molecular simulations:
- Track motion: Keep an eye on ligands, residues, or entire groups of atoms as they move along a specific path.
- Visualize center-of-mass displacement: Easily see how the COM of atom groups shifts within complex systems.
- Analyze key behaviors: Dive into phenomena like ligand unbinding, domain motions in proteins, or diffusion processes.
Getting Started with Pathlines
To begin using Pathlines, make sure you’ve installed the SAMSON Extension. If not, simply head to the Pathlines Extension page, click Add, and restart SAMSON to activate it. Once ready, here’s how you can visualize COM motion in a few quick steps:
Step 1: Load the Sample System
For practice, download a sample system provided by SAMSON, which contains the structural model of Lactose permease (1PV7) with its ligand Thiodigalactosid (TDG) and pre-generated unbinding paths. Use the document link here to access the system. Below is a quick preview of the setup:

Step 2: Select Atoms and Paths
For Pathlines, you’ll need to select atoms and paths to visualize their motion. Open the Document view, select the group of atoms whose motion interests you, and select one or more paths. You can use the Ctrl/Cmd key to make multiple selections. If you don’t select any specific atoms or paths, SAMSON will automatically use the entire system or all available paths.

Step 3: Create a Pathline Visual Model
Once you’ve selected your atoms and paths, create the visual model:
- Go to Visualization > Visual model > More… (or use the shortcut Ctrl/Cmd + Shift + V).
- Select Pathline of the center of mass in the dialog box and click OK.
This will generate a visual representation of your selected COM motion. Here’s an example of what it might look like:
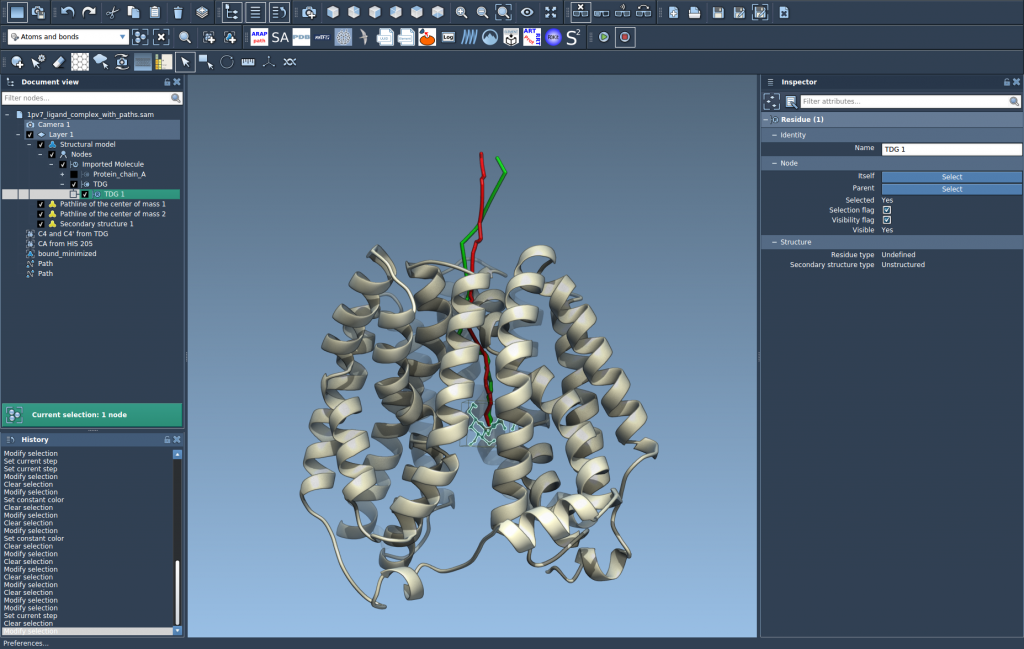
Step 4: Explore and Customize
Pathlines can be tailored to suit your analysis needs:
- Explore the paths: Double-click a path in the Document view to start or stop it. Alternatively, access options via the path’s context menu.
- Inspect and adjust properties: Use the Inspector (Ctrl/Cmd + 2) to modify attributes like color and thickness of pathlines.
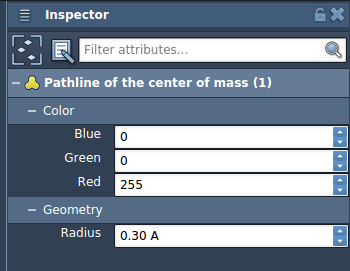
Applications of Pathline Visualization
Pathlines aren’t just versatile; they’re indispensable for various use cases:
- Understand and display ligand unbinding and rebinding routes.
- Analyze protein domain motions in complexes.
- Observe collective center-of-mass movements in simulations.
Next Steps
With your pathlines set up, you can further optimize paths using tools like SAMSON’s P-NEB Extension. To explore in greater detail, check out the related documentation page.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Get SAMSON here.





