Author: OneAngstrom
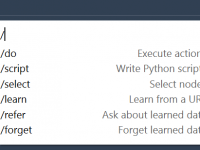
Quickly Select Charged or Polar Residues Using NSL in SAMSON
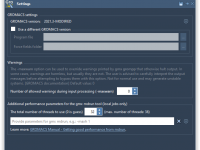
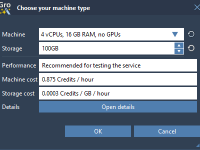
Making the Most of Idle Time: Run GROMACS Simulations in the Cloud
How to Quickly Select Molecular Paths with a Specific Number of Atoms in SAMSON
Avoid Chain Conflicts When Generating CG Models with Martinize2
When working with coarse-grained (CG) molecular models, it’s common to simulate multiple copies—or replicas—of the same protein in a single system. This setup is especially useful for studies involving protein aggregation, diffusion, or encapsulation. However, many researchers new to coarse-graining…