Author: OneAngstrom
From SMILES to 3D in Seconds: Generating 3D Structures of Molecular Analogs in SAMSON
Avoid Losing Atom Paths in Animations: A Quick Guide to Recording Trajectories in SAMSON
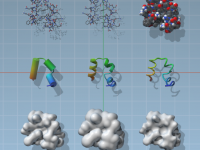
Seeing Molecules Clearly: A Guide to Visual Models in SAMSON
Animating Atomic Motion Without Scripts or Code
One of the common challenges molecular modelers face is effectively communicating molecular transformations over time. Whether you’re preparing educational content, designing nanosystems, or capturing animations for scientific presentations, showing how atoms or molecules change position step-by-step usually requires external video…