Saving Time in Molecular Modeling with Multiple Camera Views
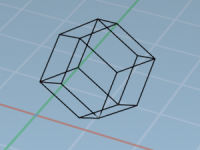
Switching between different molecular orientations and projections is a common—and sometimes frustrating—task in molecular modeling. Whether you’re aligning molecules, creating visuals, or inspecting specific structural features like active sites or crystal lattices, constantly rotating, zooming, and panning your model can…