Category: Uncategorized
Why Labels Disappear (and How to Control When They Reappear)
Quickly Spot and Fix Unfavorable Residues with an Interactive Ramachandran Plot
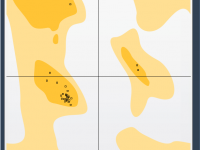
Protein modelers frequently run into a recurring challenge: identifying and correcting residues with unfavorable backbone conformations. These problematic residues can undermine the structural integrity of simulations, reduce model accuracy, and even produce misleading results in docking or dynamics. Fortunately, SAMSON’s…
Making Your Molecular Path Walk Backwards: A Quick Guide to Reverse Path Playback
Get Immediate Feedback with Interactive Molecular Simulations in SAMSON
Visualizing Subtle Molecular Motions: When to Use Rock Instead of Rotate
Preventing Simulation Setup Errors with Auto Input in GROMACS Wizard
Ligand unbinding paths: why the sampling box matters more than you think
When simulating ligand unbinding pathways in protein-ligand complexes, one surprisingly common source of difficulty isn’t always the algorithm or even the protein structure itself—it’s defining the sampling region. Many molecular modelers spend hours tuning energy parameters, refining force fields, and…
A Better Way to Animate Molecular Presentations in SAMSON
Clearer Scientific Storytelling: Making Molecular Models Appear on Cue
Why Labels Disappear (and How to Control When They Reappear)
Quickly Spot and Fix Unfavorable Residues with an Interactive Ramachandran Plot

Protein modelers frequently run into a recurring challenge: identifying and correcting residues with unfavorable backbone conformations. These problematic residues can undermine the structural integrity of simulations, reduce model accuracy, and even produce misleading results in docking or dynamics. Fortunately, SAMSON’s…
Making Your Molecular Path Walk Backwards: A Quick Guide to Reverse Path Playback
Get Immediate Feedback with Interactive Molecular Simulations in SAMSON
Visualizing Subtle Molecular Motions: When to Use Rock Instead of Rotate
Preventing Simulation Setup Errors with Auto Input in GROMACS Wizard
Ligand unbinding paths: why the sampling box matters more than you think

When simulating ligand unbinding pathways in protein-ligand complexes, one surprisingly common source of difficulty isn’t always the algorithm or even the protein structure itself—it’s defining the sampling region. Many molecular modelers spend hours tuning energy parameters, refining force fields, and…