For molecular modelers, one of the most common challenges is turning chemical information encoded in SMILES (Simplified Molecular Input Line Entry System) strings into 3D molecular structures. These structures are essential for simulations, visualizations, and property predictions. But how do you streamline this process while minimizing manual efforts? Enter the SMILES Manager extension for SAMSON, a tool designed to simplify the generation of 3D molecular structures.
The SMILES Manager is an intuitive tool based on the widely-used RDKit cheminformatics library. One of its standout features is the ability to transform a list of SMILES strings into precise 3D representations, directly integrated into SAMSON. Below, we’ll guide you through how this feature works, how it solves a key pain point, and why it can elevate your workflow.
What’s the Pain Point?
Molecular modelers need accurate 3D molecular models for calculations, docking simulations, and structure-activity relationship studies. While SMILES strings provide a compact way to describe molecules, converting these into 3D structures can be tedious and error-prone, especially when multiple molecules or complex SMILES strings are involved. Many tools require external file conversions and extensive post-processing, leading to inefficiencies.
How the SMILES Manager Helps
The SMILES Manager makes 3D structure generation quick and hassle-free. Here’s how you can use it to generate 3D structures from SMILES codes:
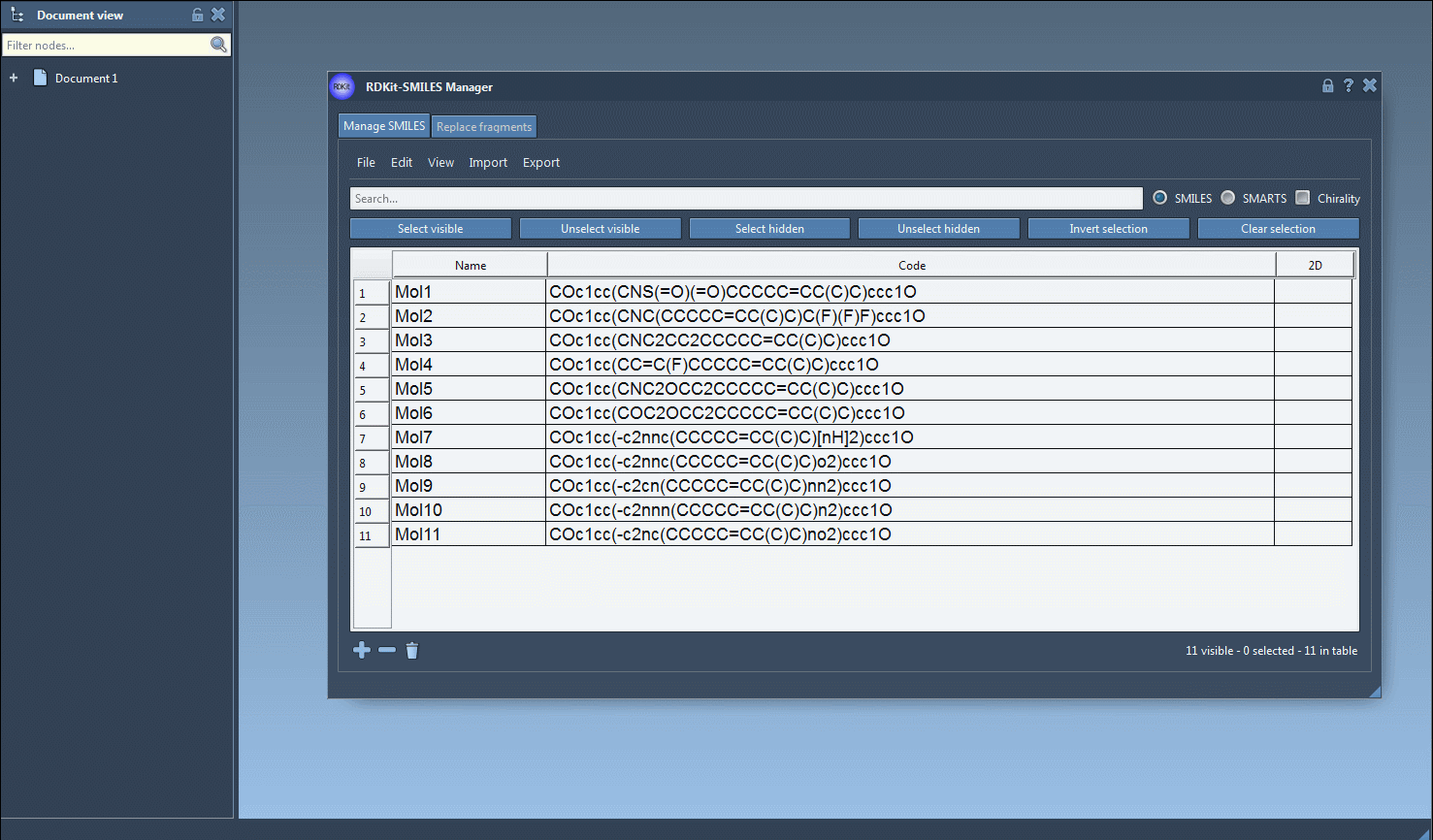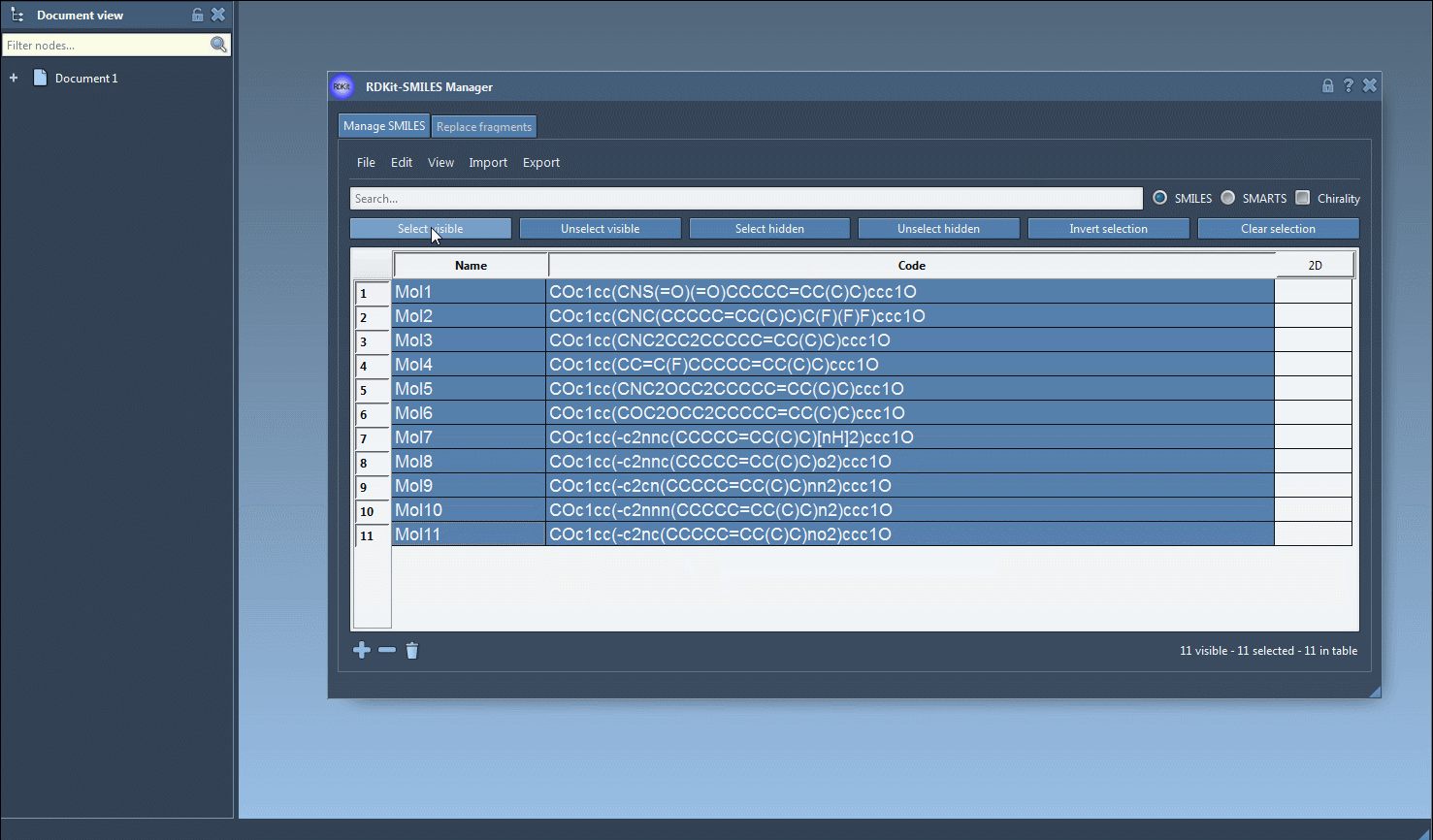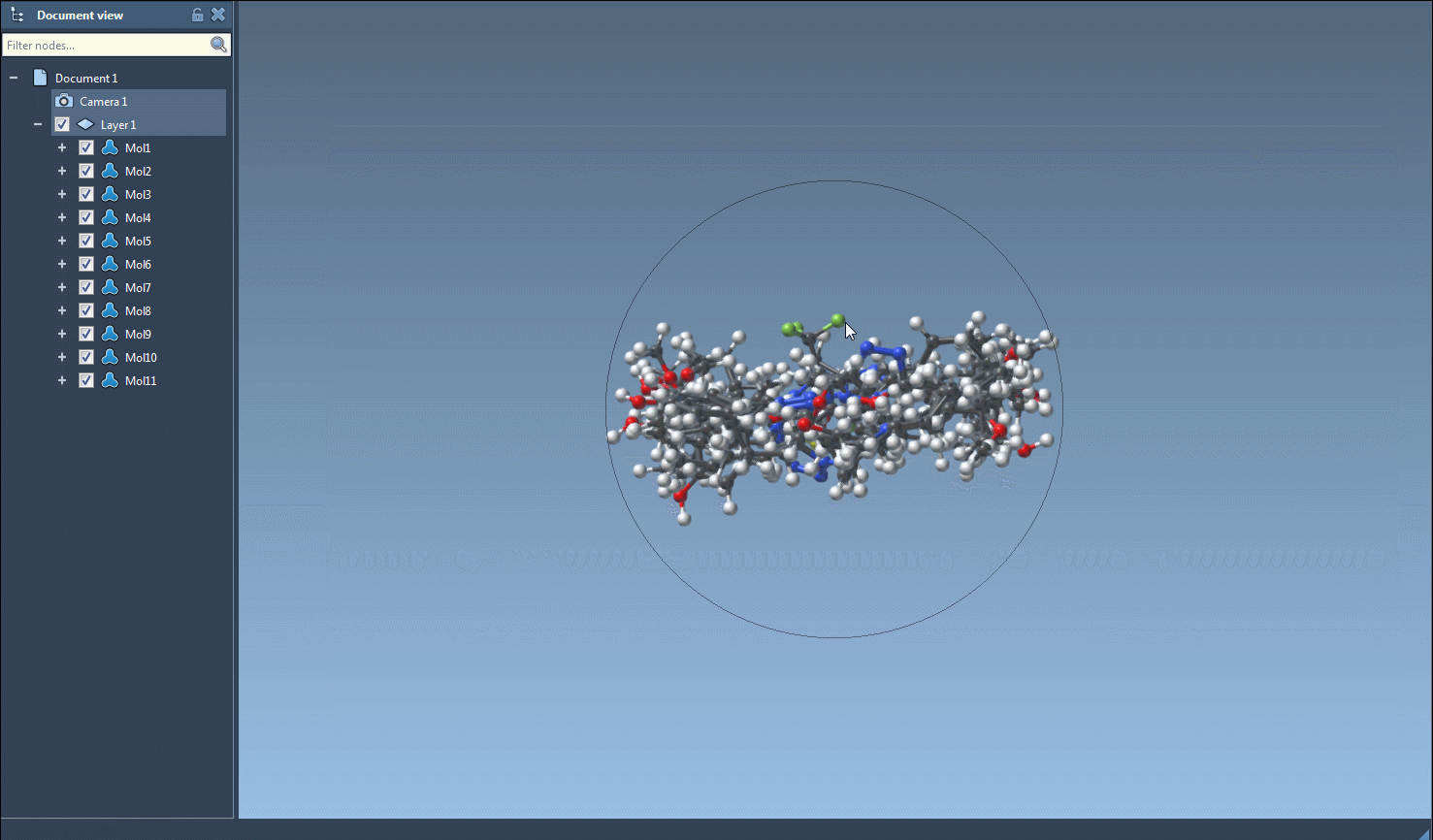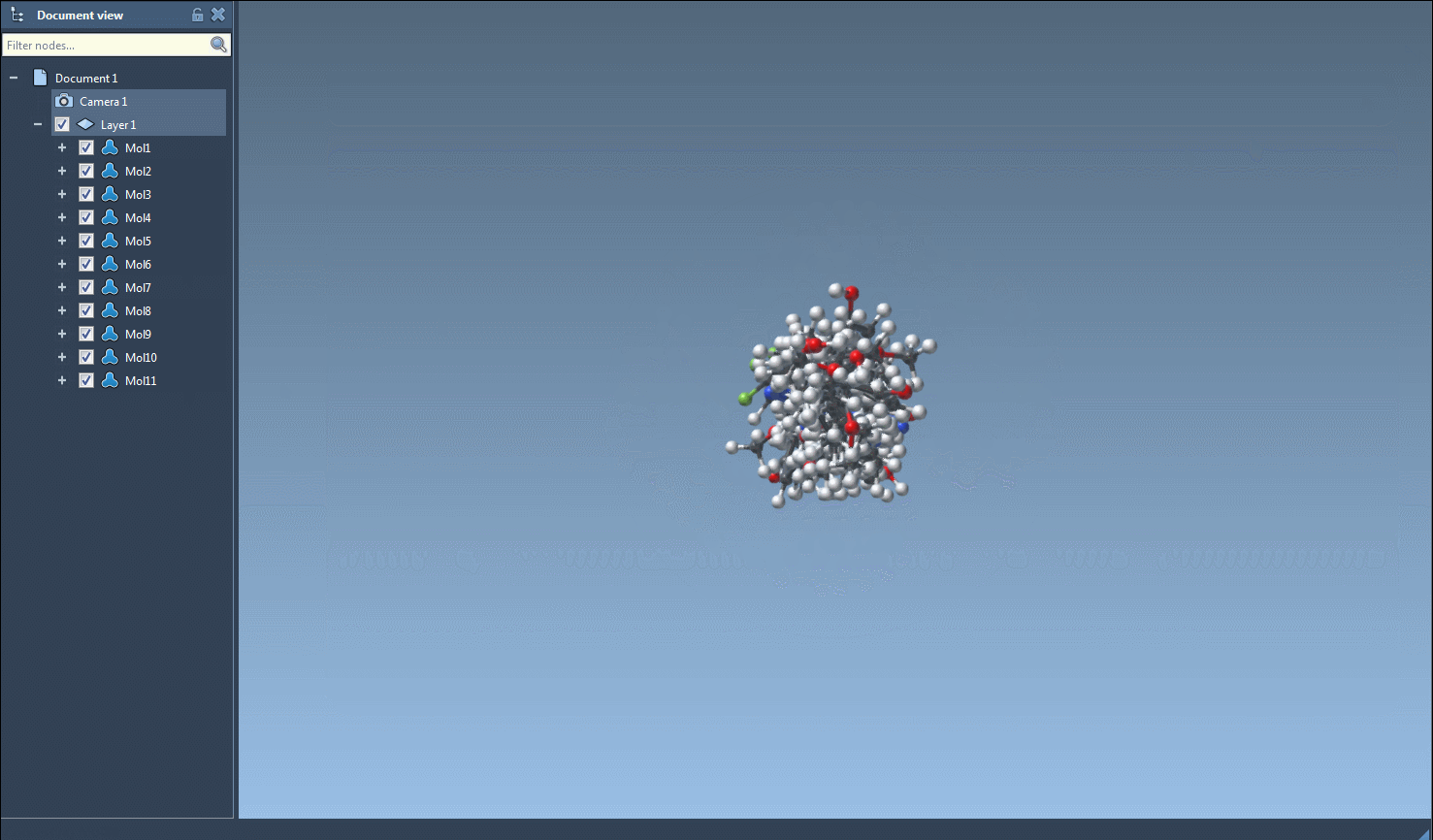
- Begin by selecting the SMILES strings in the table interface of the SMILES Manager. You can import these strings from a file or add them manually.
- Navigate to the Export drop-down menu at the top and click on Selected SMILES string to Document. This will instantly generate 3D structures for the selected molecules.
- The resulting 3D structures, along with their corresponding names, will appear in the active SAMSON document, ready for analysis or visualization.
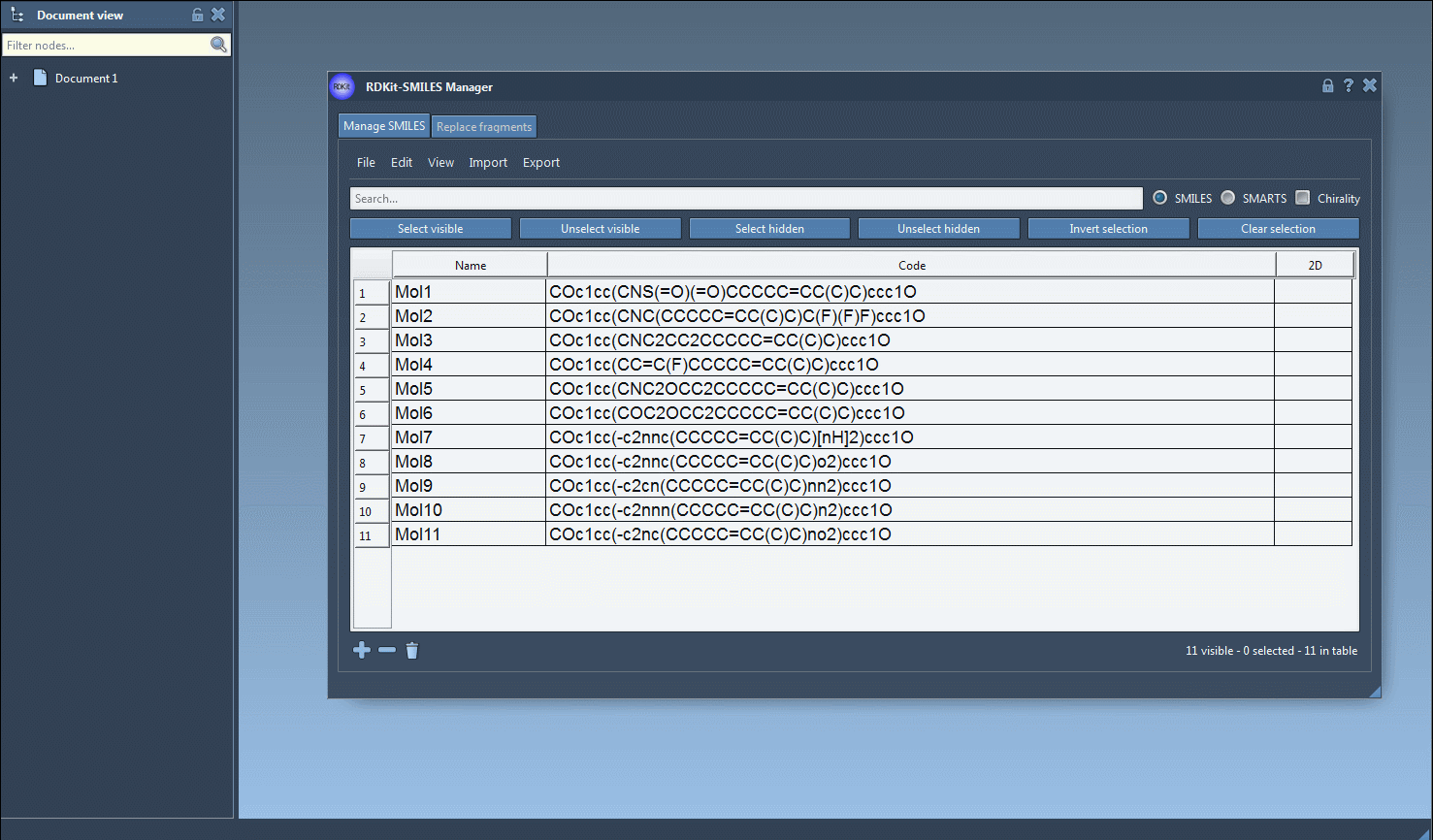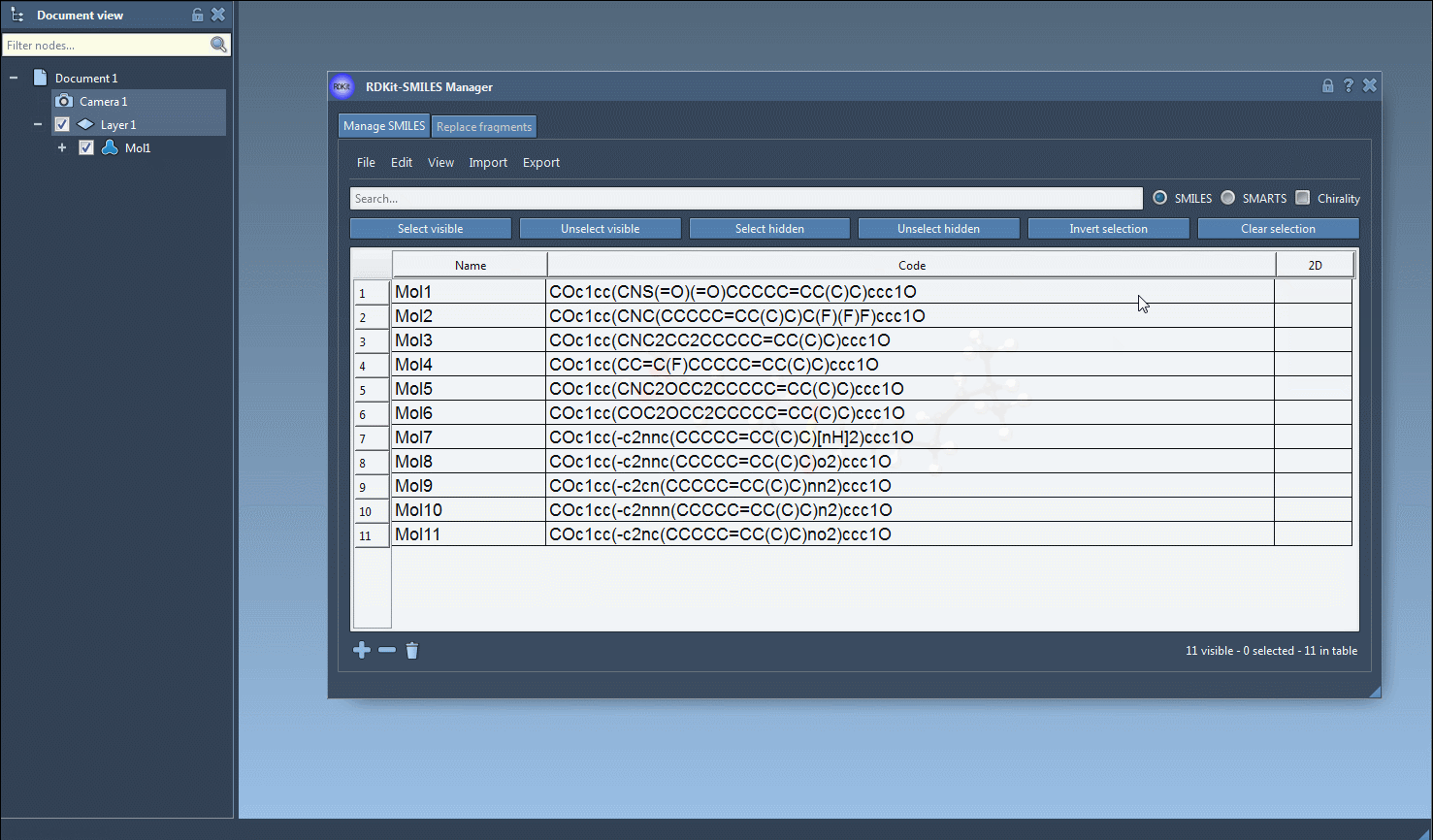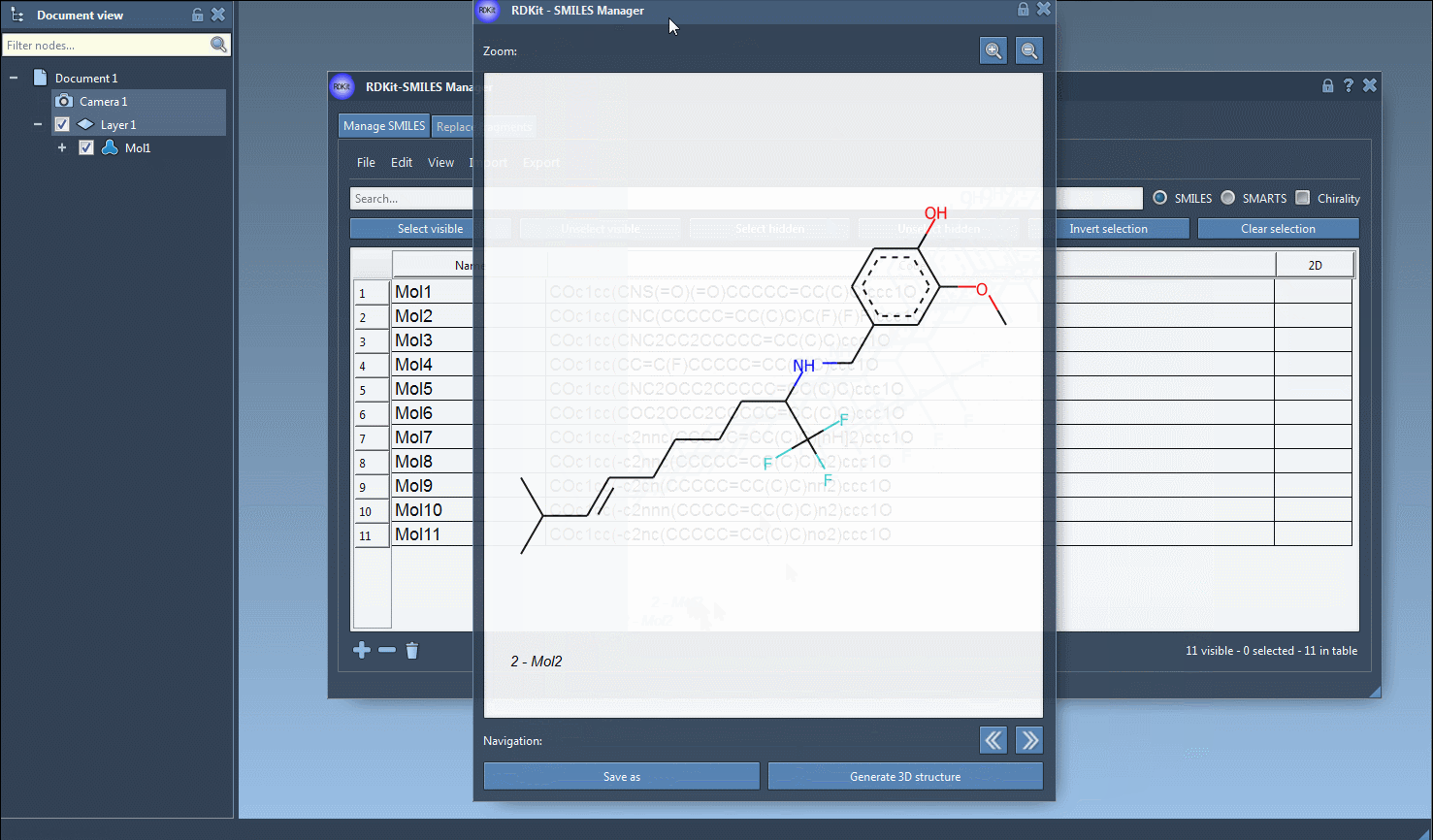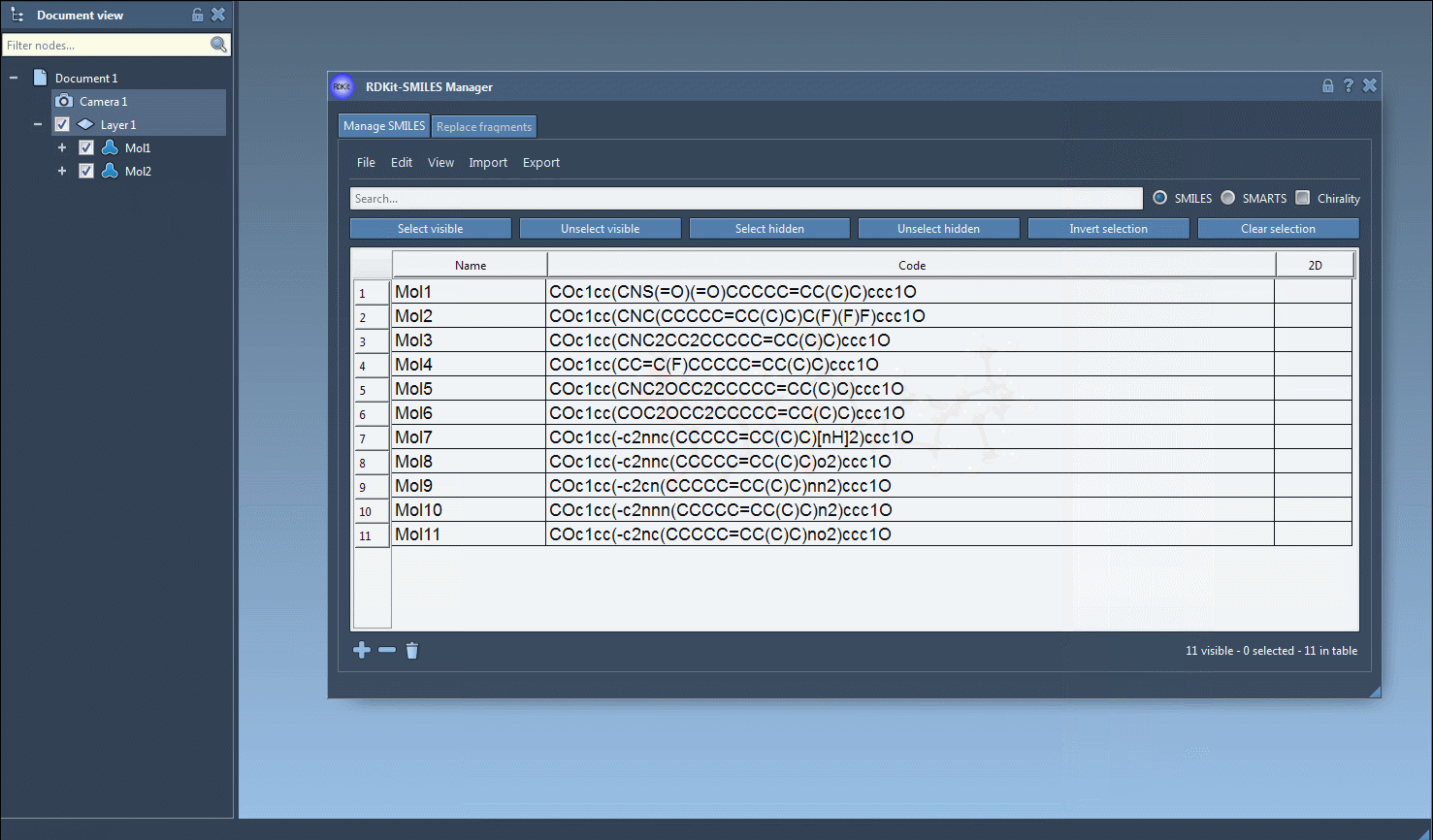
If you prefer to generate 3D structures for individual molecules, you can either right-click on a SMILES string and select Generate 3D Structure from the context menu or use the Generate 3D Structure button in the large view window.
Want to see it in action? Here’s an animation of how this module works:
Features to Streamline Your Workflow
What sets the SMILES Manager apart is how it automates processes and minimizes manual intervention. Here are some highlights:
- Immediate Feedback: Invalid SMILES strings are flagged, so you can correct issues before generating structures. Errors are visually represented in the table.
- Automatic Updates: Alter a SMILES string in the table, and the corresponding 3D structure is immediately updated as well.
- Interactive Visualization: You can open a larger view of any molecule and explore it in detail. Multiple molecules can also be directly added to the active SAMSON document for comparative analyses.
Check out this second animation, which demonstrates the generation of a 3D structure for a single molecule:
Why Molecular Modelers Love This
The SMILES Manager eliminates the need for specialized knowledge or external intermediate tools to convert SMILES into 3D models, making it highly accessible even for beginners. The integration within SAMSON means you can move seamlessly from SMILES handling to advanced structural analysis without leaving the platform. For researchers working on large datasets or complex molecules, this can be a game-changer.
Want to learn more about the SMILES Manager and its other powerful features? Visit the official documentation page.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON now at samson-connect.net.





