One of the common challenges molecular modelers face is efficiently manipulating and editing molecular structures without disrupting complex topologies. Whether it’s creating or breaking covalent bonds, adjusting bond orders, or redefining atom types, maintaining a seamless workflow can be challenging. This is where the Interactive Modeling Universal Force Field (IM-UFF) comes in, a SAMSON Extension designed specifically for interactive editing of molecular systems. Let’s explore how IM-UFF can address these common pain points and enhance your molecular modeling workflow.
What Is IM-UFF and Why Do You Need It?
IM-UFF expands upon the Universal Force Field (UFF), adding capabilities for smooth interactive modeling. Unlike traditional static modeling workflows, IM-UFF allows molecules to undergo dynamic topological changes during simulations. With this tool, you can:
- Create and break covalent bonds.
- Change bond orders.
- Adjust atom typizations dynamically as you manipulate structures.
These capabilities make it significantly easier to model molecular systems and refine topologies based on physical inter-atomic forces, all in real-time.
Getting Started: Requirements and Setup
To use IM-UFF, you’ll first need to add the extension in SAMSON. Once installed, follow these steps to set it up for molecular editing:
- Open a document with the molecular system you wish to simulate.
- Add a simulator via the menu (
Edit > Simulate > Add simulator) or use the shortcutCtrl + Shift + M(orCmd + Shift + Mfor Mac users). - Select “Interactive Modeling Universal Force Field” from the list of interaction models.
- Choose a state updater, such as the FIRE algorithm for energy minimizations.
- Press the OK button to finalize your setup.
How IM-UFF Simplifies Molecular Editing
Once set up, IM-UFF’s dynamic capabilities offer a highly interactive and flexible modeling experience. Below are some of the transformative features you’ll encounter while working with this tool:
- Real-time Topology Updates: Displacing an atom slightly causes the system to adjust locally, preserving the current topology. However, significant movements result in bond breakages, with the topology automatically updating to reflect the change.
- Automatic Bond Formation: Similarly, when an atom moves closer to others, new bonds may form dynamically based on the proximity and overall system configuration.
For example, imagine you’re rearranging a molecular system and need to test multiple configurations. Doing this manually with static workflows might involve tedious recalculations and resetting of bonds. With IM-UFF, these adjustments occur seamlessly in the background, saving time and effort. The following image illustrates the dynamic nature of IM-UFF during molecular manipulation:
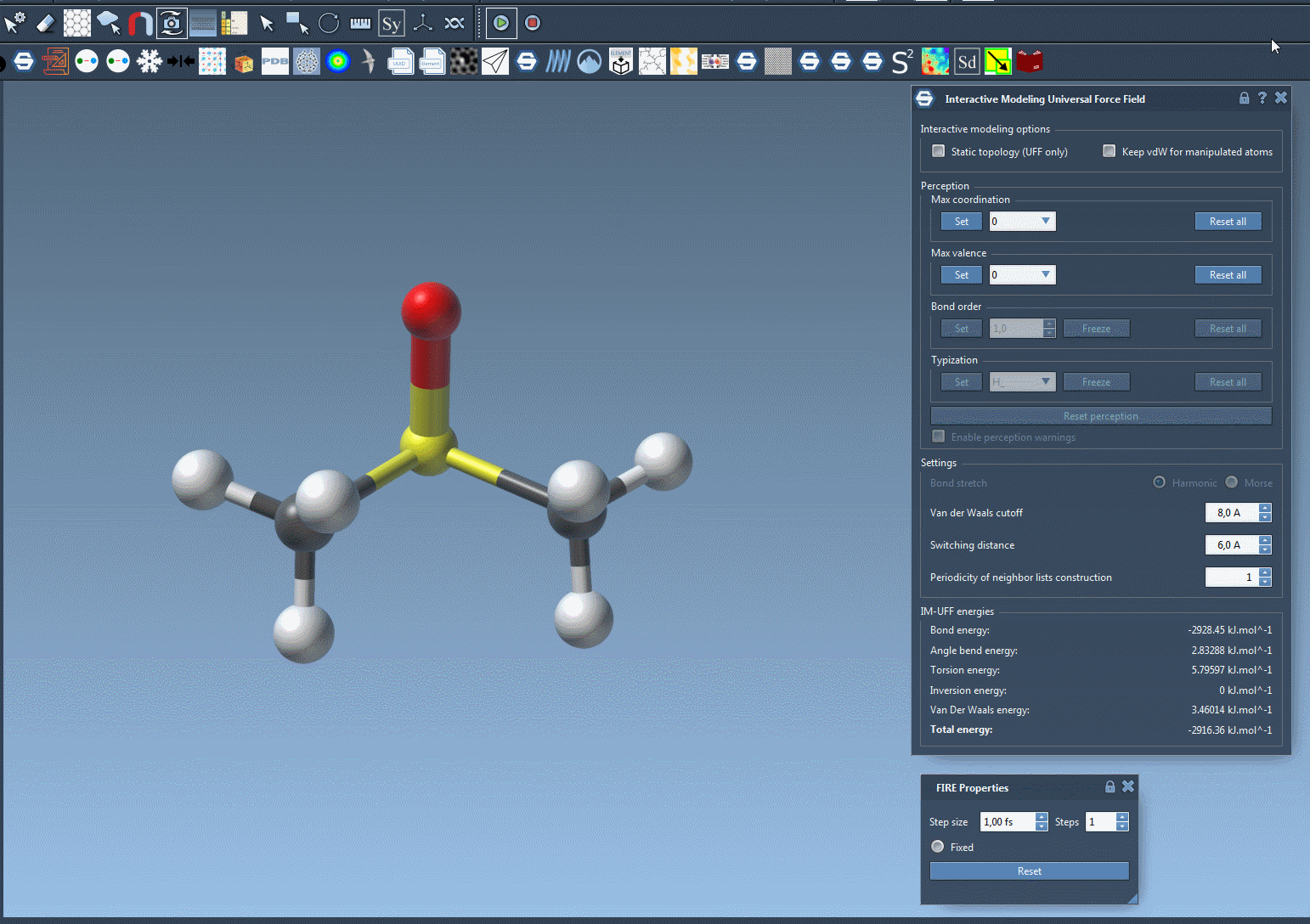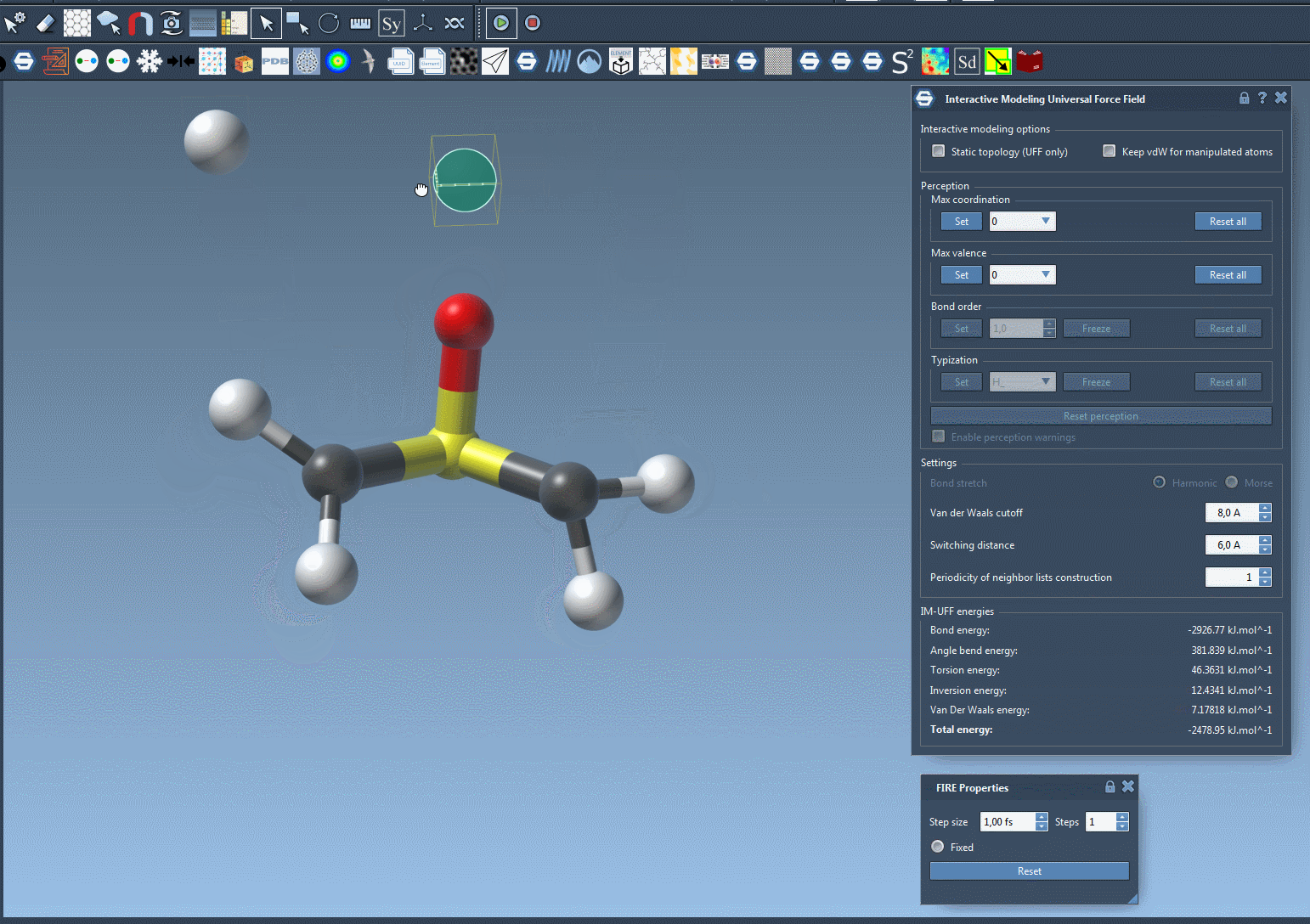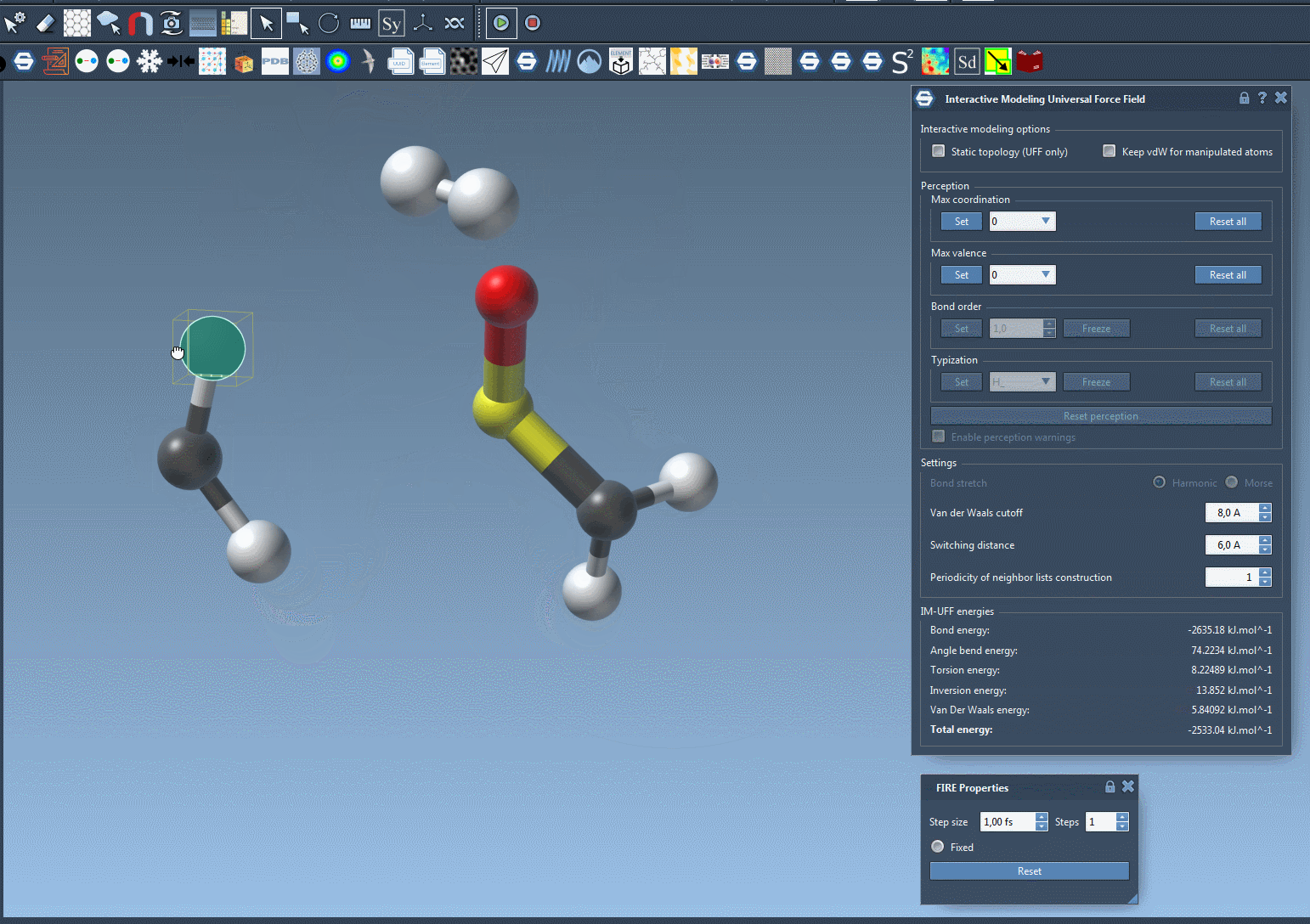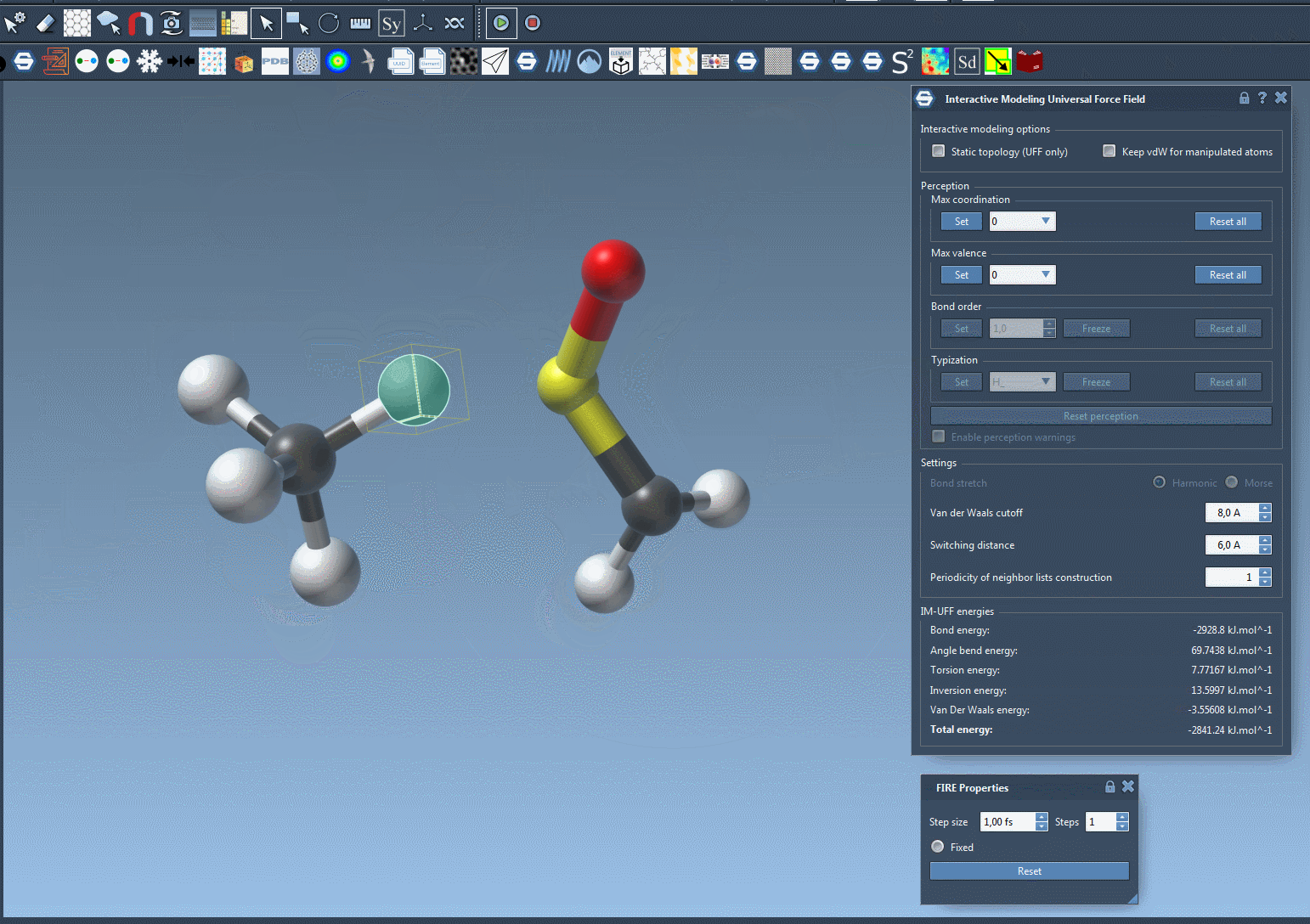
Tips for Optimizing Your Workflow
To make the most of IM-UFF, keep these tips in mind:
- Start Small: Begin with a small or medium-sized molecular system to easily observe topology changes during manipulations.
- Adjust Parameters: Use the IM-UFF parameter window to customize settings like van der Waals cutoffs and interaction forces for more nuanced simulations.
- Utilize Interactive Options: For more control, explore interactive modeling options like “Static topology (UFF only)” and “Keep vdW for manipulated.” These settings can help fine-tune how the system behaves during editing.
Conclusion
IM-UFF is a powerful tool for molecular modelers looking to simplify interactive modeling tasks. With its ability to handle dynamic topological changes and seamlessly integrate with SAMSON’s interface, it’s an invaluable asset for improving productivity and enhancing accuracy in molecular design projects. To learn more and access detailed instructions, visit the original documentation page at https://documentation.samson-connect.net/tutorials/uff/im-uff/.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Get SAMSON today at https://www.samson-connect.net.





