Tracking the movement of molecules during molecular simulations, such as ligand unbinding or macromolecular conformational changes, is essential for understanding biological processes. However, visualizing this motion effectively can be a challenge for molecular modelers. The Pathlines extension in the SAMSON platform provides a solution by enabling users to display center-of-mass (COM) trajectories of selected atoms along predefined paths. This blog post will guide you step-by-step through the process of using Pathlines to visualize COM motion and enhance your molecular modeling workflow.
What Are Pathlines?
Pathlines are visual models that plot the COM trajectory of a selected group of atoms along a specific molecular path. These visualizations are particularly useful for analyzing processes such as:
- Ligand unbinding and rebinding routes in molecular dynamics simulations.
- Macromolecular conformational changes, such as domain motions in proteins.
- Tracking atomic motion across reaction coordinate workflows.
This approach simplifies the understanding of complex systems by providing a clear, graphical representation of motion over time. Now, let’s dive into how to use it.
Getting Started
- Log into SAMSON Connect.
- Visit the Pathlines Extension page and click Add.
- Restart SAMSON to activate the extension.
Step-by-Step Guide to Visualizing COM Motion
Step 1: Load Your Sample System
Start by loading a pre-prepared system like the Lactose permease structure:
- Navigate to Home > Download in SAMSON.
- Paste the following URL: https://www.samson-connect.net/documents/046f1acd-c799-40f6-8185-cb4847eff795.
- Click Download to load the sample document.
This document contains a structural model of Lactose permease (1PV7) with its ligand Thiodigalactosid (TDG) and unbinding paths generated using the Ligand Path Finder.

Step 2: Select Atoms and Paths
In the Document view, select the atoms or groups you want to analyze and the paths along which you want to visualize the COM motion. You can hold the Ctrl/Cmd key to select multiple nodes. If you do not select specific atoms or paths, the entire system and all available paths will be used.

Step 3: Create a Pathline Visual Model
Once you have your selections:
- Go to Visualization > Visual model > More… (shortcut: Ctrl/Cmd + Shift + V).
- In the dialog, select Pathline of the center of mass and confirm by clicking OK.
This will create a visual pathline showing the motion of the selected atoms’ center of mass along the chosen paths.
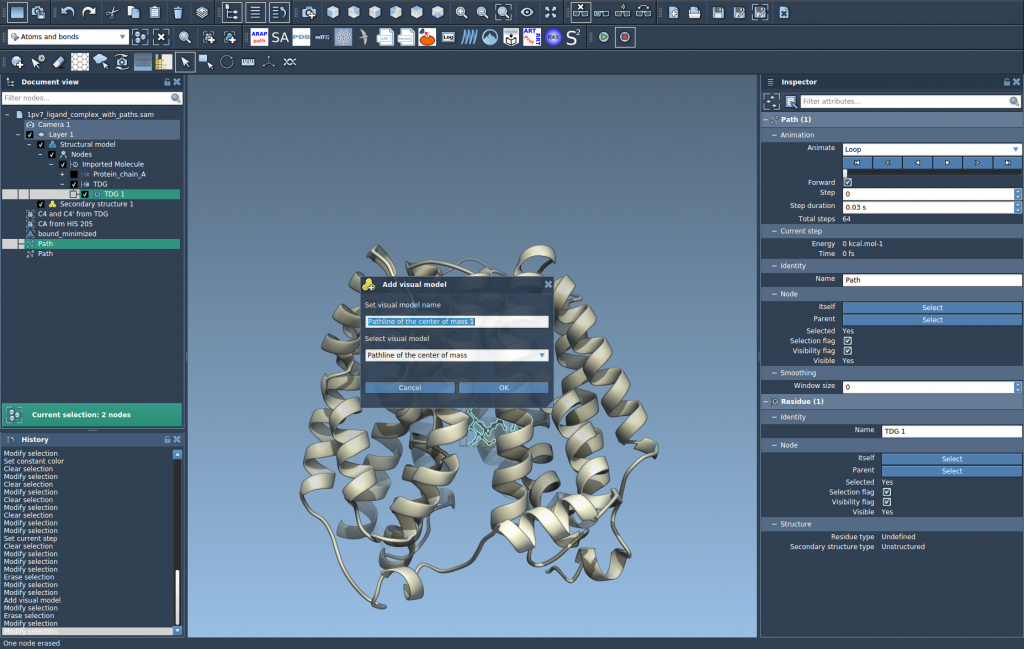
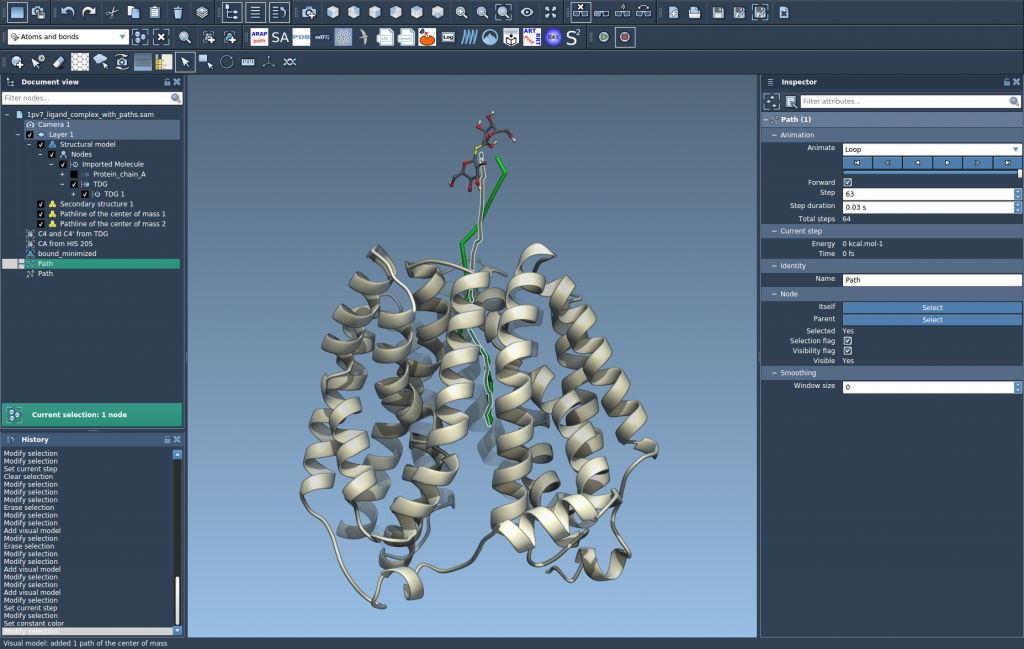
Step 4: Explore and Customize
Finally, explore and tweak the resulting pathlines to suit your analysis:
- Double-click a path in the Document view to start or stop it.
- Right-click a path for more options under Path > ….
- Adjust visual attributes such as color and thickness by selecting the pathline and accessing the Inspector (Ctrl/Cmd + 2).
Benefits of Pathlines
Using Pathlines, you can:
- Identify unbinding pathways for ligands.
- Analyze collective domain motions in proteins.
- Develop insights into center-of-mass movement in reaction workflows.
By incorporating this visualization method, molecular modelers can enhance their understanding of dynamic molecule behavior with clarity and precision.
To learn more, visit the full documentation page: Visualize Center-of-Mass Motion with Pathlines.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Download SAMSON at https://www.samson-connect.net.





