Substructure searching is a common challenge for molecular modelers. Whether you’re analyzing complex molecules or attempting to identify specific patterns within a chemical library, the process can often feel tedious and prone to error. If this sounds familiar, SAMSON’s SMILES Manager may have the solution you need.
The new version of SAMSON’s SMILES Manager provides an improved Substructure Search feature, allowing you to efficiently search for combinations of molecular patterns. In this blog post, we’ll explore how this tool can be used to find exactly what you’re looking for—quickly, accurately, and with minimal effort.
What Is Substructure Search?
Substructure search allows users to identify specific chemical patterns, or substructures, within a larger set of molecules. For example, you might want to find all molecules in your dataset that contain both a nitrogen atom (N) and a fluorine atom (F). These kinds of searches are vital in tasks ranging from drug discovery to material design, but they typically require sophisticated tools and know-how to execute correctly.
How Does It Work in SAMSON?
The updated SMILES Manager in SAMSON leverages RDKit-based functions to make substructure searching intuitive. Here’s a simple breakdown of how to use this feature:
-
Start your search: Open the Search bar in the SMILES Manager interface. Begin by entering the pattern you’re looking for, such as
Ffor fluorine atoms. After pressing Select visible, only molecules containing fluorine will be selected, and their corresponding SMILES strings will be highlighted in the list. -
Combine search terms: Clear the search bar and enter a new term—e.g.,
Nfor nitrogen atoms. This search result will include only the previously selected molecules containing fluorine and nitrogen. Use the Unselect hidden button to refine your selection, ensuring only molecules visible in the results are retained. - Adjust as needed: By combining multiple search terms in this way, complex substructure searches can be performed effectively without requiring separate scripts or external tools.
The functionality of the tool is showcased in the image below:
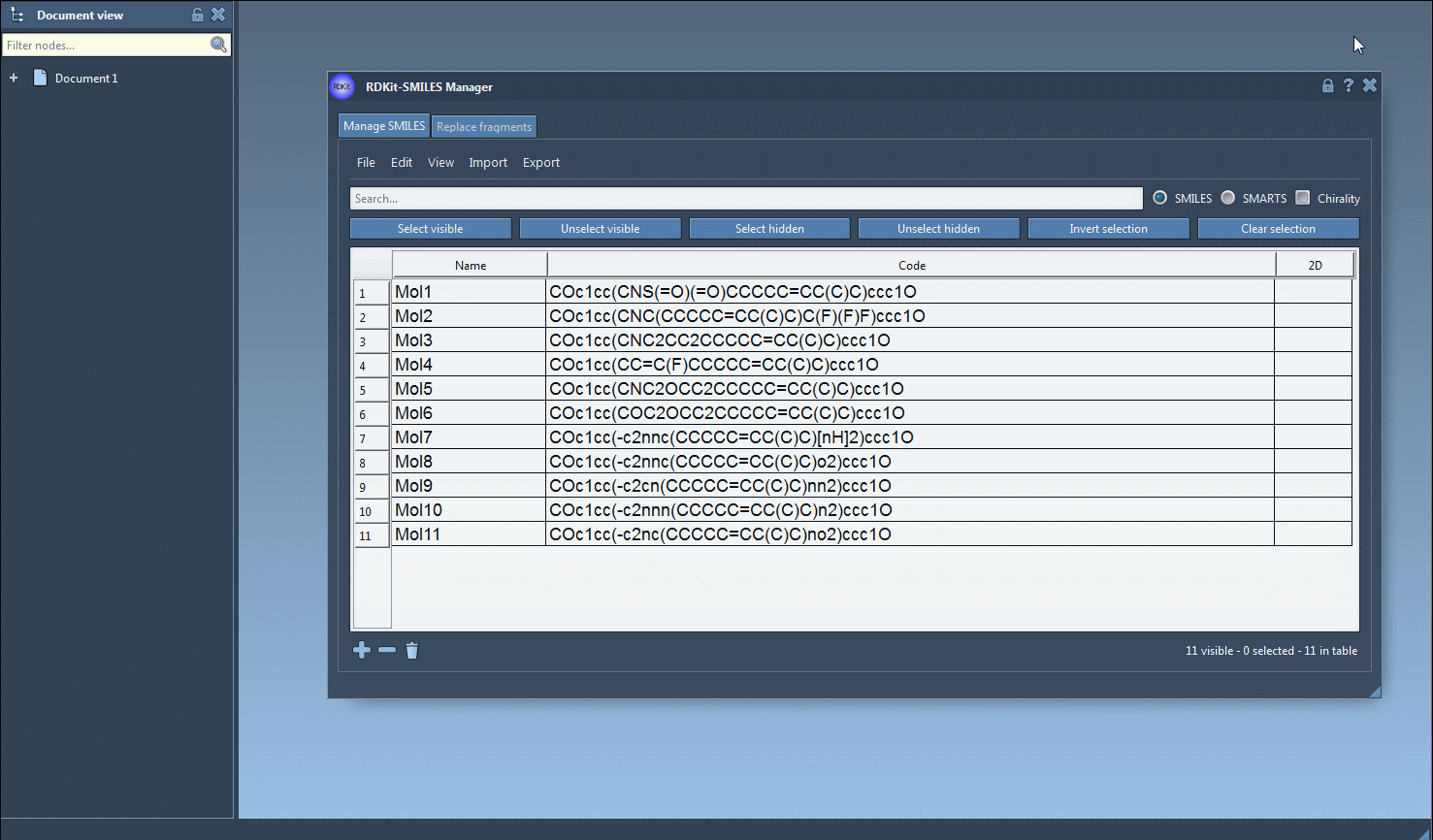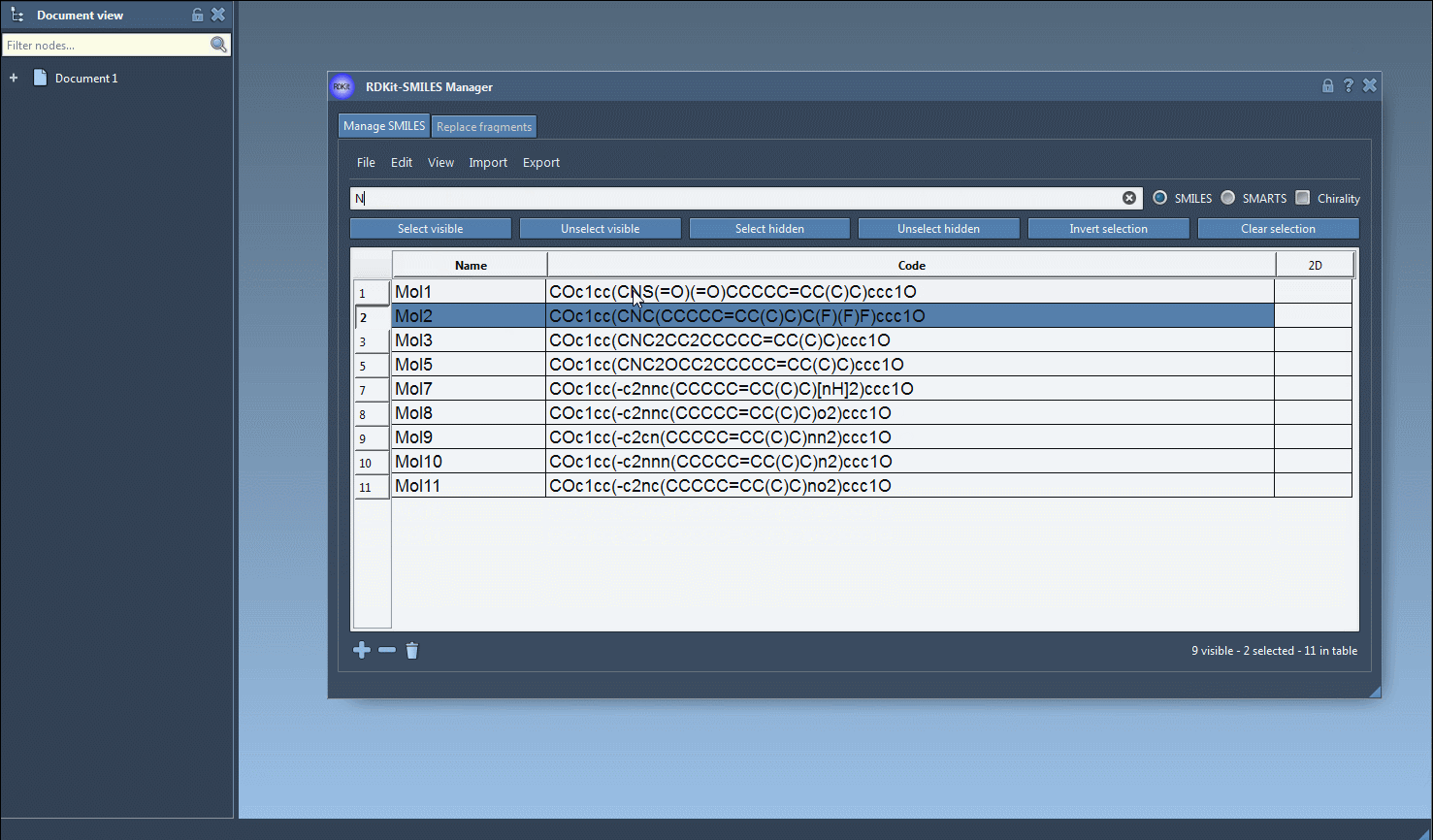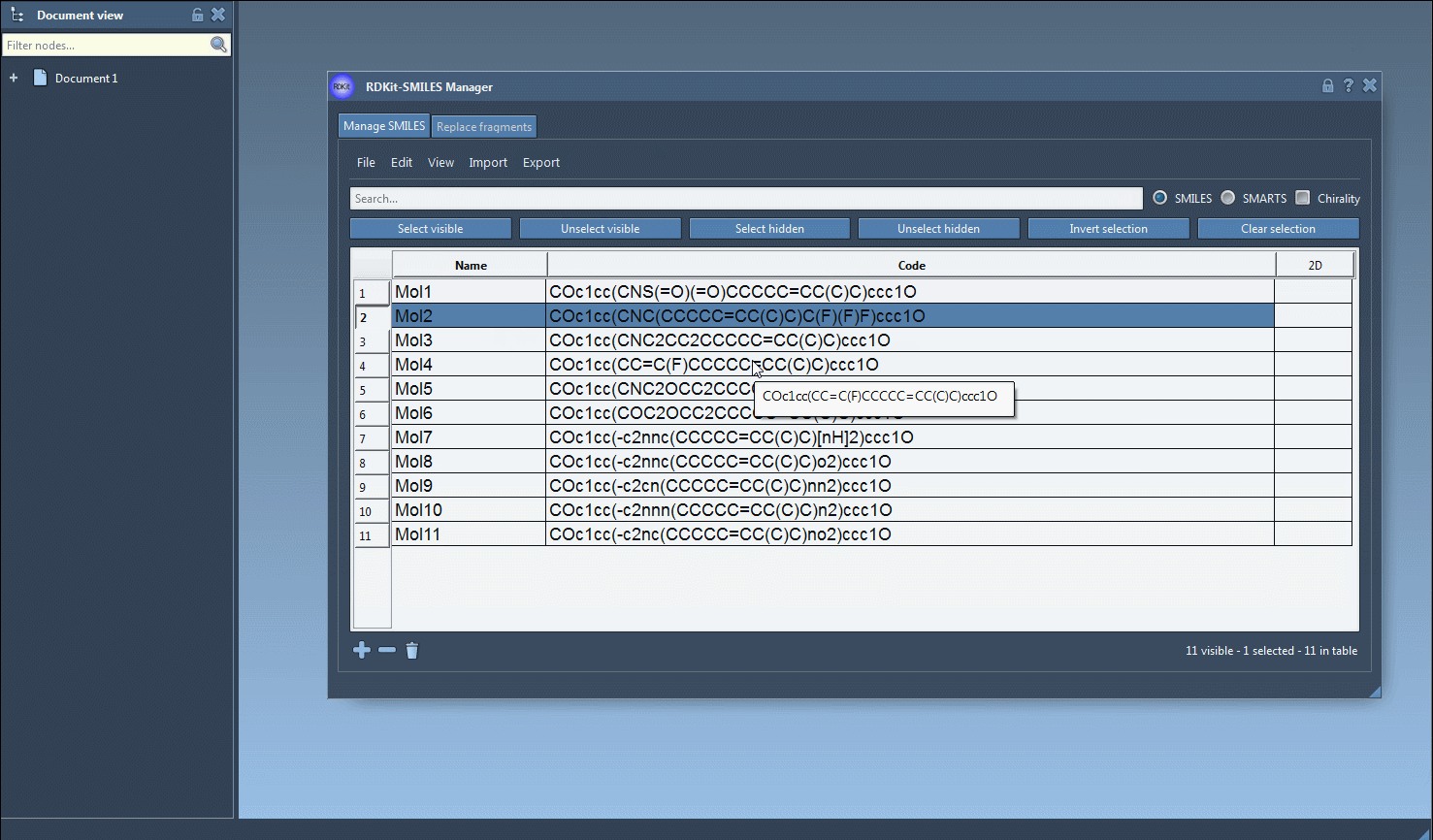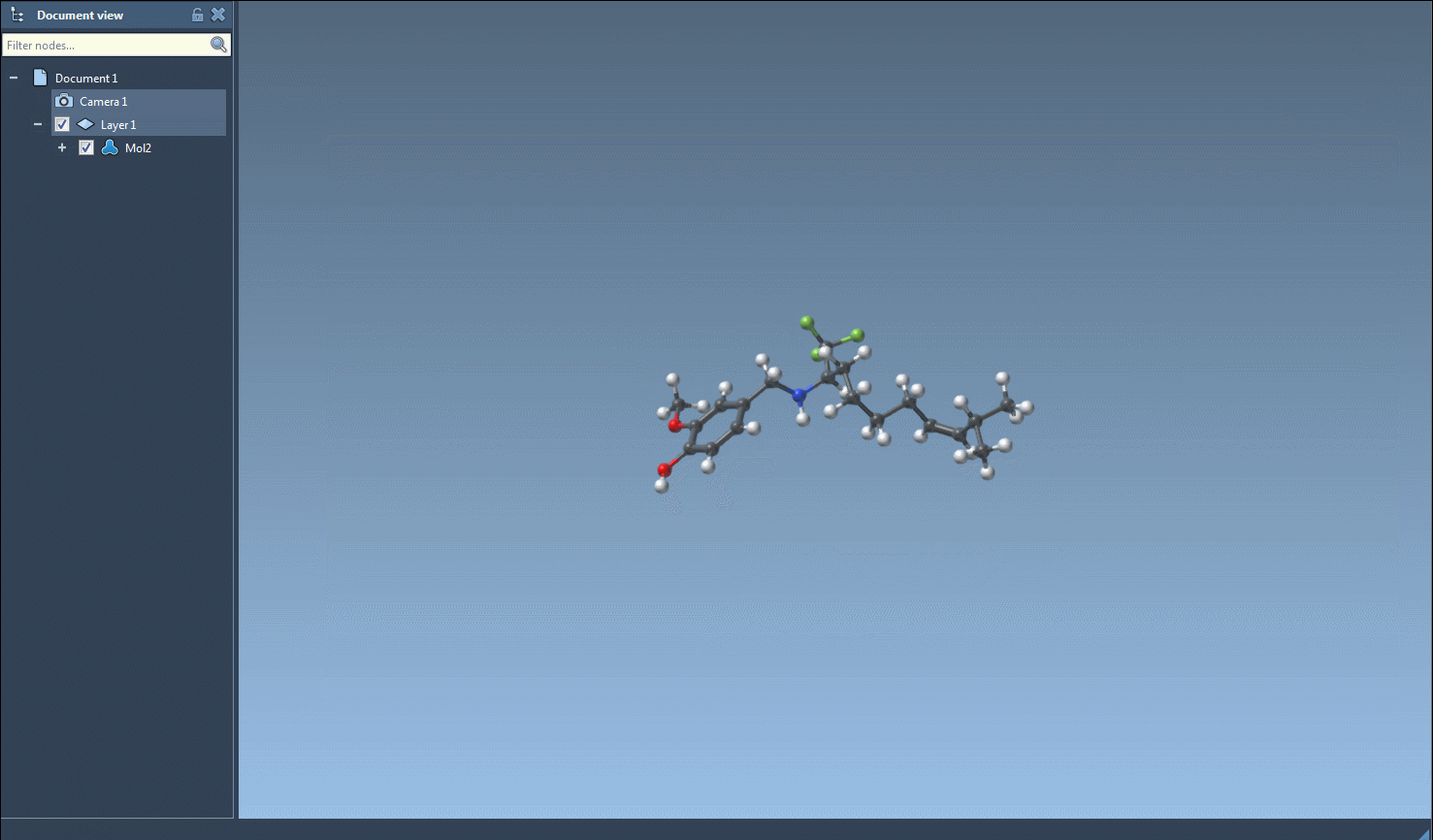
Additional Flexibility
By default, RDKit doesn’t use stereochemical information for substructure searches. However, for users who require this information, SAMSON’s SMILES Manager offers the option to enable chirality during searches. This ensures that the results align perfectly with your project’s requirements, whether or not stereochemistry is crucial.
Who Should Use This Feature?
If you work with chemical libraries or need to identify molecular patterns quickly, the substructure search feature in SAMSON is designed for you. It eliminates the guesswork and manual effort often associated with traditional methods, enabling smoother workflows for both beginners and advanced users alike.
Whether you’re analyzing materials, designing new drugs, or studying chemical reactions, this tool empowers you to screen for relevant molecular patterns more efficiently, ultimately saving time and effort for more complex tasks in your pipeline.
Learn more about how SAMSON’s SMILES Manager can improve your molecular modeling experience by visiting the full documentation page here.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can get access to SAMSON at www.samson-connect.net.





