Molecular modeling often requires understanding the motion of atoms or groups of atoms, tracking ligand unbinding, or analyzing macromolecular conformational changes across simulations. This can be a challenging task, especially in complex systems. With SAMSON’s Pathlines visual model, you can streamline this process and visualize the motion of the center of mass (COM) of selected atoms along defined paths. Here’s a step-by-step guide to help you navigate this tool.
What Are Pathlines?
Pathlines in SAMSON provide a clear representation of the motion of the center of mass for selected atoms. This is particularly useful for:
- Tracking ligand movement during binding and unbinding events.
- Visualizing center-of-mass dynamics in complex biological systems.
- Analyzing macromolecular domain movements or diffusion events.
It’s a highly effective way to analyze and interpret molecular trajectories across time in an intuitive visual format.
Step-by-Step: How to Create Pathlines
Here’s how you can generate and explore pathlines:
Step 1: Prepare the System
Loading a Sample: To follow this tutorial, you can load a sample system of Lactose permease (1PV7) with its ligand, Thiodigalactoside (TDG). This includes paths generated using the Ligand Path Finder extension. You can access the sample system via this link:
- Click Home > Download in SAMSON.
- Paste the following link: https://www.samson-connect.net/documents/046f1acd-c799-40f6-8185-cb4847eff795.

This sample document includes a protein model, a ligand, and predefined paths for analysis, making it easier to get started.
Step 2: Select Atoms and Paths
Specify the Atoms: Select the group of atoms for which you want to analyze the center-of-mass motion. You can do this by using the Document view. To select multiple nodes, hold the Ctrl (Windows) or Cmd (Mac) key. If you don’t select specific atoms, the entire system’s motion will be visualized.

Similarly, select one or more paths to visualize specific trajectories. If no paths are selected, SAMSON uses all available paths in the document.
Step 3: Create the Pathline Visual Model
Generate the Visualization: Navigate to Visualization > Visual model > More… (shortcut: Ctrl/Cmd + Shift + V), and choose the Pathline of the center of mass option.
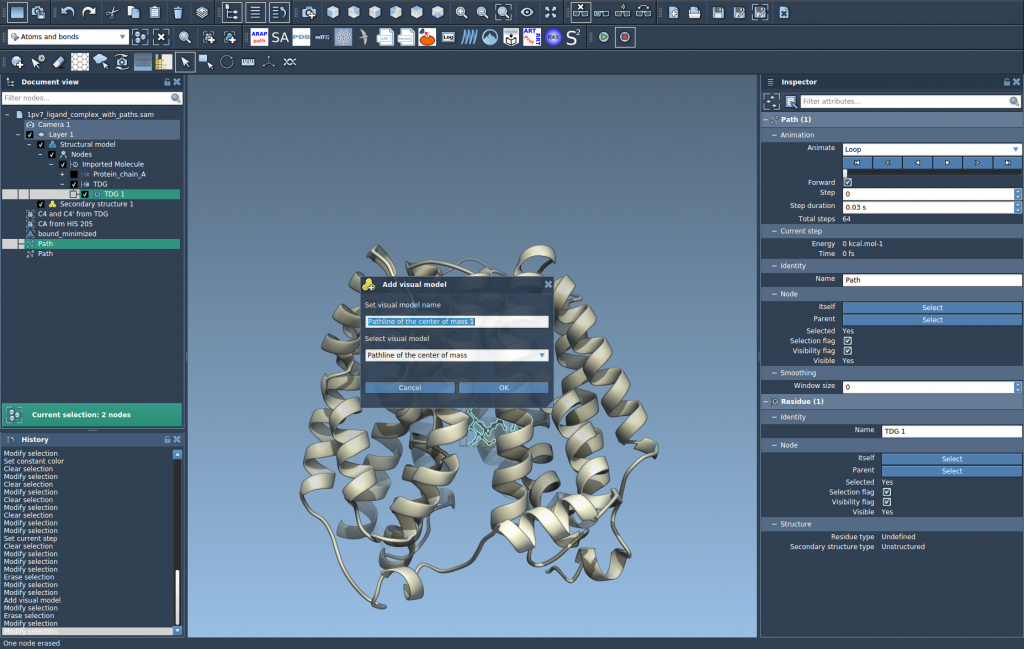
Click OK and observe as SAMSON generates a visual model displaying the motion of the selected atoms’ center of mass along the chosen path(s).
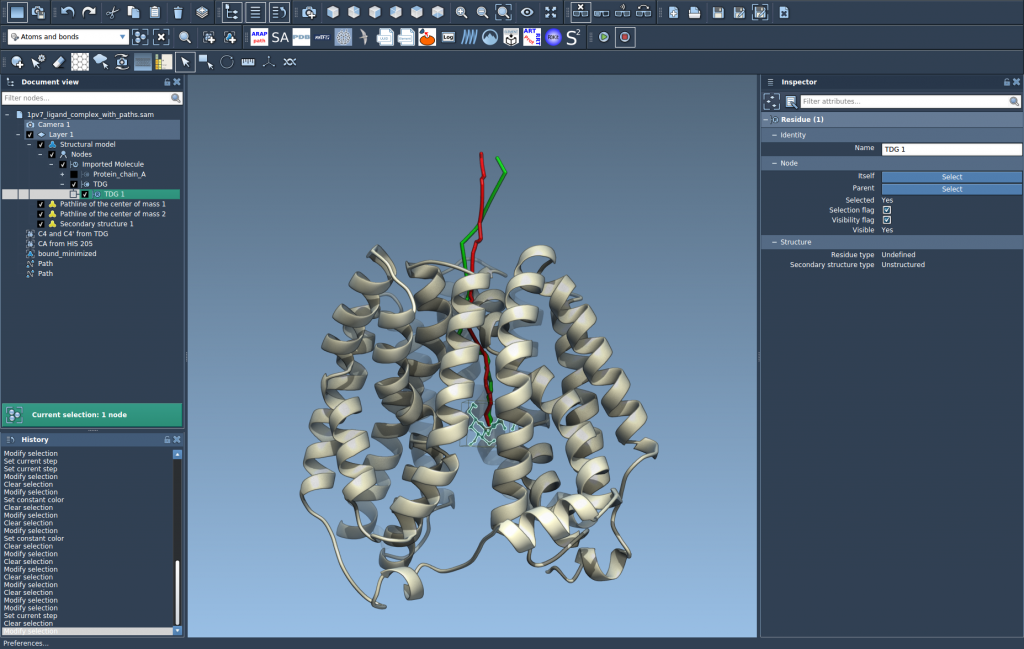
Step 4: Explore and Fine-Tune
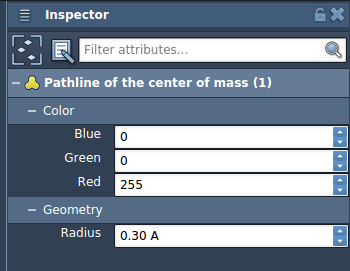
Customize and Analyze: You can interact with the pathlines to explore various frames, adjust visual properties like color and thickness, and control the trajectory. Use the Inspector (Ctrl/Cmd + 2) to refine the settings further.
Applications
Pathlines are versatile and can be employed in several scenarios:
- Track ligand unbinding/rebinding routes during drug design.
- Analyze collective domain motion in protein complexes.
- Understand center-of-mass shifts in reaction coordinate workflows.
Learn More
Ready to integrate pathlines into your workflow? Visit the official documentation for more detailed steps and tips: Pathlines Documentation.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Ready to start? Get SAMSON at SAMSON Connect.





