For molecular modelers, navigating large molecular structures and selecting specific components can often feel like finding a needle in a haystack. Fortunately, SAMSON introduces the powerful Node Specification Language (NSL) to streamline this task. NSL enables users to create precise and efficient queries, allowing them to select nodes, molecules, and other structural components effortlessly based on a wide range of parameters such as type, name, proximity, and more.
The Power of NSL in the Find Command
Imagine needing to isolate specific residues, atoms, or functional groups in a large biomolecular structure. SAMSON’s Find command leverages NSL to help you achieve this quickly. Using a simple string query, you can extract exactly the elements you need, no matter how intricate the molecular organization might seem.
Here’s how it works:
- You enter an NSL query in the Find command’s search box.
- The system provides context-sensitive completions when you press the Tab key, simplifying complex searches.
- The result is a filtered, precise selection that matches your criteria.
For example, let’s consider searching for alanine residues. Enter "ALA and press Tab. You’ll see suggestions like:
"ALA 22 Backbone""ALA 22 Side chain""ALA 28"
This functionality makes NSL remarkably user-friendly, even for beginners.
Enhance Your Workflow with AI Assistance
SAMSON further simplifies node selection through its integrated AI Assistant. If you’re unsure how to frame your NSL query, just click on the ![]() Ask AI button. The AI Assistant understands your document’s hierarchy and can generate tailored NSL expressions, saving you time and effort. This is especially useful for new users learning to harness the full power of NSL.
Ask AI button. The AI Assistant understands your document’s hierarchy and can generate tailored NSL expressions, saving you time and effort. This is especially useful for new users learning to harness the full power of NSL.
Visualize and Filter Your Selections
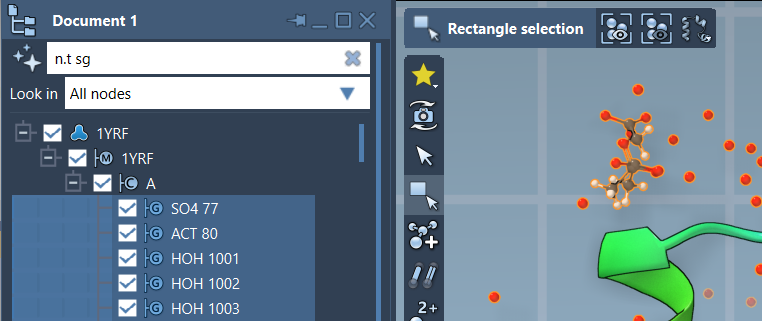
Besides the Find command, NSL can also be employed in SAMSON’s Document view. Here, you can filter nodes based on NSL criteria and refine your selection further. For instance, applying the query node.type structuralGroup (or its short version, n.t sg) highlights all structural groups, allowing you to examine specific molecular assemblies with ease.
Once filtered, pressing Enter completes the selection. The capability to visually navigate filtered molecular components makes it easy to focus on relevant areas for your modeling tasks. An example image, presented below, demonstrates this filtering approach in action:
Why Use NSL?
Utilizing the Node Specification Language empowers molecular modelers to:
- Save time by locating specific nodes without manually browsing structures.
- Handle complex molecular data with ease by leveraging logical and proximity operators.
- Focus on the essential molecular components for precise analyses and designs.
Whether you’re filtering residues by name, identifying atoms within a certain distance, or selecting structural groups with specific properties, NSL becomes an indispensable tool in your molecular modeling toolbox.
To dive deeper into NSL, explore the original documentation page here: https://documentation.samson-connect.net/users/latest/nsl/.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Get SAMSON here.





