Finding transition paths between molecular states can often be a challenging task for molecular modelers. Whether you’re working on uncovering reaction mechanisms or exploring conformational changes, capturing these pathways accurately and efficiently is essential. This is where the Parallel Nudged Elastic Band (P-NEB) app in the SAMSON molecular design platform can prove invaluable. Let’s dive into how to streamline transition path optimization using this tool.
Why Is Transition Path Optimization Important?
Molecular systems often exist in different states, such as folded and unfolded protein conformations or bound and unbound states in ligand interactions. Transition paths reveal the intermediate steps that connect these states, providing a detailed understanding of the molecular mechanism. However, accurately identifying these paths, particularly those with minimal energy, can be computationally intensive and time-consuming.
The P-NEB method offers a structured solution to this problem. It works by optimizing a number of intermediate “images” (representations of molecular states) along the path using the NEB method, ensuring minimal energy pathways with evenly spaced conformations.
Getting Started with P-NEB
The P-NEB app, part of the SAMSON platform, simplifies this process by implementing the climbing image NEB method, making it easy to optimize existing transition paths or generate them from conformations. Follow these steps to get started:
1. Load a sample input model:
To experiment with a predefined system, head to Home > Download in SAMSON and use one of the following sample documents:
- Zinc ligand unbinding trajectory – A smaller system ideal for tests.
- Protein-ligand complex of Lactose permease and Thiodigalactosid – A more complex example with unbinding paths derived from the Ligand Path Finder tutorial.

By loading one of these documents, you can follow along with the tutorial on how to improve ligand unbinding pathways using the P-NEB app.
P-NEB App Settings at a Glance
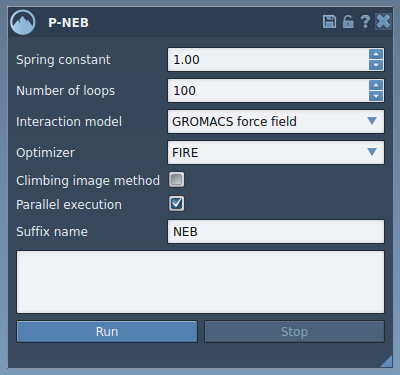
After starting the P-NEB app (Home > Apps > All > P-NEB), you’ll find several configuration options to tailor the optimization process:
- Spring constant: Defines the spring force between intermediate images (e.g., 1.00).
- Number of loops: Specifies how many optimization steps should be performed (e.g., 100).
- Interaction model: Choose the force field for energy and force computations (e.g., Universal Force Field).
- Optimizer: Select the algorithm for optimizing the path (e.g., FIRE).
- Climbing image method: Toggle the use of climbing image strategy to pinpoint saddle points. You can leave this unchecked initially and try it later.
- Parallel execution: For faster computation, enable this option to run threads for each conformation within the path.
- Suffix name: Define a custom suffix name (e.g., NEB) for the generated path or conformations.
Optimizing Transition Paths
Using the P-NEB app is straightforward. First, select a path node in the Document view, or create one by combining individual conformations (Conformation > Create path from conformations). Once selected, click on the Run button in the app. A dialog will appear asking whether to use existing bonds. Confirm it and proceed.
As the optimization progresses, you can monitor the status in real-time through the status bar, which indicates computation progress:

When the optimization finishes, a new path will appear in the Document view, ready for inspection. You can explore options such as visualizing the path’s animation or analyzing it with the Inspector.
Learn More
The P-NEB app offers a robust method to efficiently optimize transition paths—be it for smaller test cases or complex molecular systems. To explore the full step-by-step guide, visit the official documentation page.
SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON at SAMSON Connect.





