For molecular modelers, selecting the right sets of atoms, residues, or nodes is a frequent and crucial task. Whether you’re isolating ligand-receptor complexes, homing in on specific residue types, or filtering structures within a defined distance, efficient node selection can save hours of repetitive work. Luckily, SAMSON’s Node Specification Language (NSL) offers a powerful way to select nodes based on complex queries, leveraging the platform’s sophisticated molecular modeling tools.
Let’s take a closer look at how NSL can simplify and enhance the task of molecular selection in SAMSON.
The Basics: Why Use NSL?
SAMSON’s Node Specification Language is a query system that allows users to select molecular structures like atoms, residues, ligands, bonds, and more, based on their properties. You can use NSL in the Find command or to filter nodes in the Document view. For example, you can filter residues to identify all instances of alanines or locate atoms within a certain distance from a binding pocket.
Quick Example: To find all residues within 5 angstroms of a sulfur atom, you could enter:
|
1 |
residue within 5A of S |
With NSL, even complex queries are intuitive. For instance, let’s say you wish to exclude alanines:
|
1 |
residue and not residue.type ALA |
To avoid surprises, you can further refine this to ensure only residues are selected (e.g., ignoring folders, which are also nodes):
|
1 |
node.type residue and not residue.type ALA |
Find Command: Context-Sensitive Search
The Find command in SAMSON is a primary spot for using NSL. You simply enter your query in the search box, and pressing Tab offers context-sensitive completion. This feature is especially helpful when searching for node names or attributes.
For example, entering a partial name like:
|
1 |
"ALA |
and pressing Tab will suggest all matches, such as:
- “ALA 22 Backbone”
- “ALA 28 Side Chain”
- …and more.

Filtering Nodes in the Document View
NSL is equally useful when you’re working in the Document view. With a well-crafted query, you can filter nodes directly and refine your model in real-time. For example, here’s how you can select structural groups using an NSL expression:
|
1 |
node.type structuralGroup |
Or its shortened version:
|
1 |
n.t sg |
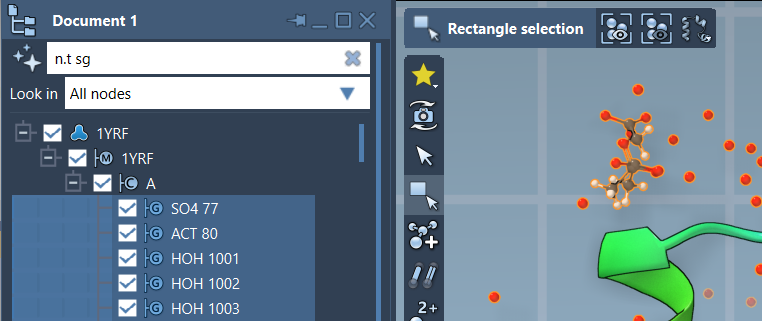
Pressing Enter after applying the filter will select the matching nodes for further operations in SAMSON.
Tips to Supercharge Your Selections
With its AI-powered capabilities, SAMSON makes it even easier to master NSL. You can use the “Ask AI” button to automatically generate NSL expressions tailored to your active document’s hierarchy. Simply click the star-marked ![]() icon and interact with the intuitive AI Assistant to refine your queries. This feature is especially useful for those new to NSL or for accelerating complex searches.
icon and interact with the intuitive AI Assistant to refine your queries. This feature is especially useful for those new to NSL or for accelerating complex searches.
Conclusion
The Node Specification Language is an incredible tool for molecular modelers looking to maximize efficiency and precision in node selection. Whether you’re working on large molecular assemblies or small organic compounds, mastering NSL will streamline your workflow in SAMSON.
For a deeper dive into NSL, check out the full documentation at https://documentation.samson-connect.net/users/latest/nsl/.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Download it today at https://www.samson-connect.net.





