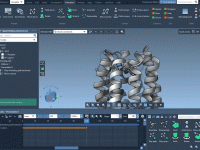
Simplifying Animation Node Management with SAMSON’s Animation Attributes
Managing animation nodes during molecular modeling projects can be a time-consuming process when you’re dealing with complex molecular structures. SAMSON, the integrative molecular design platform, offers a structured way to tackle this challenge using animation attributes. If you’ve ever struggled…