No Admin Rights? No Problem: Installing SAMSON Seamlessly
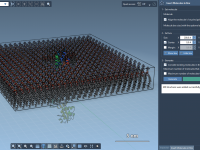
Building Lipid Bilayers Around Proteins in SAMSON
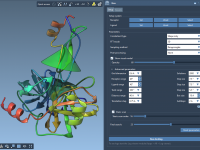
Choosing the Right Color Palette in Molecular Modeling: A Guide to Discrete Palettes in SAMSON
Quickly Filter and Control Light Nodes in SAMSON Using the NSL
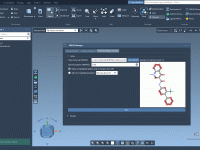
Swapping Atoms, Saving Time: Quick Analogue Generation in SAMSON
FASTA In, Structure Out: Predicting Proteins with AlphaFold-2 in SAMSON
Predicting protein structures used to be a long, resource-heavy process, often requiring local installations or complex data preparation. Today, especially with the introduction of AlphaFold-2, molecular modelers have access to powerful prediction engines remotely. However, integrating these capabilities into real…