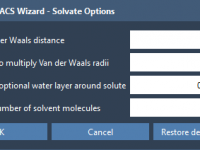
Solvating MARTINI Coarse-Grained Systems Without the Headaches
A Quick Way to Emphasize Molecular Events: Using the Flash Animation in SAMSON
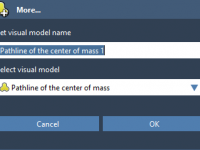
Visualizing Ligand Unbinding Routes with Pathlines in SAMSON
Quickly Find What’s Visible in Your Molecular Scene in SAMSON
How to Filter Side Chains by Atom Count in Molecular Models
When working with large molecular systems, quickly locating and analyzing specific side chains based on their atomic composition can become a challenge. Whether you’re interested in simplifying your view, isolating chemically relevant groups, or preparing a structure for coarse-grained simulations,…