Choosing the Right Symmetry Group in Molecular Assemblies
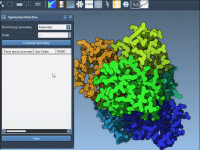
When working with large molecular assemblies like protein complexes or viral capsids, symmetry can act as a powerful ally. Recognizing repeating patterns through symmetry detection can simplify analyses, reduce computational demands, and enhance interpretation of experimental data. But what happens…
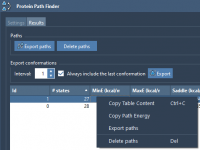
Exporting Molecular Data Doesn’t Have to Be a Hassle
Exporting molecular structures and visualizations is a common task for molecular modelers—whether to share simulation results, create publication-ready images, or prepare input files for other tools. Yet, dealing with incompatible formats and clunky export pipelines can easily become frustrating. SAMSON,…