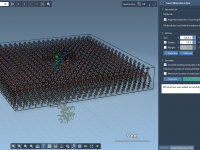
From Ring to Tube: Constructing Carbon Nanotubes Manually in SAMSON
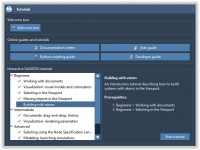
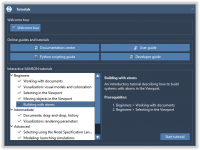
Designing nanostructures like carbon nanotubes can be tedious when you’re confined to repetitive manual replication and alignment. For computational chemists, nanomodelers, and molecular designers, the lack of intuitive pattern generation tools has been a long-standing bottleneck—especially when aiming to preserve…