If you’ve ever needed to analyze molecular trajectories for your simulations, such as tracking ligand movements through defined pathways or preparing input files for free energy calculations, SAMSON’s Export Along Paths extension might just be what you’re looking for. This tool simplifies the complex task of exporting atomic coordinates along specific paths, saving both time and effort. Curious how it works? Let’s dive in.
Why Export Atom Trajectories?
Tracking atomic movements is essential in understanding molecular processes like ligand unbinding or reaction pathways. With SAMSON’s Export Along Paths, you can export coordinates for all or selected atoms as they travel along paths, enabling you to:
- Generate reaction coordinate files for free energy profiling.
- Export ligand entry/exit trajectories for enhanced sampling simulations.
- Visualize intermediate states along a pathway.
- Focus on specific atoms, such as a ligand or region of interest.
By using this feature, you can easily extract valuable trajectory data for further analysis, simulation, or visualization.
Getting Started
To begin, ensure the Export Along Paths app is installed and ready in your SAMSON environment. Follow these steps:
- Log into SAMSON Connect.
- Visit the Export Along Paths extension page and click Add.
- Restart SAMSON so that the extension is accessible in your workspace.
Exporting Atom Trajectories: Step-by-Step
To explain the workflow, we’ll use the provided tutorial sample. This includes a molecular model, Lactose permease (1PV7), its ligand TDG, and unbinding paths generated using the Ligand Path Finder extension.
Loading the Sample System
First, load the tutorial sample system:
- Go to Home > Download.
- Paste the provided link: https://www.samson-connect.net/documents/046f1acd-c799-40f6-8185-cb4847eff795.
- Click Download.

You’ll now see a structural model containing the protein Protein_chain_A, the ligand TDG, and predefined unbinding paths.
Exporting the Trajectories
Once your system is loaded, open the Export Along Paths app (Home > Apps > All > Export Along Paths):
![]()
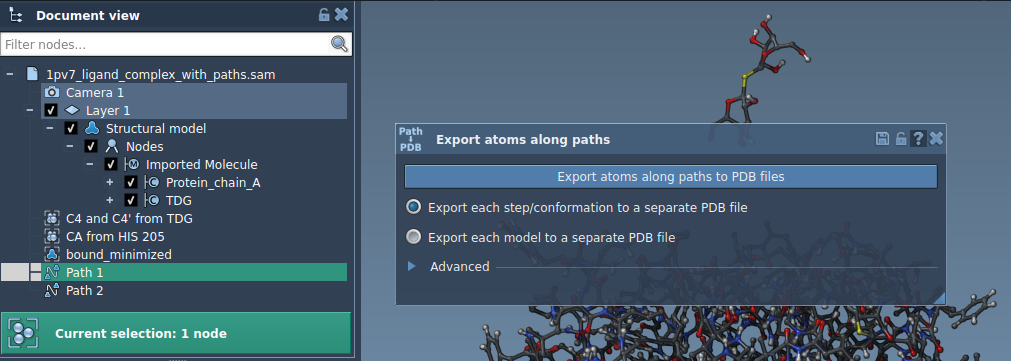
The export process is flexible and has two primary options:
Option 1: Export All Atoms
- Select one or more paths in the Document view.
- Choose your export mode:
- All frames in a single PDB file.
- Each frame as a separate PDB file.
- Click Export atoms along paths to PDB files and specify the destination folder and file name prefix.
Option 2: Export a Subset of Atoms
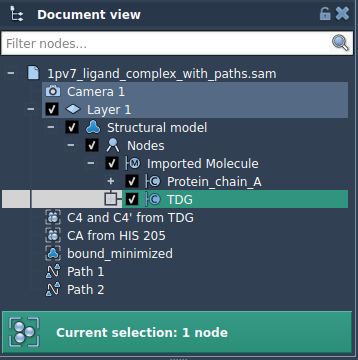
If you’re only interested in specific atoms (e.g., the ligand):
- Expand the Advanced panel in the app.
- Select the molecule of interest (e.g.,
TDG) in the Document view. - Click Add to include it as a model for export.
You can rename models, add multiple sets of atoms, and adjust the frame export intervals as needed.
Once configured:
- Choose export format (single or multiple PDB files).
- Select the relevant path(s).
- Click Export atoms along paths to PDB files.
It’s that simple!
Conclusion
Exporting atomic trajectories is now a streamlined process with the Export Along Paths extension in SAMSON. Whether you’re working with ligand pathways or want to export subsets of atoms for detailed analysis, this tool equips you with the functionality you need for free energy profiling, enhanced simulations, and more.
For more detailed instructions, you can explore the full documentation page here.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. Get SAMSON at https://www.samson-connect.net.





