For molecular modelers working with proteins, understanding symmetry is often a key step in designing molecular structures or simulating interactions. Crystallographic units and biological assemblies provide critical insights into protein organization, but deriving symmetry mates from these data can sometimes be a challenging task. Enter the Symmetry Mate Editor in SAMSON, a user-friendly tool that simplifies the process of generating symmetric replicas of proteins from PDB files.
The Pain Point: Decoding Protein Symmetry
When exploring protein structures, researchers frequently encounter two kinds of symmetry-related information in PDB files: CRYST1 (which describes symmetry within the crystal lattice) and BIOMT (which provides biological assembly annotations). Reconstructing protein symmetries, visualizing biological assemblies, or creating complex structures often requires painstaking manual work or reliance on fragmented tools. This can slow down workflows significantly.
The Symmetry Mate Editor in SAMSON addresses this pain by allowing users to effortlessly generate symmetry mates and explore protein symmetries directly in an intuitive graphical interface.
How the Symmetry Mate Editor Works
Once you’ve added the Symmetry Mate Editor extension to SAMSON and activated it, you’ll gain access to powerful tools for exploring and generating symmetry mates. Here’s an overview of the main features:
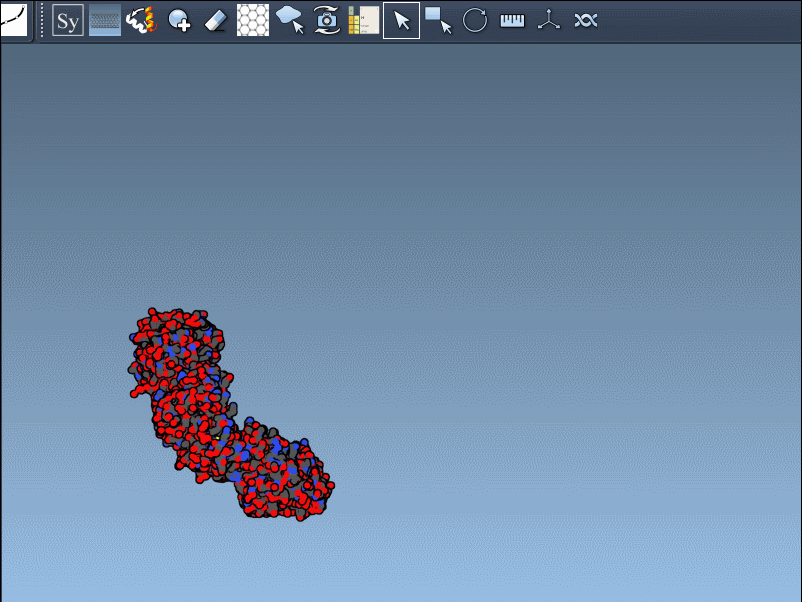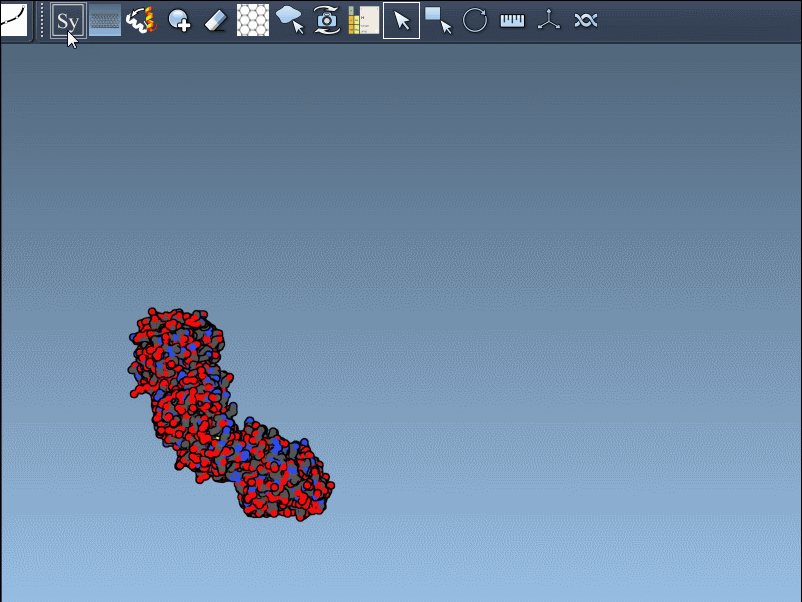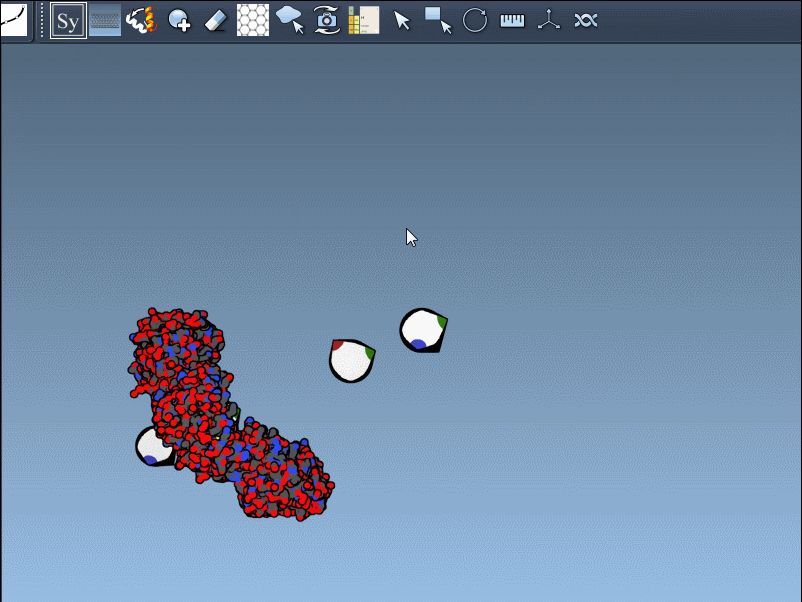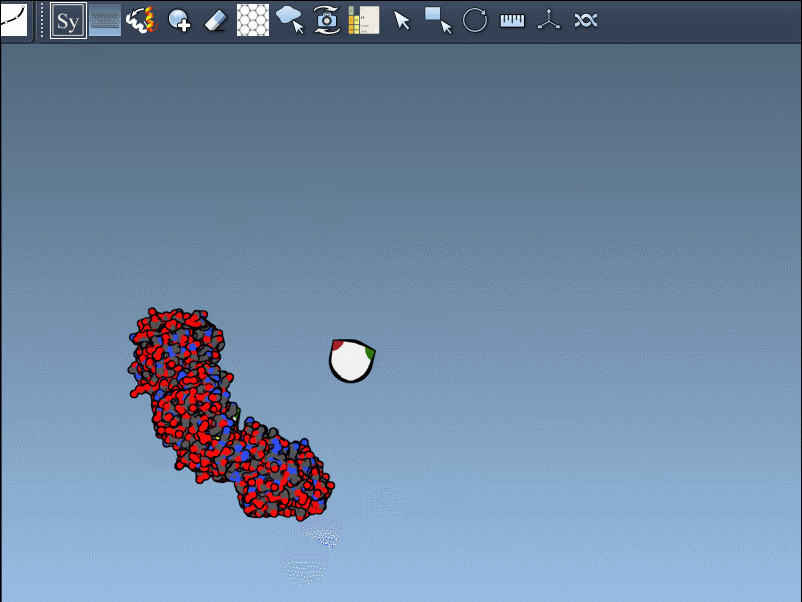
- Control nodes: When you activate the editor within SAMSON (via Find Everything or the Editors Menu), control nodes representing symmetry transformations become visible in the viewport. These nodes offer interactive options for previewing or generating replicas.
- Scaling control nodes: You can scale the number of visible control nodes by using Ctrl/Cmd + mouse wheel, allowing finer adjustments to your visual exploration of symmetry.
- Preview and click: By hovering over a node, you can preview the corresponding symmetric replica in real time. Left-clicking a node permanently generates that replica, enabling step-by-step construction of composite protein structures.
CRYST1 and BIOMT – A Versatile Implementation
The Symmetry Mate Editor supports both CRYST1 and BIOMT records, covering a broad spectrum of structural symmetries. Each record type is color-coded in the interface for clarity:
| Record type | Description | Widget color |
|---|---|---|
| CRYST1 | Symmetry from crystal lattice | White |
| BIOMT | Symmetry from biological assembly annotations | Yellow |
In practice, switching between CRYST1 and BIOMT is straightforward; it’s as simple as toggling between options in the editor’s interface. Here’s an example showcasing this dual functionality:
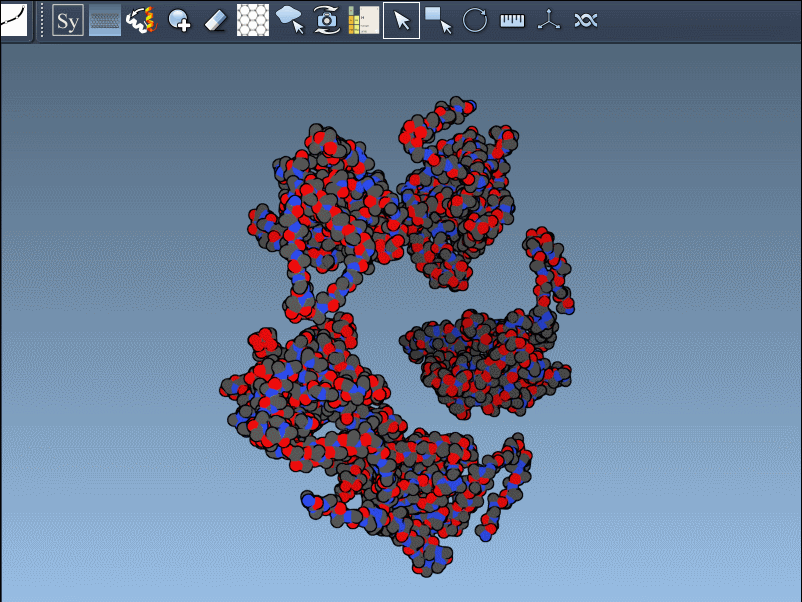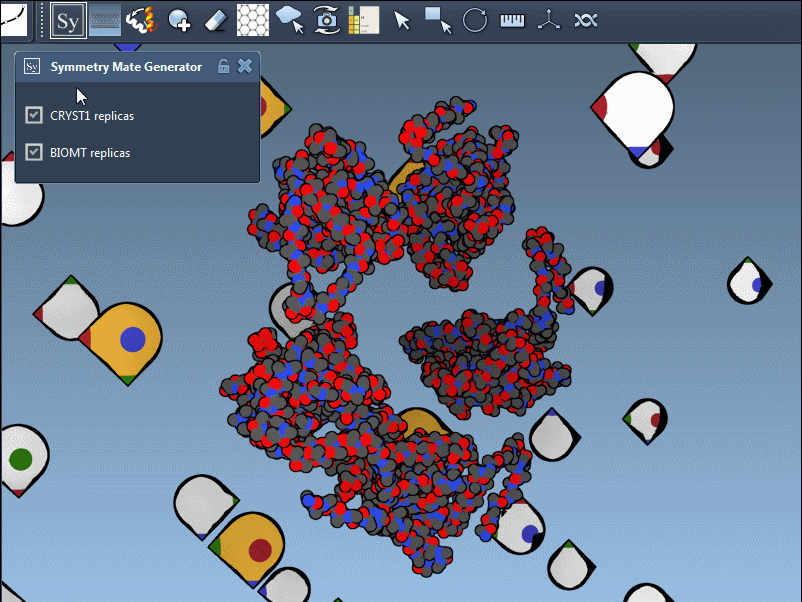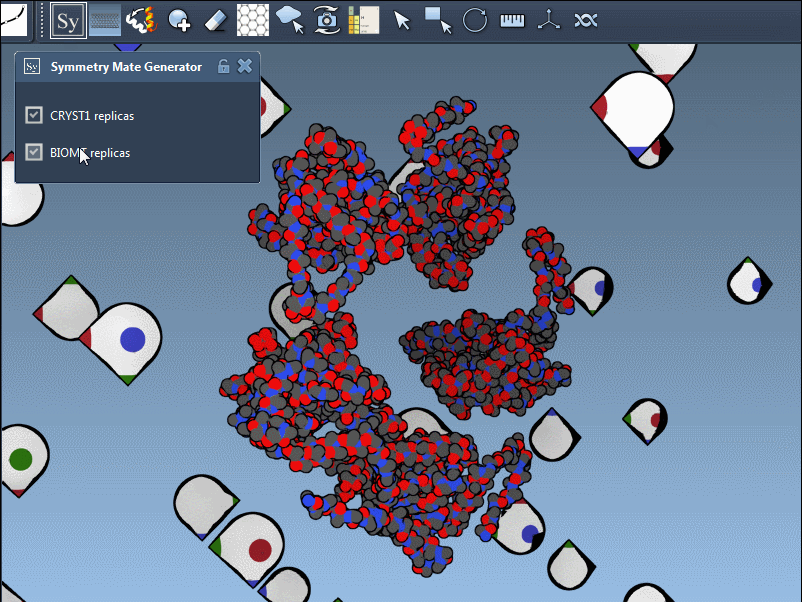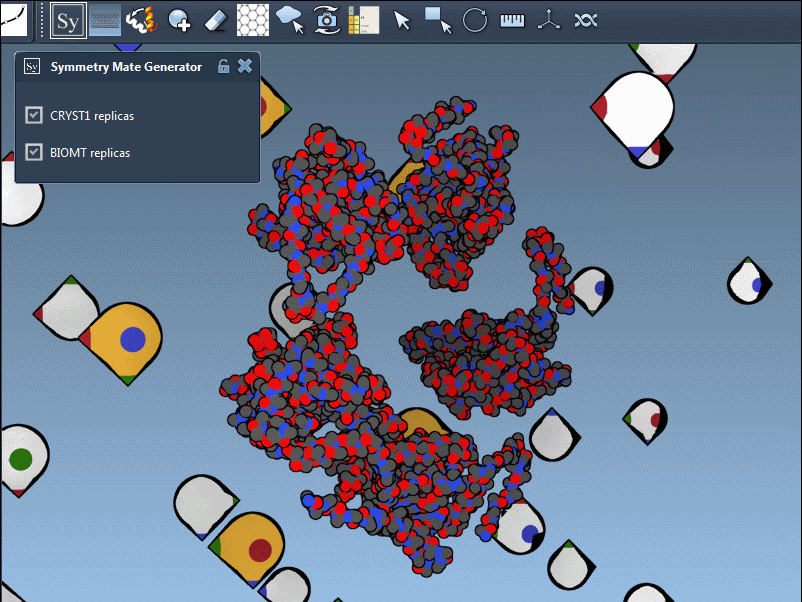
Why This Matters
Generating symmetry mates in a highly interactive environment is not just about convenience—it opens up possibilities for advanced molecular modeling workflows:
- Reconstruct complete biological assemblies from asymmetric units.
- Explore protein-protein interfaces and identify potential binding sites.
- Design symmetric protein complexes and examine their stability or function.
Learn More
The Symmetry Mate Editor is designed to smooth out a common roadblock for molecular modelers: working with protein symmetries efficiently. With features like real-time symmetry previews, record-specific support, and intuitive interaction, it helps you focus on higher-level structural analyses with less effort.
You can check out the official documentation to learn more about using the Symmetry Mate Editor.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. To get started, visit SAMSON Connect.





