If you are a molecular modeler or researcher studying SARS-CoV-2, understanding the motion of the viral spike protein is crucial. The spike protein is central to the virus’s ability to infect human cells, transitioning from a closed state to an open configuration that enables binding to the ACE2 receptor. But modeling this motion accurately can be tricky and time-consuming—especially with the complexity of the spike’s structure and its interactions. In this post, we’ll explore how SAMSON helps tackle this challenge through its computational tools, facilitating in-depth investigations of the spike protein’s behavior.
Why Model Spike Motion?
The spike is not just structural—it actively facilitates viral entry into human cells. By shifting between closed and open conformations, it unveils its receptor-binding domain, allowing it to connect with ACE2, an essential receptor on the surface of epithelial cells in the lungs and other organs. Consequently, this makes modeling its motion pivotal for understanding the mechanisms behind infection and for designing therapeutic interventions such as neutralizing antibodies or antiviral drugs.
Below, we’ll break down how SAMSON computationally models the spike protein’s transition between these states and how you can access tools to reproduce and build upon this research.
Showcasing the Spike in Motion
To model the spike’s transformative journey between open and closed states, SAMSON uses a computational pipeline combining two specialized approaches: the As-Rigid-As-Possible (ARAP) Interpolation Path module and the Parallel Nudged Elastic Band (P-NEB) module. These steps ensure both accuracy and efficiency. Here’s an outline of the workflow:
- Starting Points: The closed (PDB 6VXX) and open (PDB 6VYB) states of the spike are used. Since the structures differ slightly in the number of residues involved, the first step involves refining them. Bond orders and hydrogen atoms are adjusted, and minimization steps are carried out to ensure consistency.
- Interpolating Motion with ARAP: The ARAP module generates a smooth interpolation path between the open and closed configurations. This intermediate trajectory allows modelers to visualize the motion dynamically.
- Refining the Path with P-NEB: To increase accuracy, the P-NEB module enhances the interpolated path by optimizing the structures along the trajectory, creating realistic intermediate forms as the spike transitions between states.
The result? You get a detailed representation of the spike protein’s dynamics, enabling deeper insights into its biological behavior during infection.
Interactive Visualization Examples
The animations below depict the spike transition between its closed (down) and open (up) states. These views illustrate how the spike prepares itself to bind with the ACE2 receptor:
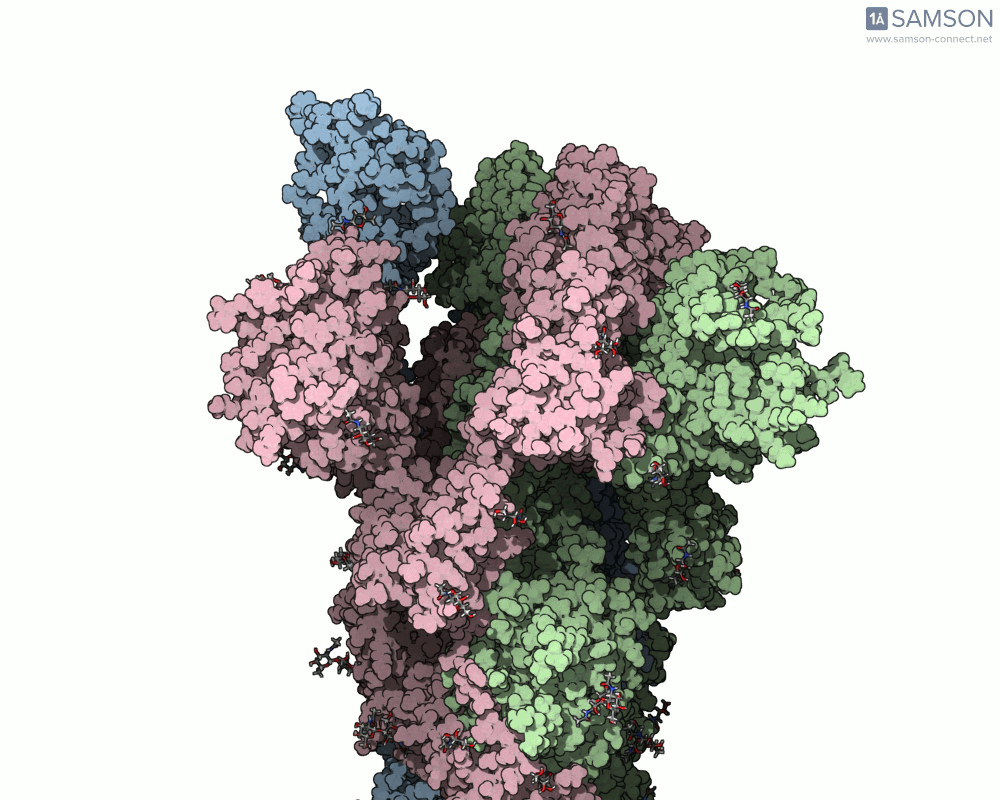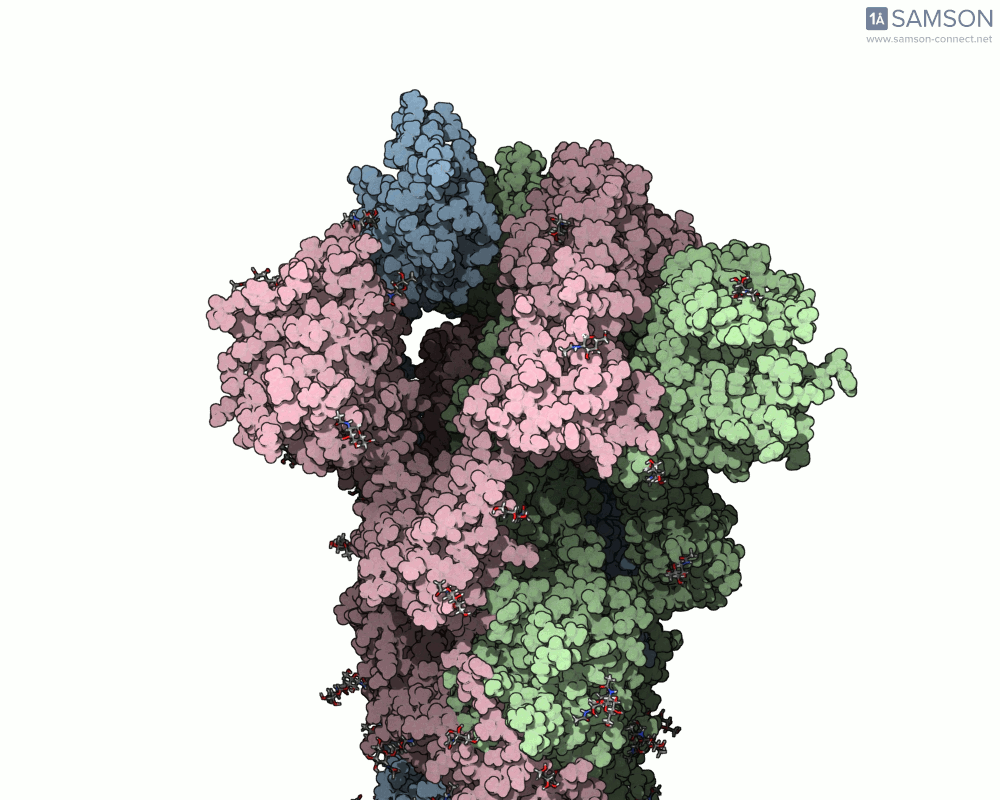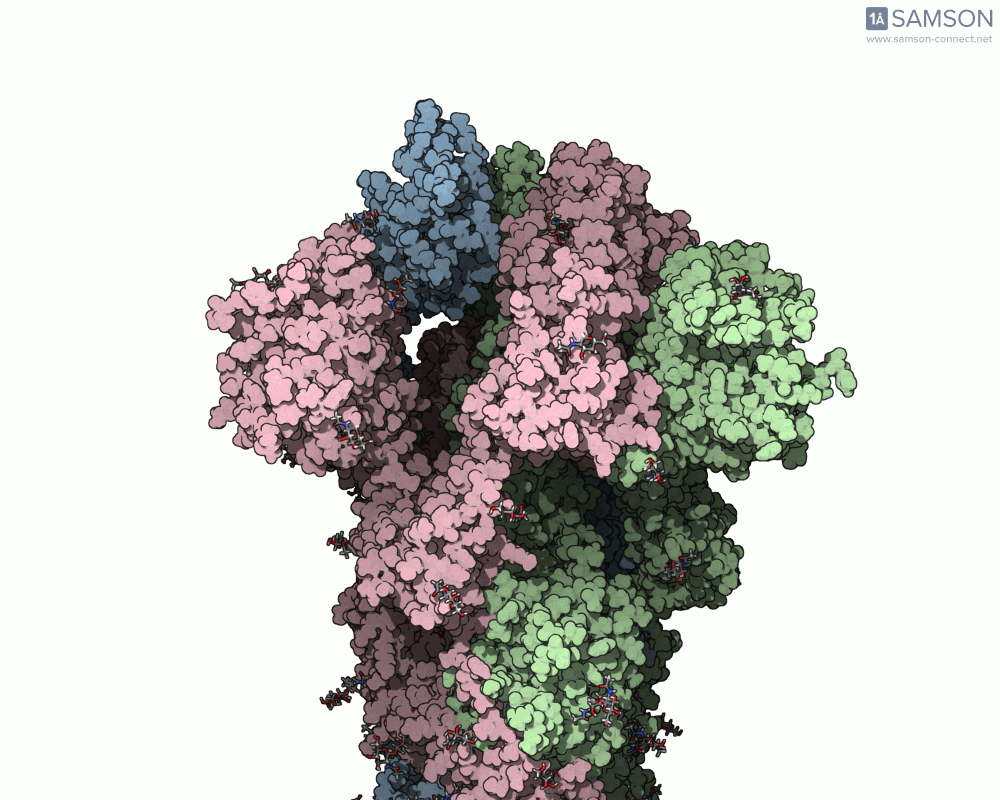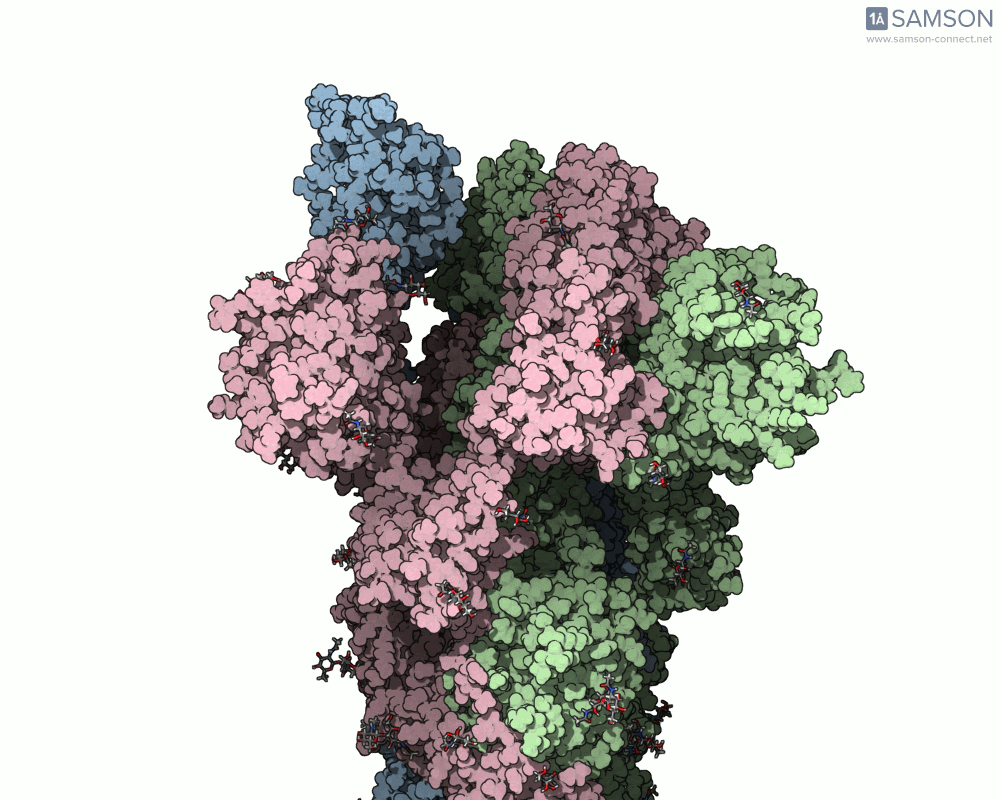
These animations are particularly useful for researchers aiming to understand how molecular interactions occur during the opening and closing of the spike protein.
How to Access These Tools
For modelers interested in recreating or building upon this research, SAMSON offers free access to the ARAP and P-NEB modules during the outbreak. You can sign up on SAMSON Connect, download the platform, and install the tools. Additionally, trajectory files such as PDB formats of the motion are available directly here for further experiments and analyses.
To learn more, visit the full documentation: https://documentation.samson-connect.net/tutorials/sars-cov-2/coronavirus-computing-the-opening-motion-of-the-sars-cov-2-spike/.
Note: SAMSON and all SAMSON Extensions are free for non-commercial use. You can get SAMSON at https://www.samson-connect.net.





