Author: OneAngstrom
Working with Many Conformations at Once in Molecular Simulations
Avoid Restart Headaches: Smart Input Selection for NVT Equilibration in GROMACS Wizard
When Your Molecules Start to Rock: A Simple Way to Show Local Motion in 3D
Stop Losing Your Molecules: How to Keep the Camera Focused on Moving Atoms
Integrating External Tools in SAMSON: A Flexible Approach to Molecular Modeling
Orbiting Molecules: Making Cleaner Molecular Presentations with the Orbit Camera
Preparing Multiple Proteins in One Go with SAMSON: A Quick Guide to Batch Protein Preparation
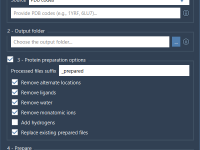
If you’re working with molecular docking, structural analysis, or simulations across many proteins, you’ve likely run into this challenge: preparing dozens or even hundreds of raw PDB files manually. Stripping water, removing unnecessary ligands, adding hydrogens, resolving alternate locations… doing…
A Simple Way to Animate Molecular Docking in Your Presentations
How to Explore Protein Transition Paths and Visualize Their Energies
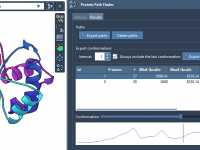
Understanding how proteins change shape between two conformations is a challenge faced by many molecular modelers. These conformational changes are essential for protein function, but identifying plausible transition pathways and analyzing their energy profiles can be time-consuming and complex. The…