Author: OneAngstrom
Mastering Horizontal Camera Movements in Molecular Visualization
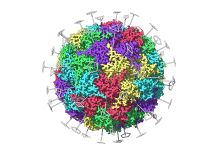
Why Symmetry Detection Matters in Molecular Complexes
Mastering Node Selection with NSL in SAMSON.
Smooth Protein Transitions with ARAP Interpolation: A Guide
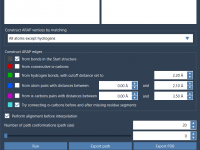
For molecular modelers, creating realistic and biologically meaningful transition paths between protein conformations can be challenging and time-consuming. Whether you’re working on conformational analysis, transition state modeling, or setting up an umbrella sampling workflow, an efficient and precise interpolation method…