Category: Uncategorized
Speeding up Nanotube Design with Pattern Editors in SAMSON
For molecular modelers working on nanostructures, building carbon nanotubes and similar architectures atom by atom is time-consuming and error-prone. Small misalignments between atoms can create structurally incoherent models and increase the time required for cleanup and minimization. If your work…
Quickly Filter Molecular Side Chains by Atom Type in SAMSON
When dealing with large biomolecular models, pinpointing specific side chains with certain atomic compositions can be tedious and time-consuming. SAMSON’s Node Specification Language (NSL) provides a concise way to filter and select these molecular entities efficiently. Whether you’re preparing systems…
Why Protein Cleanup Matters Before Structural Interpolation
From Steel to Glass: Fine-Tuning Molecular Materials with Cycles in SAMSON
When Molecules Fade Away: Using ‘Conceal atoms’ to Focus Your Story
Python Scripting in SAMSON: Automate, Simulate, Repeat
How to Select Charged, Polar, or Hydrophobic Residues in Seconds
Tired of Misaligned Molecules? Align Structures in Global Reference Frame in Seconds
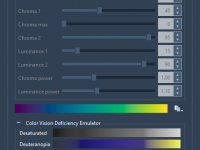
What Molecular Modelers Should Know About HCL Color Palettes in SAMSON
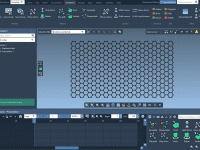
Speeding up Nanotube Design with Pattern Editors in SAMSON

For molecular modelers working on nanostructures, building carbon nanotubes and similar architectures atom by atom is time-consuming and error-prone. Small misalignments between atoms can create structurally incoherent models and increase the time required for cleanup and minimization. If your work…
Quickly Filter Molecular Side Chains by Atom Type in SAMSON
When dealing with large biomolecular models, pinpointing specific side chains with certain atomic compositions can be tedious and time-consuming. SAMSON’s Node Specification Language (NSL) provides a concise way to filter and select these molecular entities efficiently. Whether you’re preparing systems…