Category: Uncategorized
Making Time Stand Still: Using the Pause Animation in Your Scientific Presentations
Choosing the Right Symmetry Group for Protein Assemblies
Changing Atom Attributes Without Losing Context
Make Your Molecular Modeling Profile Work for You

Creating and managing a public scientific profile can be time-consuming, especially when your research contributions are spread across multiple platforms. For molecular modelers involved in interdisciplinary or collaborative projects, keeping everything centralized and accessible often becomes a task on its…
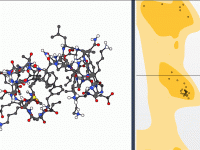
Editing Protein Backbone Geometry Directly from a Ramachandran Plot
Smaller Boxes, Faster Simulations: Choosing the Right Unit Cell Shape for Molecular Models
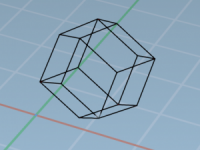
When setting up molecular simulations, one challenge that modelers often face is finding the right balance between accuracy and computational efficiency. Many simulation tasks—especially when working with spherical or flexible macromolecules like proteins—require significant solvent padding to respect periodic boundary…
Filtering Molecules by Structure: Speed Up Your Modeling Workflow
Save Time When Starting from a Single Molecule in SAMSON
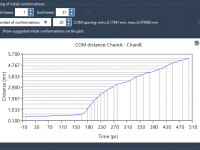
Avoid Common Mistakes When Setting Up Protein–Ligand Systems for Unbinding Pathway Analysis
A frequent challenge in molecular modeling is preparing a protein–ligand system for unbinding pathway simulations. Even experienced users often encounter issues with misassigned atoms or poorly defined conformations, which can invalidate simulation results. Fortunately, the Ligand Path Finder app in…
Making Time Stand Still: Using the Pause Animation in Your Scientific Presentations
Choosing the Right Symmetry Group for Protein Assemblies
Changing Atom Attributes Without Losing Context
Make Your Molecular Modeling Profile Work for You

Creating and managing a public scientific profile can be time-consuming, especially when your research contributions are spread across multiple platforms. For molecular modelers involved in interdisciplinary or collaborative projects, keeping everything centralized and accessible often becomes a task on its…
Editing Protein Backbone Geometry Directly from a Ramachandran Plot
Smaller Boxes, Faster Simulations: Choosing the Right Unit Cell Shape for Molecular Models

When setting up molecular simulations, one challenge that modelers often face is finding the right balance between accuracy and computational efficiency. Many simulation tasks—especially when working with spherical or flexible macromolecules like proteins—require significant solvent padding to respect periodic boundary…
Filtering Molecules by Structure: Speed Up Your Modeling Workflow
Save Time When Starting from a Single Molecule in SAMSON
Avoid Common Mistakes When Setting Up Protein–Ligand Systems for Unbinding Pathway Analysis
A frequent challenge in molecular modeling is preparing a protein–ligand system for unbinding pathway simulations. Even experienced users often encounter issues with misassigned atoms or poorly defined conformations, which can invalidate simulation results. Fortunately, the Ligand Path Finder app in…