Understanding and Using Active Editors in SAMSON
Streamline Protein-Ligand Modeling with 2D Interaction Diagrams
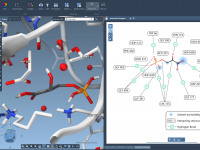
For researchers and molecular modelers working with protein-ligand interactions, visualizing and exploring molecular interactions effectively is essential yet often challenging. The SAMSON Interaction Designer offers a user-friendly way to bridge the gap between 3D and 2D molecular modeling, enabling researchers…
Mastering Transparency with the Appear Animation
Effortlessly Zoom in on Molecular Models Without Changing Your Perspective
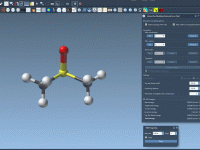
Simplifying Bond Adjustments in Molecular Models with SAMSON
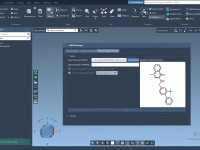
Replace Molecular Patterns with Ease Using the SMILES Manager
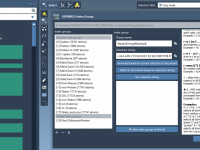
Mastering Folder Attributes for Efficient Molecular Design
For molecular modelers working with complex structures and data, managing folders effectively can significantly streamline the design process. SAMSON, the integrative molecular design platform, provides a robust and intuitive system for working with folder attributes through its Node Specification Language…