Simplifying Umbrella Sampling With GROMACS Wizard in SAMSON
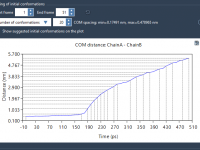
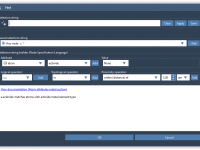
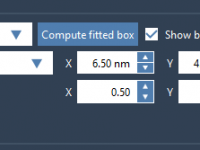
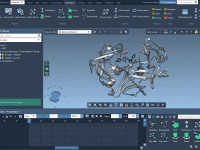
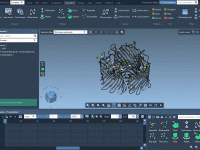
For molecular modelers exploring complex reaction coordinates, umbrella sampling can be an essential approach for free energy calculations. However, setting up umbrella sampling often means juggling multiple configurations, trajectories, and computational setups. The GROMACS Wizard in SAMSON simplifies this intricate…