Easily Export DNA Nanostructures for Simulations with Adenita
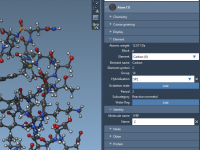
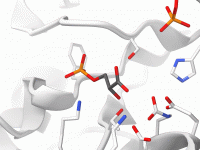
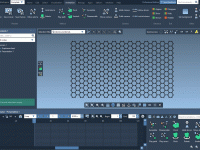
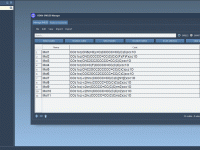
For molecular modelers working with DNA nanostructures, exporting designs for simulations can be a repetitive and complex task. Often, you must navigate different file formats, manual configurations, and inconsistencies between design tools and simulation platforms. Adenita, the DNA nanostructure modeling…